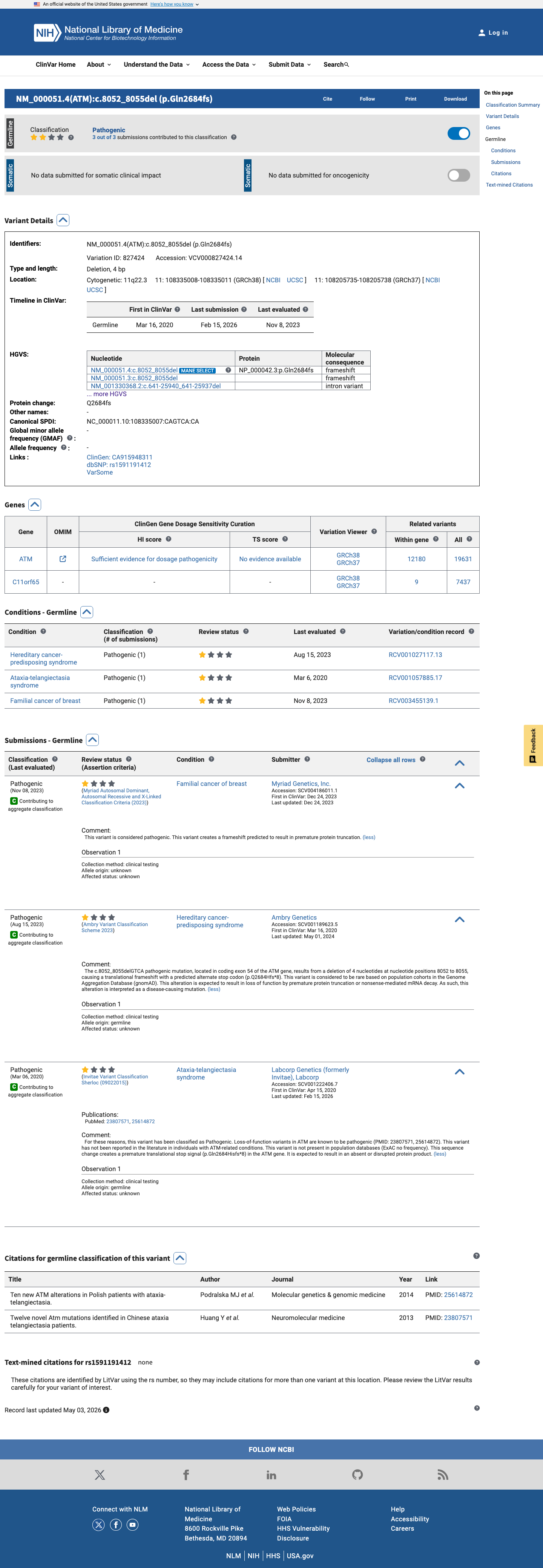
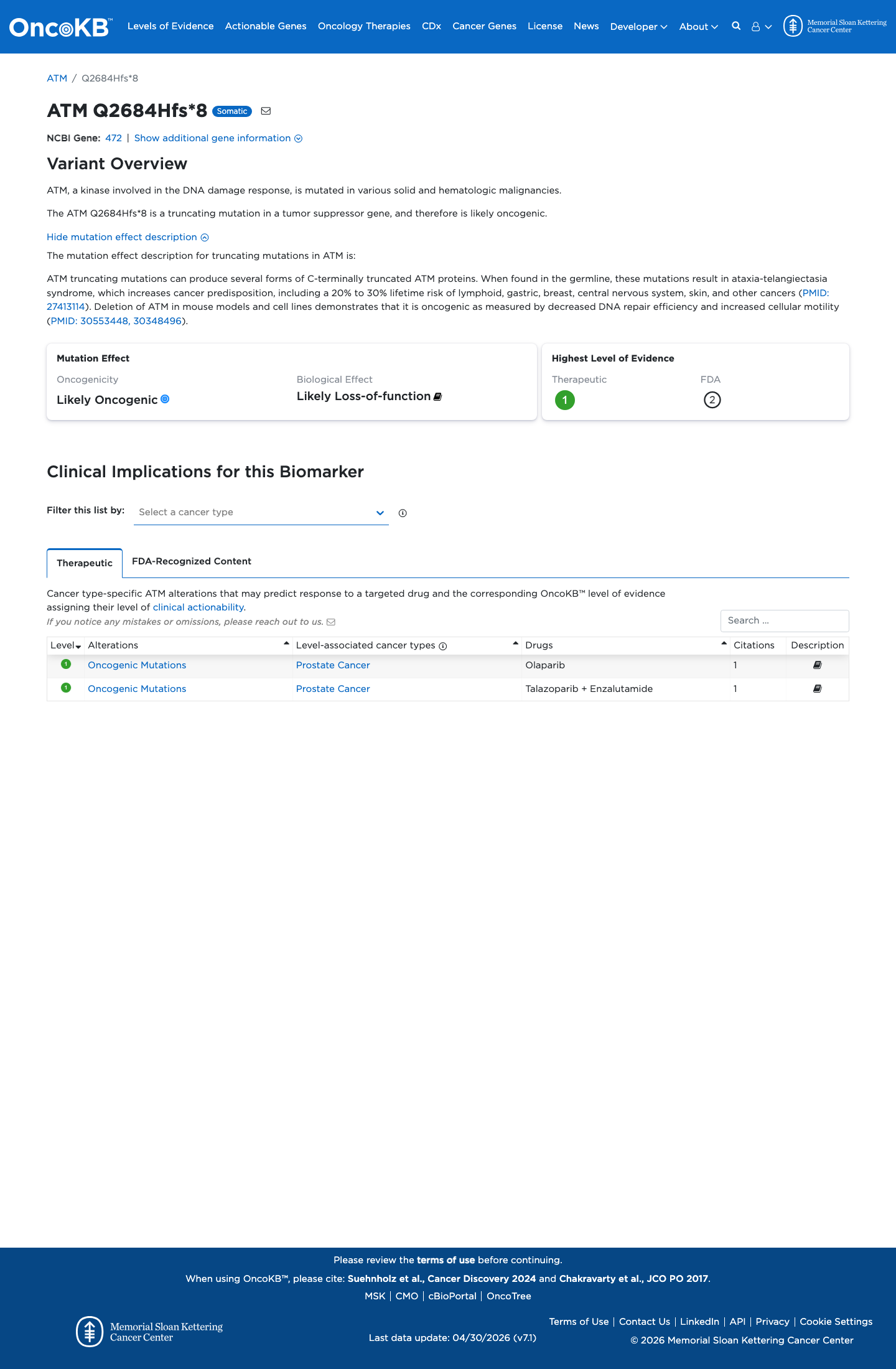
The ATM c.8052_8055del (p.Gln2684HisfsTer8; p.Q2684Hfs*8) variant has not been observed in somatic cancers in COSMIC and is reported in ClinVar as pathogenic.
