Classification rationale
1
The DNMT3A c.1312dup (p.Asp438GlyfsTer7; p.D438Gfs*7) variant has not been reported in ClinVar.
clinvar ↗2
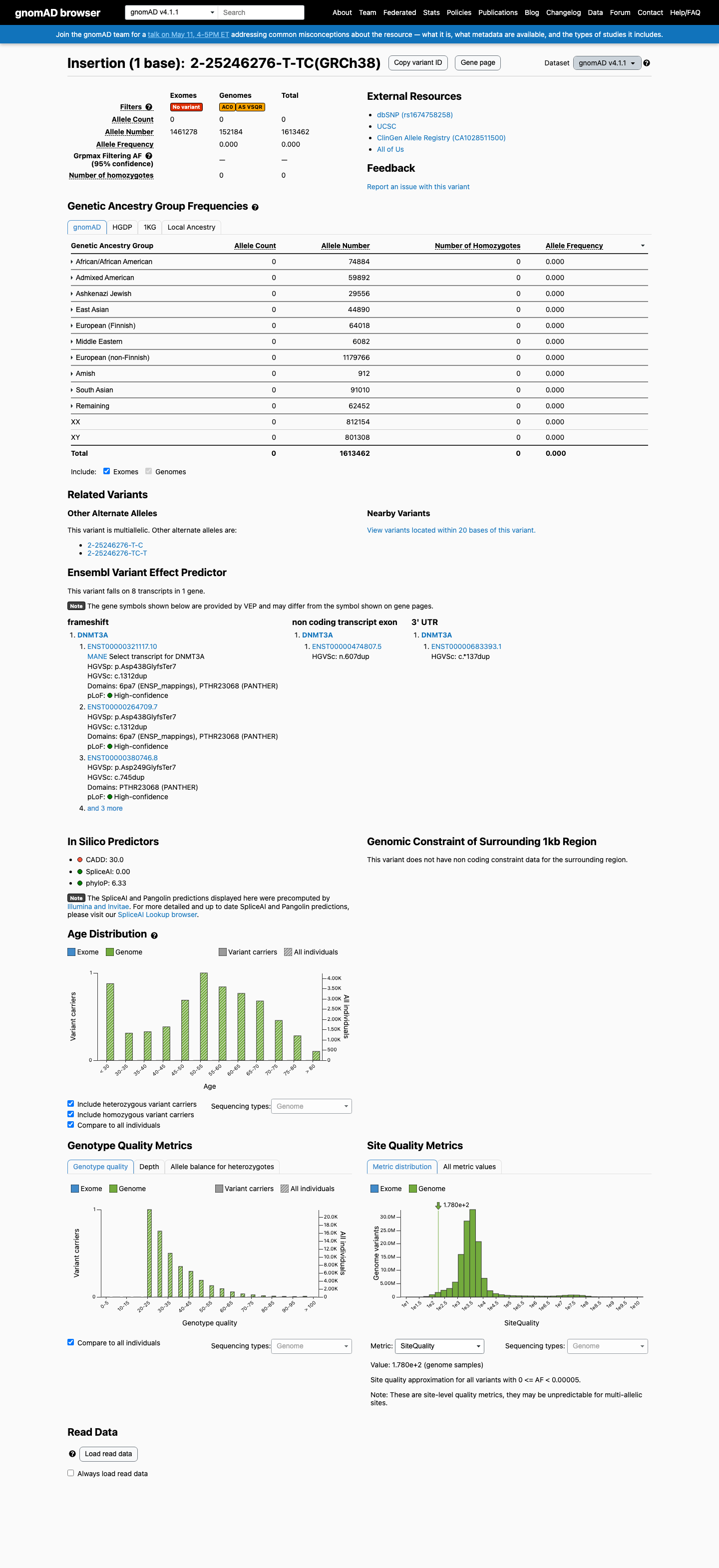
This variant is absent from gnomAD v2.1 and absent from gnomAD v4.1, with 0/1613462 alleles observed overall in gnomAD v4.1, which is below the 0.1% non-VCEP PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
Published DNMT3A germline disease literature supports loss of function as a disease mechanism, and this variant is predicted to introduce a premature stop codon through a frameshift, consistent with a truncating effect.
PMID:24614070 ↗ pvs1_generic_framework ↗4
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.06, supporting that the main predicted consequence is the frameshift truncation rather than splice disruption.
spliceai ↗