Classification rationale
1
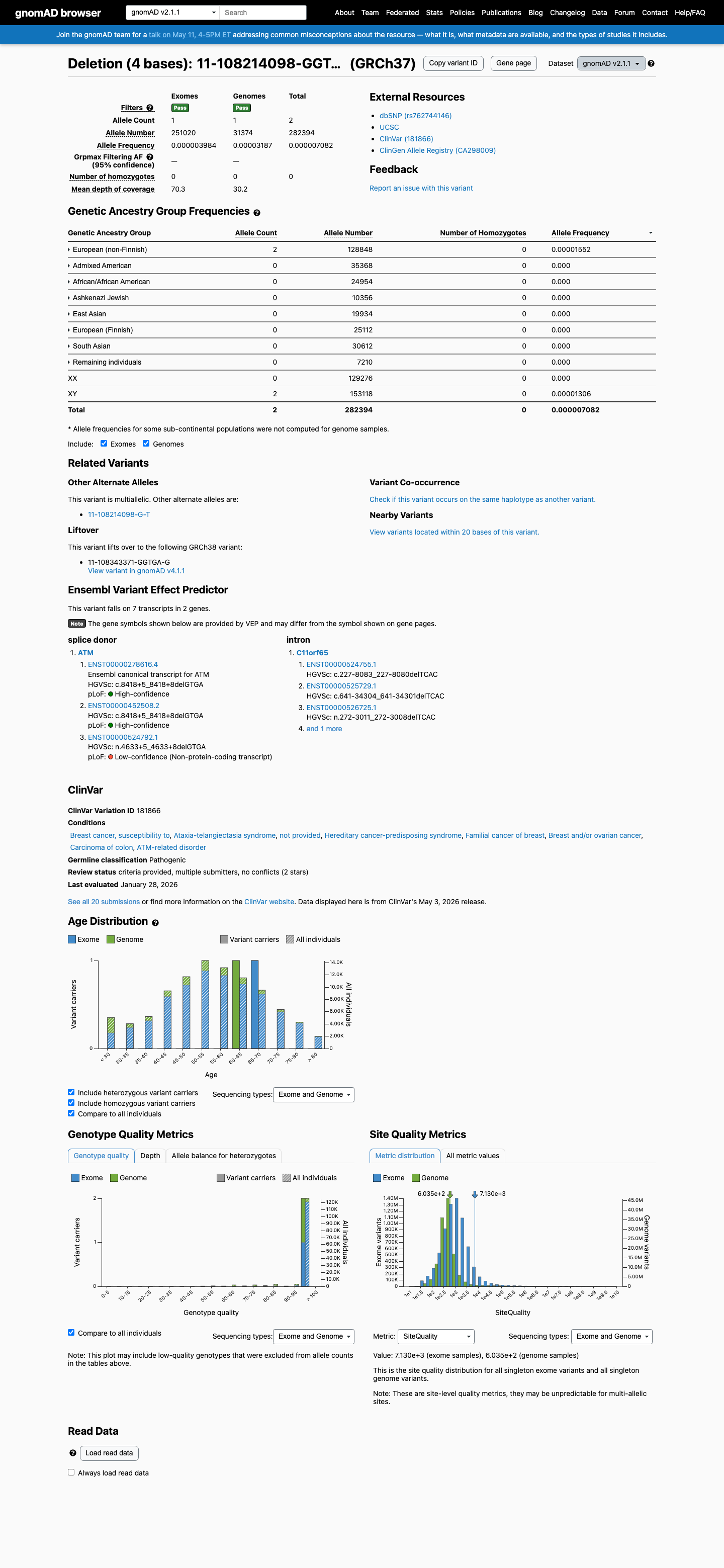
The ATM c.8418+5_8418+8del (NP_000042.3:p.?) variant has not been reported in ClinVar.