clinvar ↗This variant is absent from gnomAD v2.1 and v4.1, which is below the ATM PM2_Supporting threshold of ≤0.001% in gnomAD v4 and does not meet the BS1 (>0.05%) or BA1 (>0.5%) population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗This 4-bp deletion causes a frameshift in exon 55 with a premature stop codon, and under the ATM VCEP framework that supports PVS1 at very strong strength; because the predicted stop is upstream of p.Arg3047, it also meets the ATM-specific truncation-based PM5 rule.
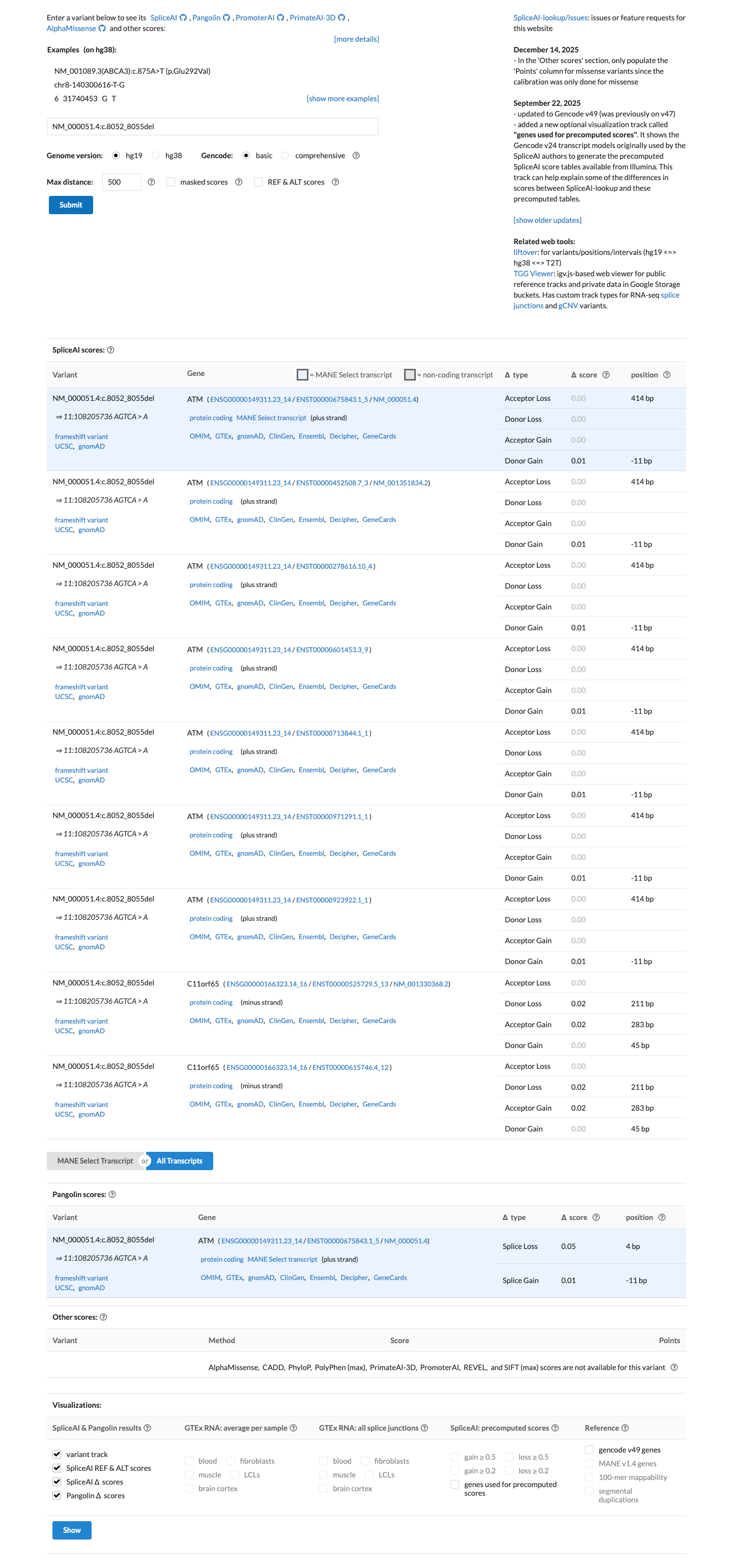
cspec ↗SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, and the ATM PP3/BP4 computational rules are not applied to this exonic frameshift variant class.
spliceai ↗ cspec ↗