Classification rationale
1
2
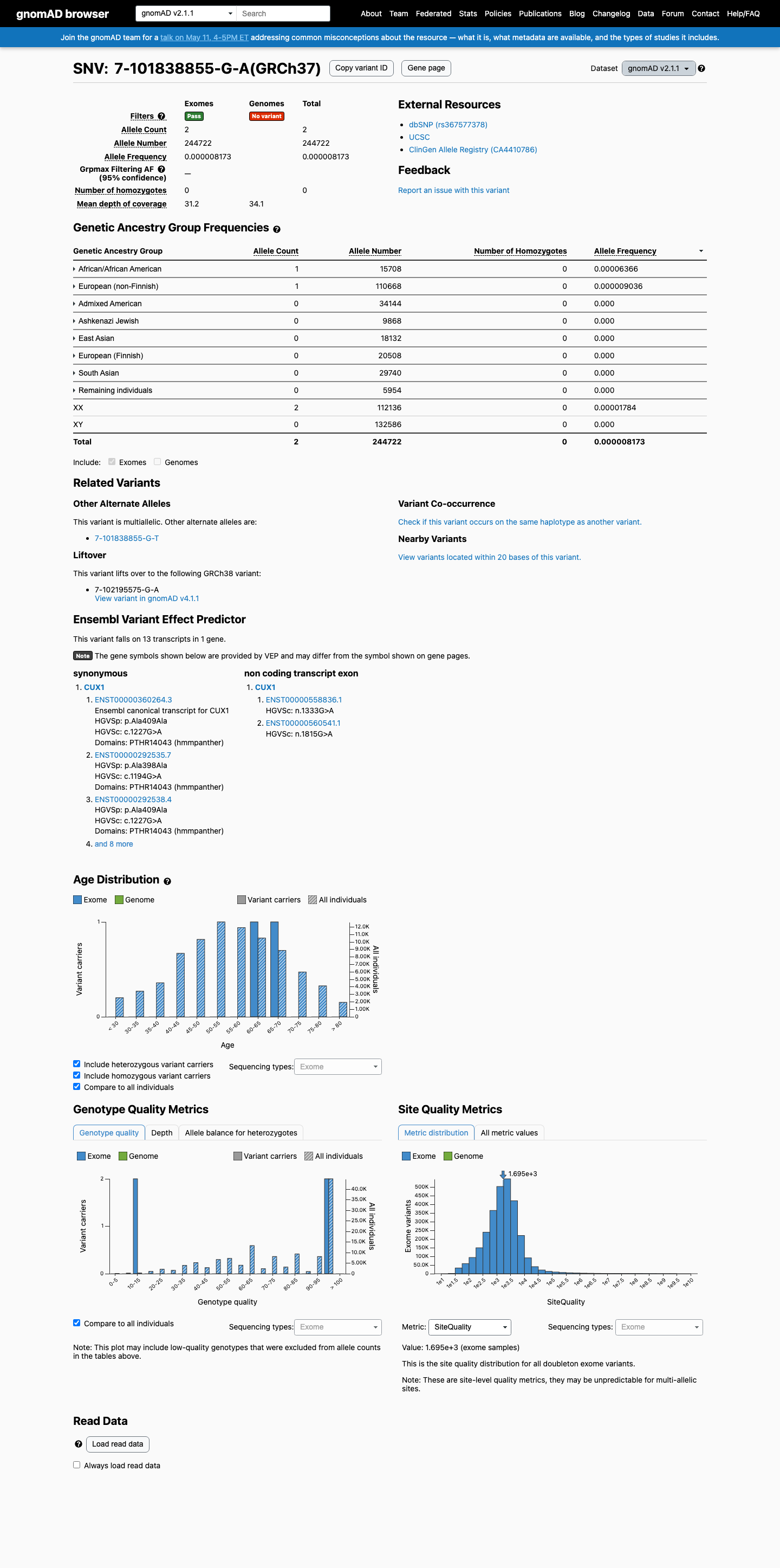
This variant is very rare in population databases, with allele frequencies of 8.17e-06 (2/244722; 0.00082%) in gnomAD v2.1 and 8.07e-06 (13/1611678; 0.00081%) in gnomAD v4.1.
gnomad_v2 ↗ gnomad_v4 ↗3
CUX1 loss of function is an established germline disease mechanism, but this synonymous variant is not a generic PVS1-eligible loss-of-function variant.
pvs1_generic_framework ↗4
Computational splice prediction does not support a damaging effect, as SpliceAI predicts no significant splice impact with a maximum delta score of 0.00.
spliceai ↗