Classification rationale
1
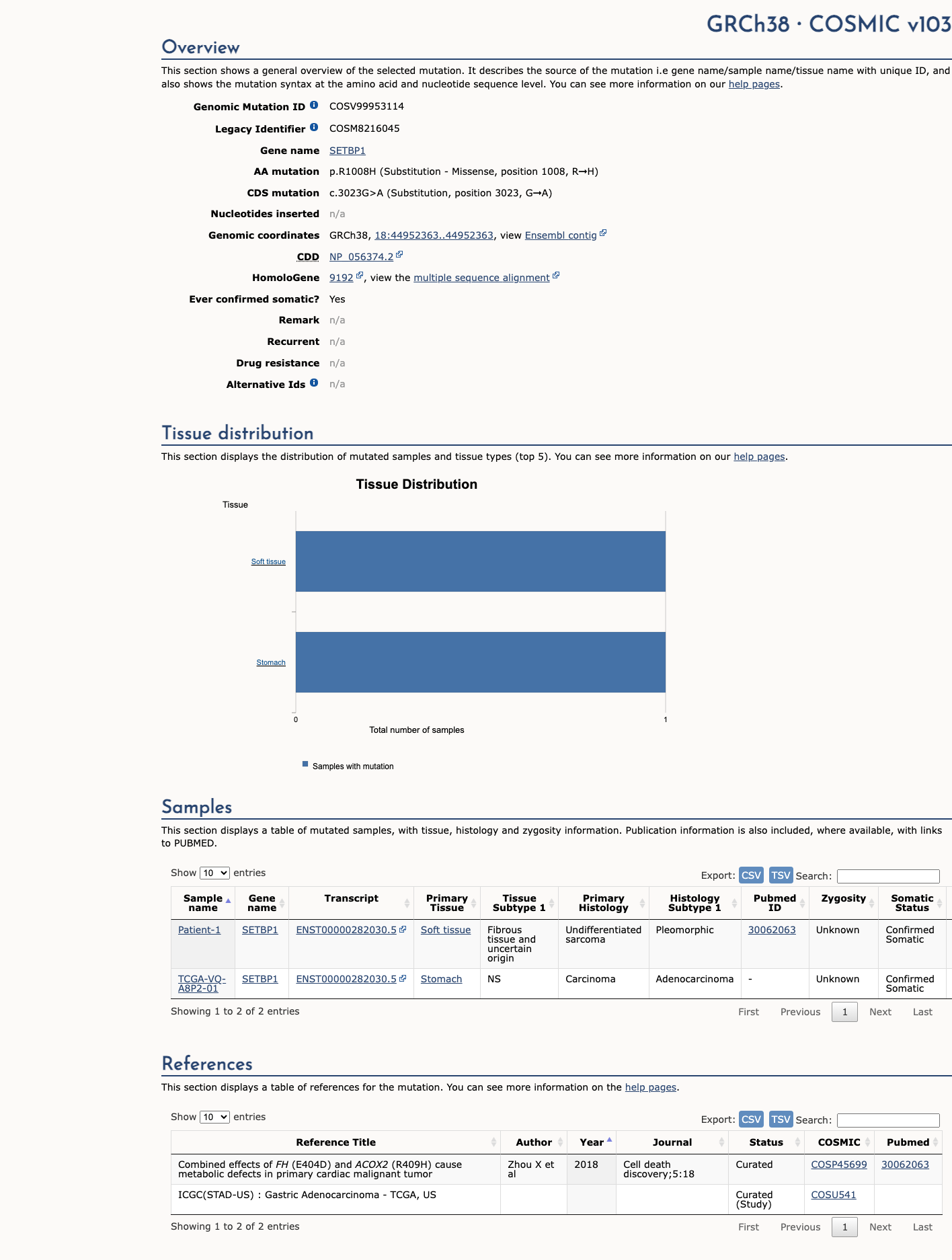
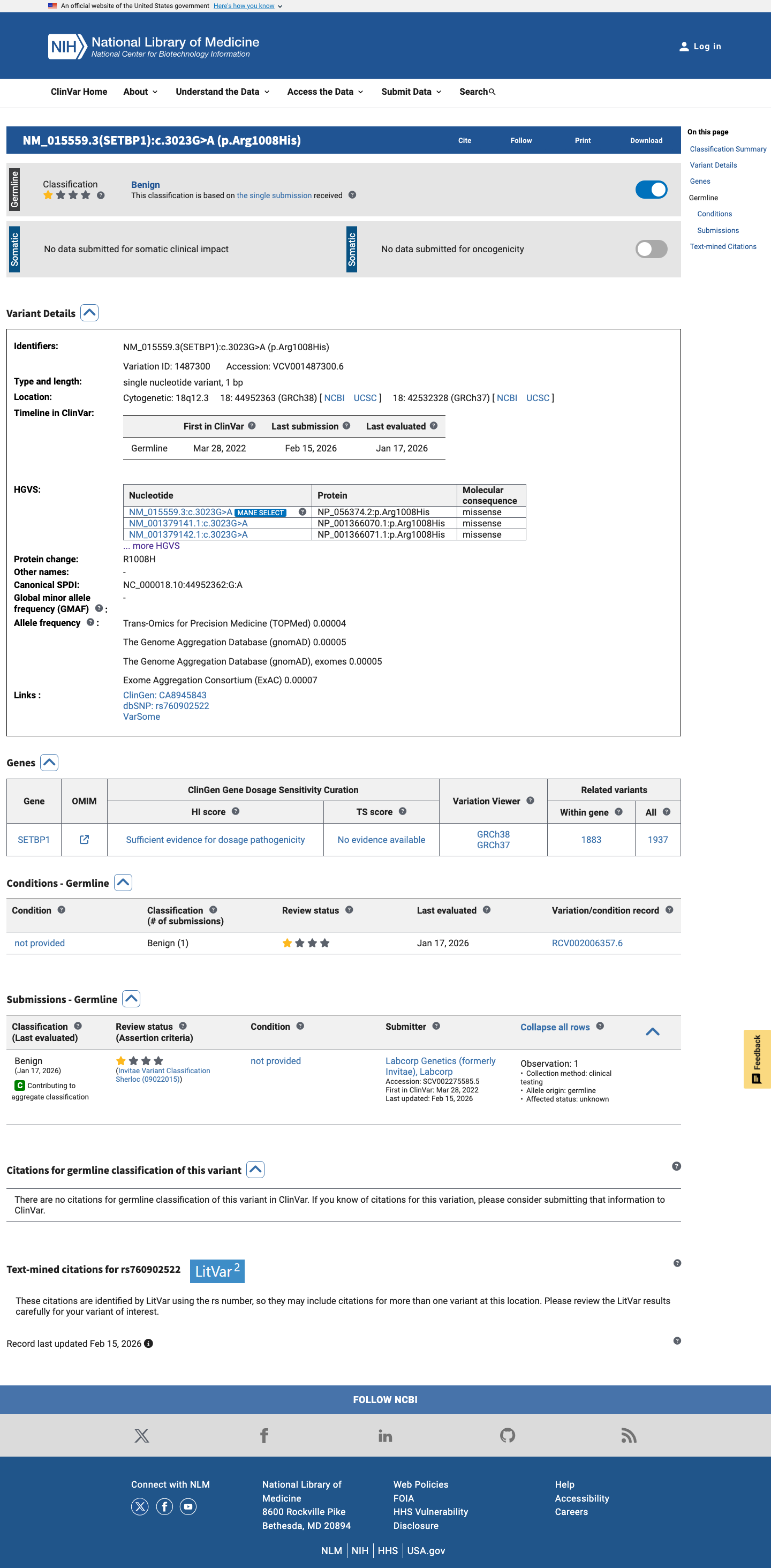
The SETBP1 c.3023G>A (p.Arg1008His) variant has been reported in ClinVar as Benign by a single clinical laboratory submitter.
