Classification rationale
1
The PTEN c.-307C>G (NP_000305.3:p.?) variant has not been observed in somatic cancers in COSMIC and has not been reported in ClinVar.
clinvar ↗2
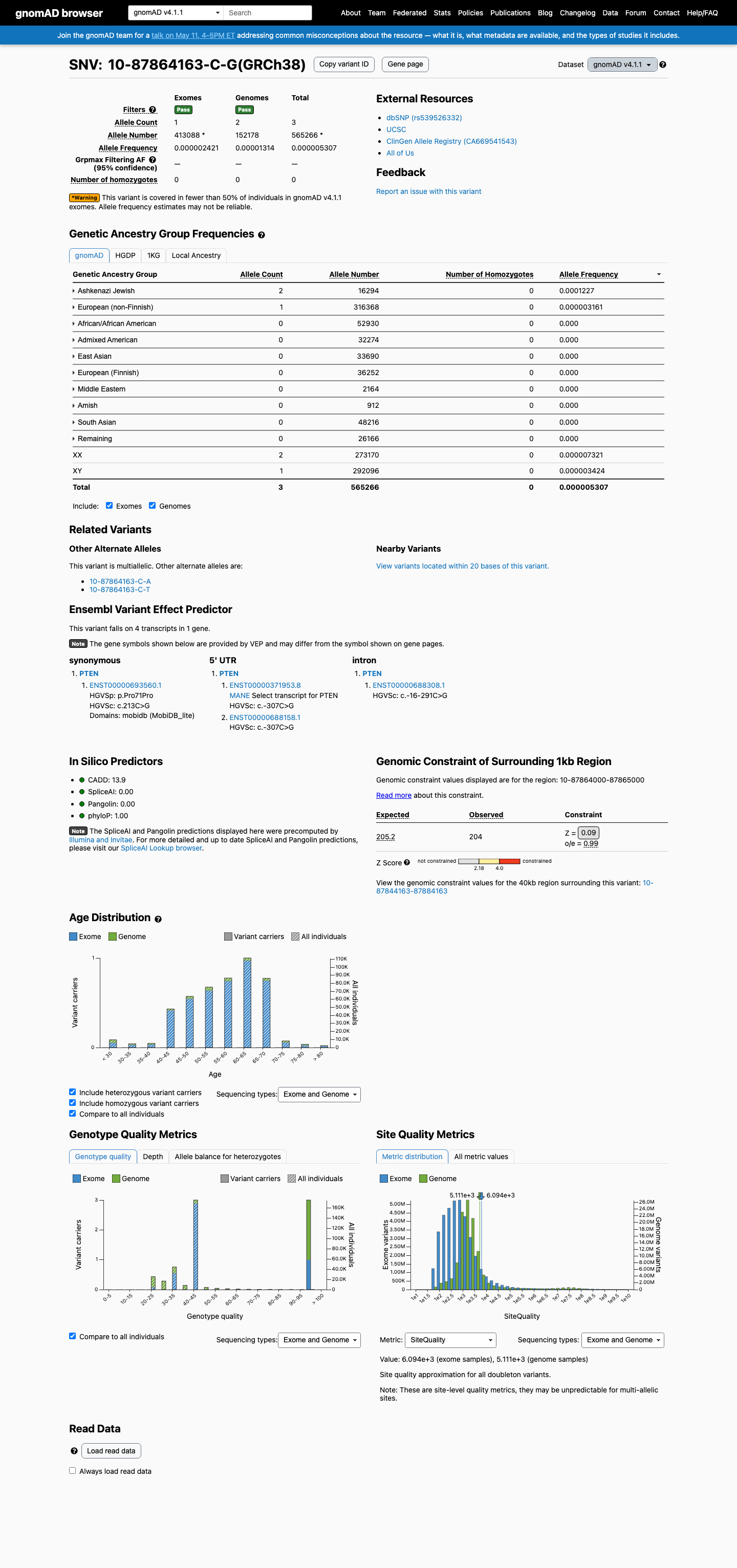
This variant is absent from gnomAD v2.1, but in gnomAD v4.1 it is present at an overall allele frequency of 5.31e-06 and reaches 1.23e-04 in the Ashkenazi Jewish