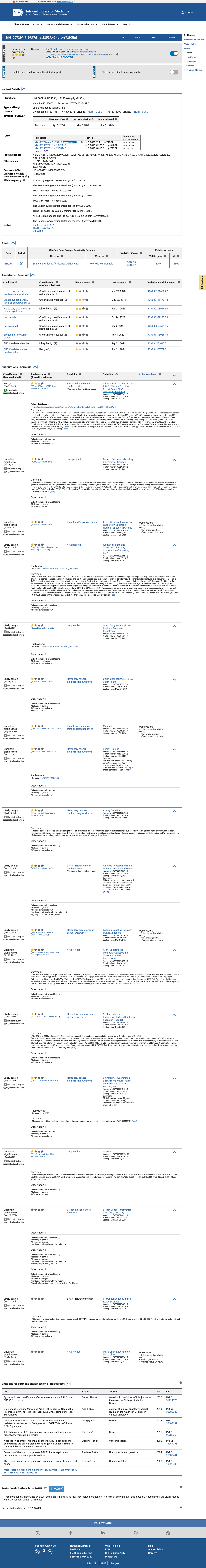
The BRCA1 NM_007294.3:c.2155A>G (NP_009225.1:p.(Lys719Glu), NP_009225.1:p.(K719E)) variant has not been observed in COSMIC and has been reported in ClinVar with a current expert-panel Benign classification, although older laboratory submissions include uncertain significance and likely benign assertions.
clinvar ↗This variant is present in population databases, including gnomAD v2.1 and v4.1, with grpmax FAF values of 0.00062195 and 0.00068503, respectively, which are above the ENIGMA BS1 threshold of 0.0001 but below the BA1 threshold of 0.001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Multifactorial clinical-history evidence is in the benign direction, with a BRCA1 clinical-history likelihood ratio of 0.0137 from 10 probands, meeting BP5_Strong and arguing against pathogenicity.
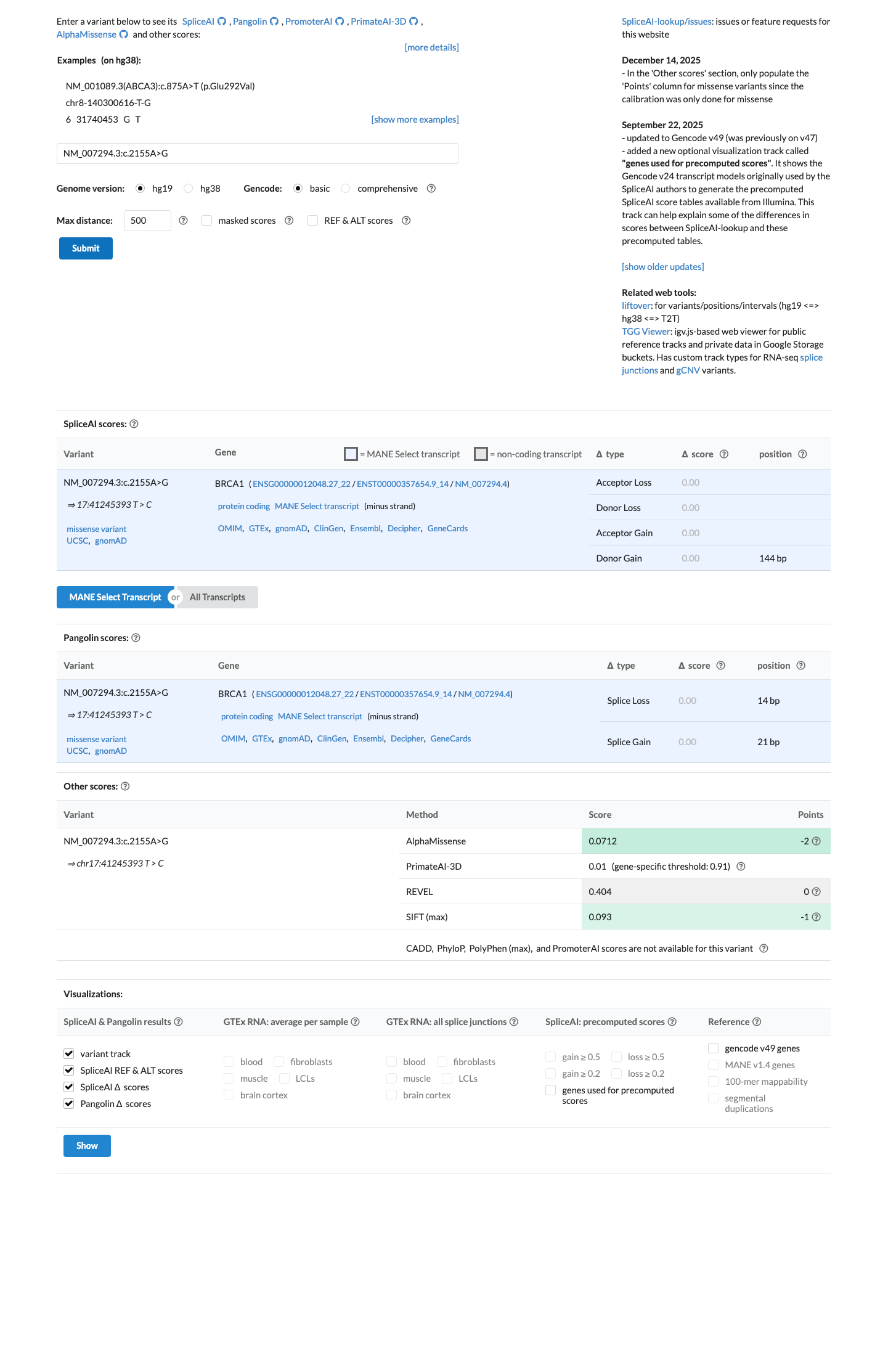
PMID:31853058 ↗ cspec ↗In silico evidence does not support a damaging effect in the ENIGMA BRCA1 framework: SpliceAI predicts no splice impact (max delta score 0.00), BayesDel is -0.216062, and the missense change lies outside the BRCA1 domains used for PP3/BP4, supporting BP1_Strong rather than PP3.
spliceai ↗ cspec ↗