clinvar ↗2
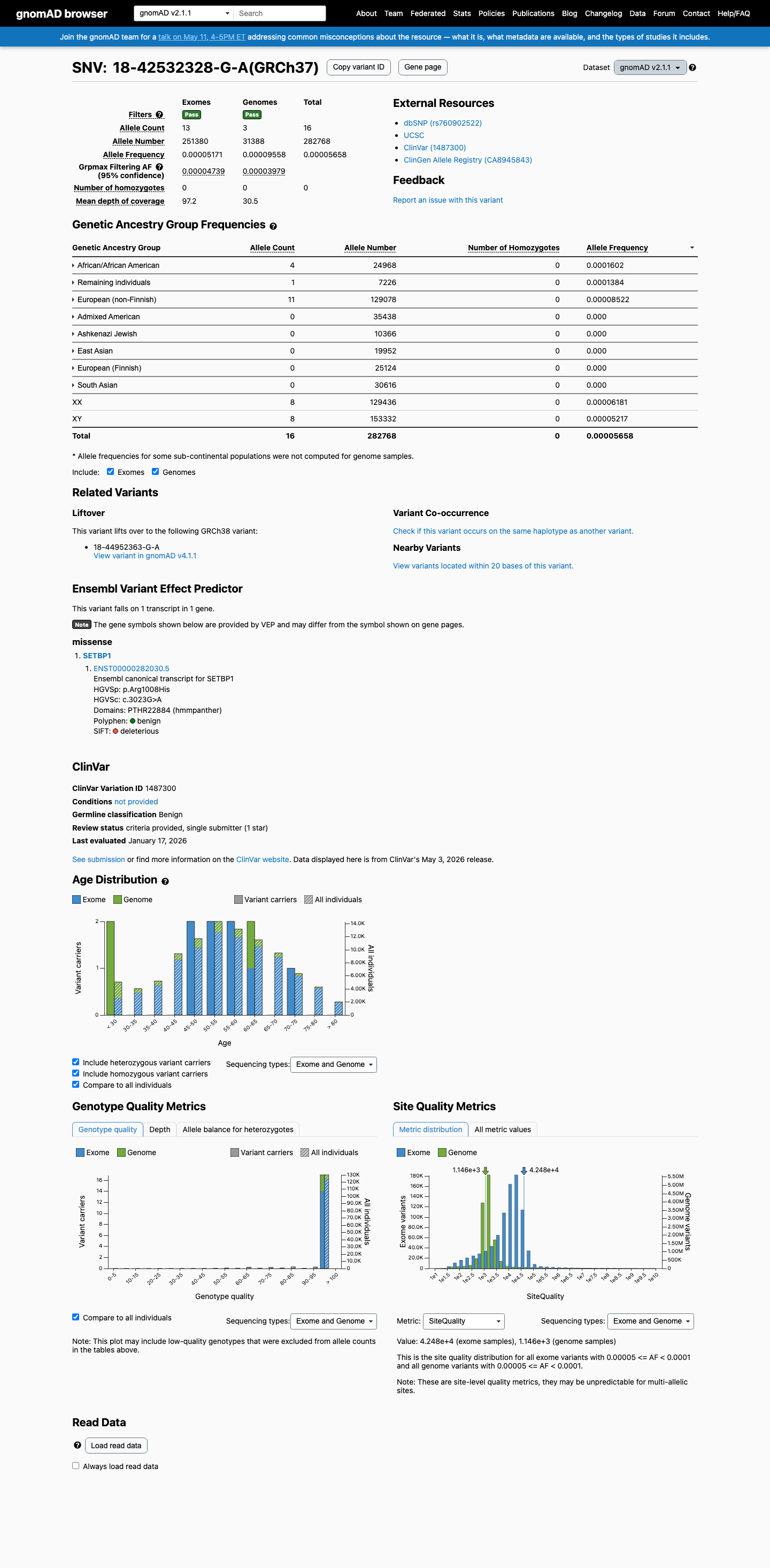
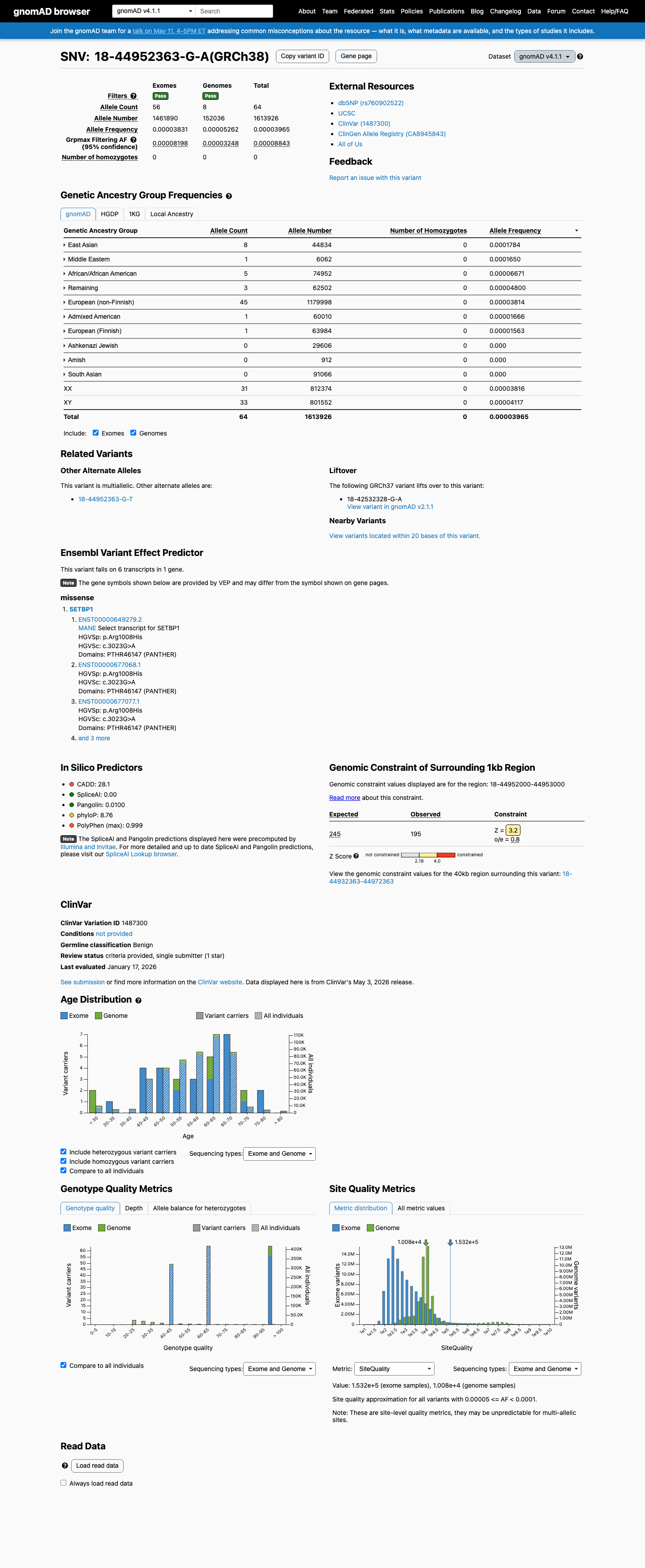
This variant is present at low frequency in population databases, with gnomAD v2.1 AF 0.0000566 (16/282768 alleles) and gnomAD v4.1 AF 0.0000397 (64/1613926 alleles), both below the 0.1% PM2 threshold and below BS1 and BA1 benign thresholds.
