Classification rationale
1
The PTEN c.956del (p.Thr319IlefsTer2; p.T319Ifs*2) variant has been reported in ClinVar as Pathogenic by one clinical laboratory.
clinvar ↗2
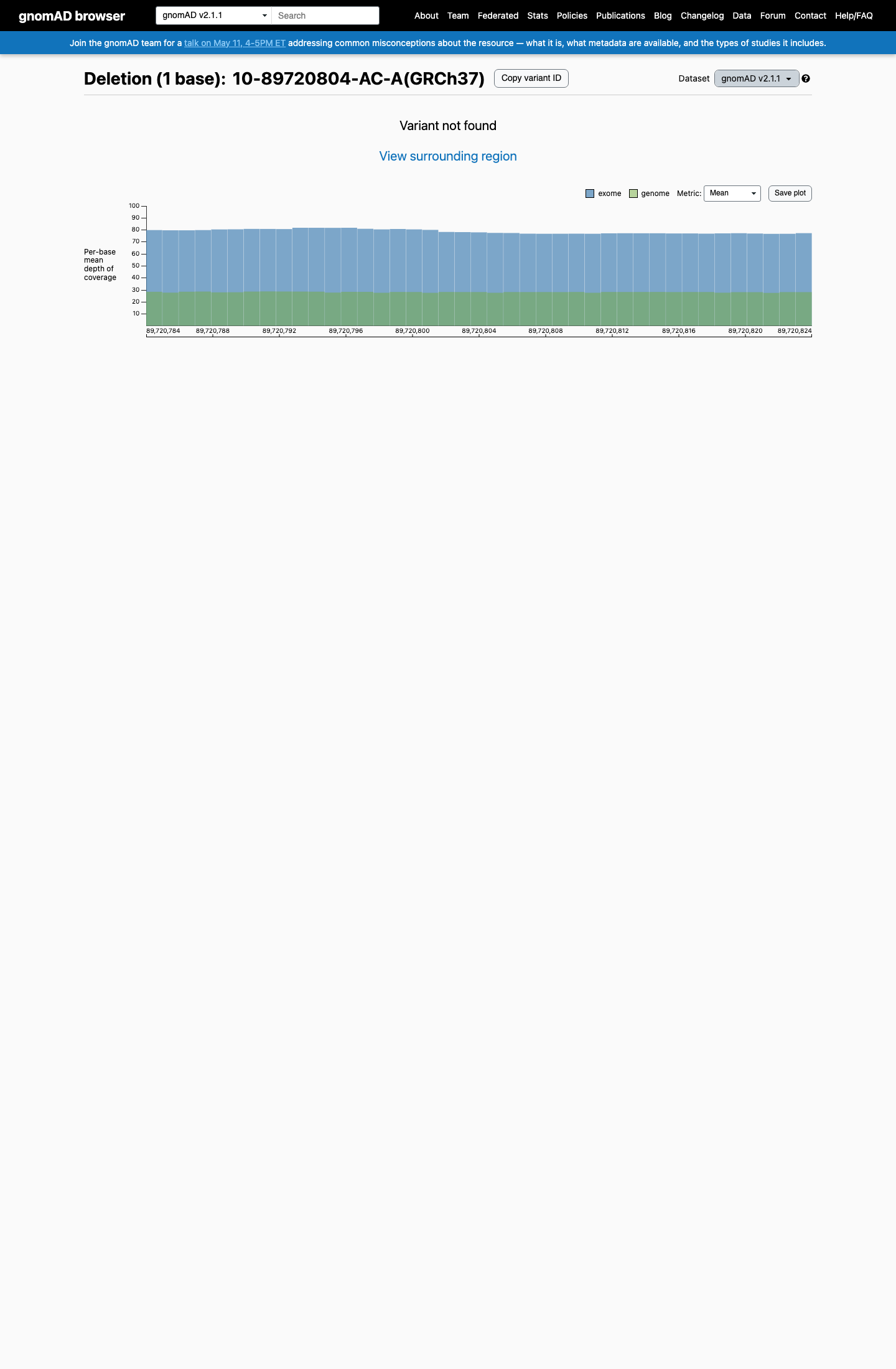
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population databases and meeting the PTEN PM2_Supporting threshold of less than 0.00001 (0.001%).
cspec ↗ gnomad_v2 ↗ gnomad_v4 ↗3
This frameshift is predicted to truncate PTEN at codon 320 in exon 8, and the PTEN-specific PVS1 decision tree assigns PVS1 to truncating variants at or 5' to p.D375 (c.1121) in transcript NM_000314.8.
cspec ↗4
SpliceAI predicts no significant splice effect for this variant (max delta score 0.02), and REVEL and BayesDel are not applicable because this is not a single-nucleotide missense variant.
spliceai ↗ cspec ↗