population, which meets the PTEN BS1 strong frequency range and is above the PTEN PM2 subpopulation threshold.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
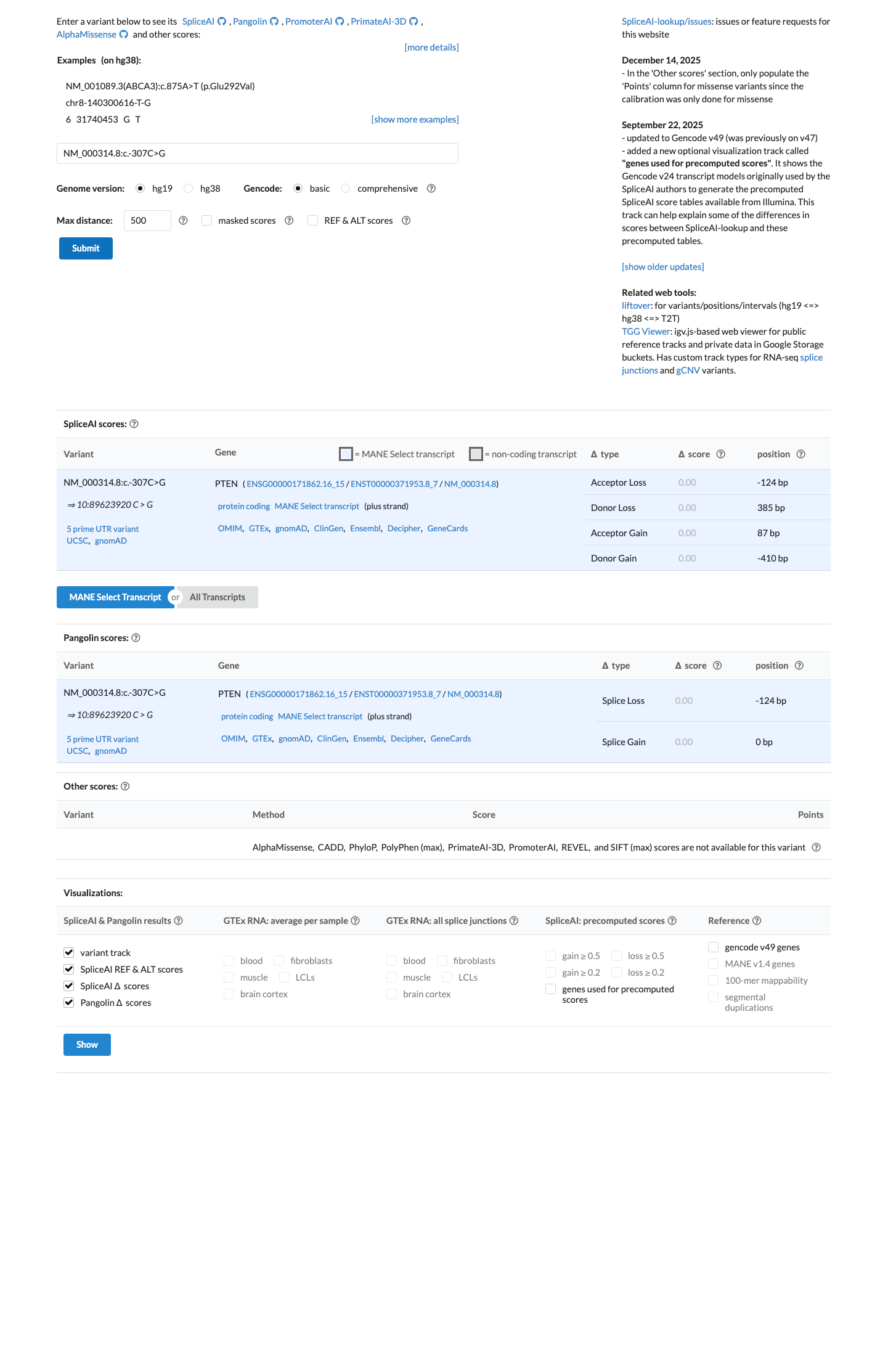
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.00, and no PTEN-specific computational evidence supporting a deleterious effect was identified.
spliceai ↗ cspec ↗