clinvar ↗2
This variant is present in gnomAD v4.1 at AF 0.00465% (75/1,613,598 alleles; grpmax FAF 0.005111%) and in gnomAD v2.1 at AF 0.00071% (2/282,394 alleles), which is above the ATM PM2_Supporting threshold of 0.001% and below the BS1 threshold of 0.05%.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗3
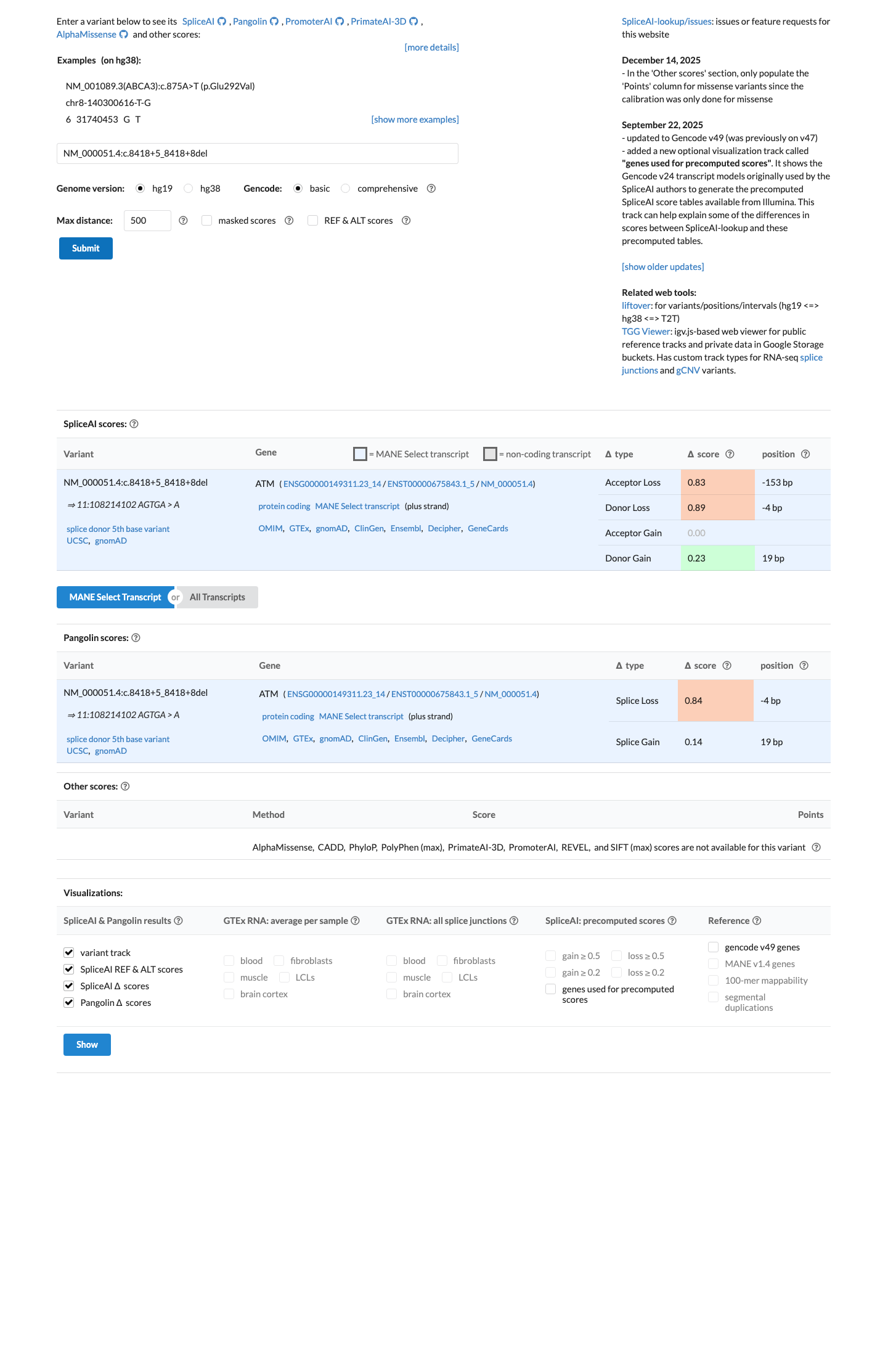
SpliceAI predicts an adverse splicing effect with a maximum delta score of 0.89, which exceeds the ATM PP3 threshold of 0.2 for intronic variants outside the donor ±1,2 positions and supports PP3.
spliceai ↗ cspec ↗