gnomad_v2 ↗ gnomad_v4 ↗3
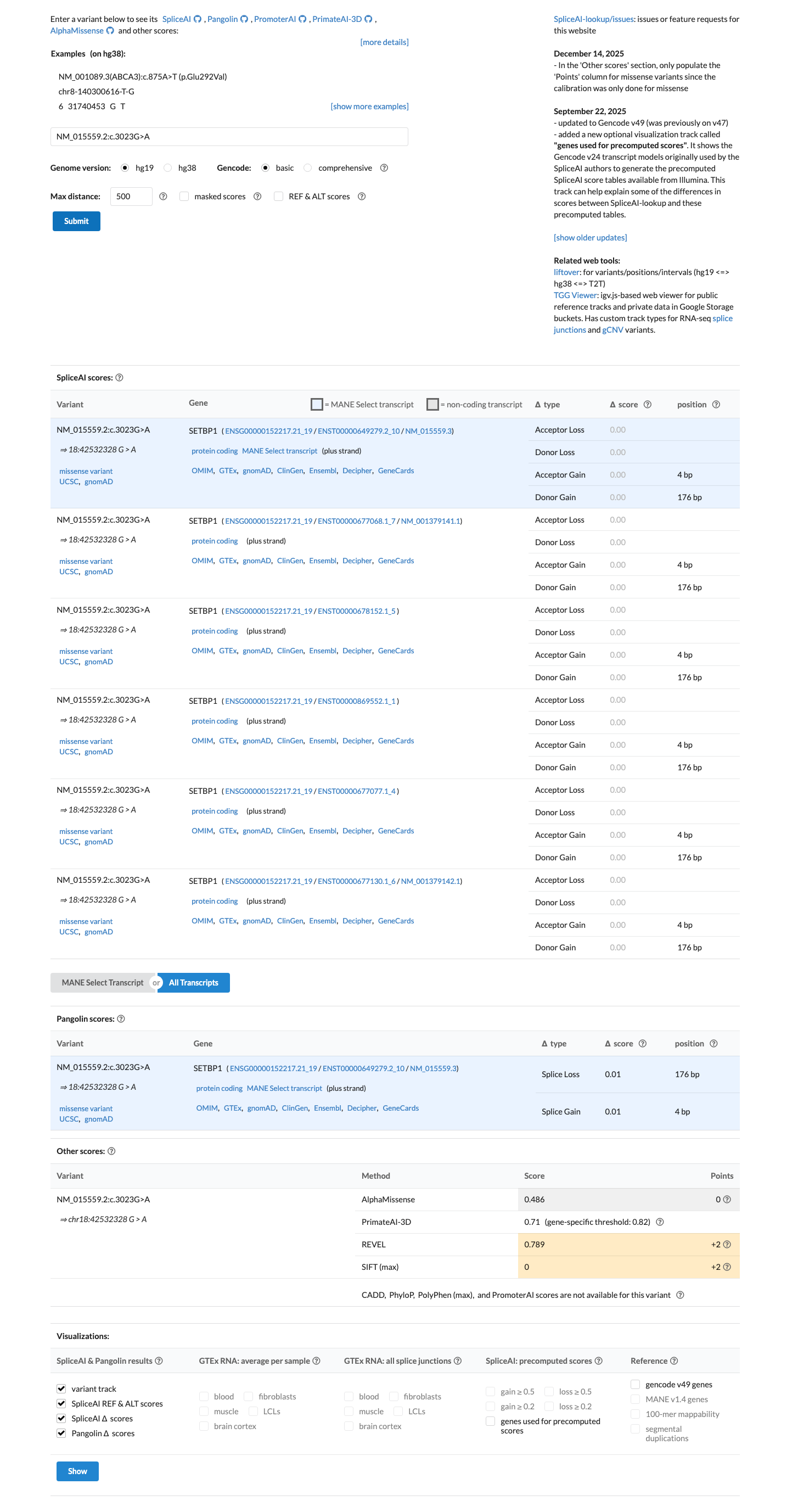
Computational evidence supports a deleterious missense effect, with REVEL 0.789 and BayesDel 0.412559, while SpliceAI predicts no significant splice impact with a max delta score of 0.00.
spliceai ↗