The PTPN11 c.53A>G (p.Asn18Ser) variant has not been observed in COSMIC and has been reported in ClinVar as Benign by the ClinGen RASopathy Variant Curation Expert Panel, with additional benign and likely benign clinical laboratory submissions.
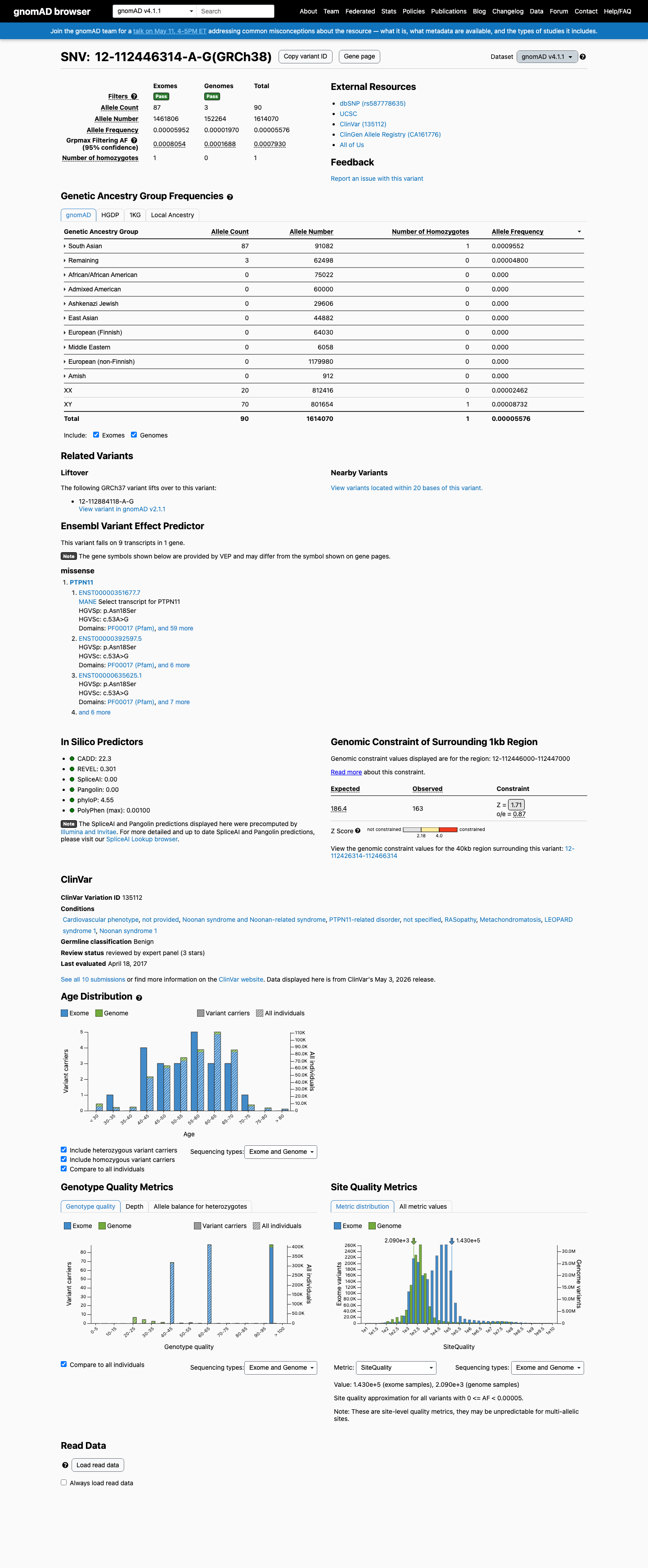
clinvar ↗This variant is present in gnomAD v2.1 and v4.1, with the highest observed South Asian frequency reaching 0.09552% in v4.1 and grpmax filtering allele frequencies of 0.05125% in v2.1 and 0.07930% in v4.1, which are above the PTPN11 RASopathy BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗No approved variant-specific functional study for p.(Asn18Ser) was identified in the RASopathy VCEP approved functional studies resource.
Computational evidence does not meet the PTPN11 missense thresholds for either PP3 or BP4: REVEL is 0.301, which is below the PP3 cutoff of 0.7 and just above the BP4 cutoff of 0.3, SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, and BayesDel is -0.292277.
cspec ↗ spliceai ↗