Classification rationale
1
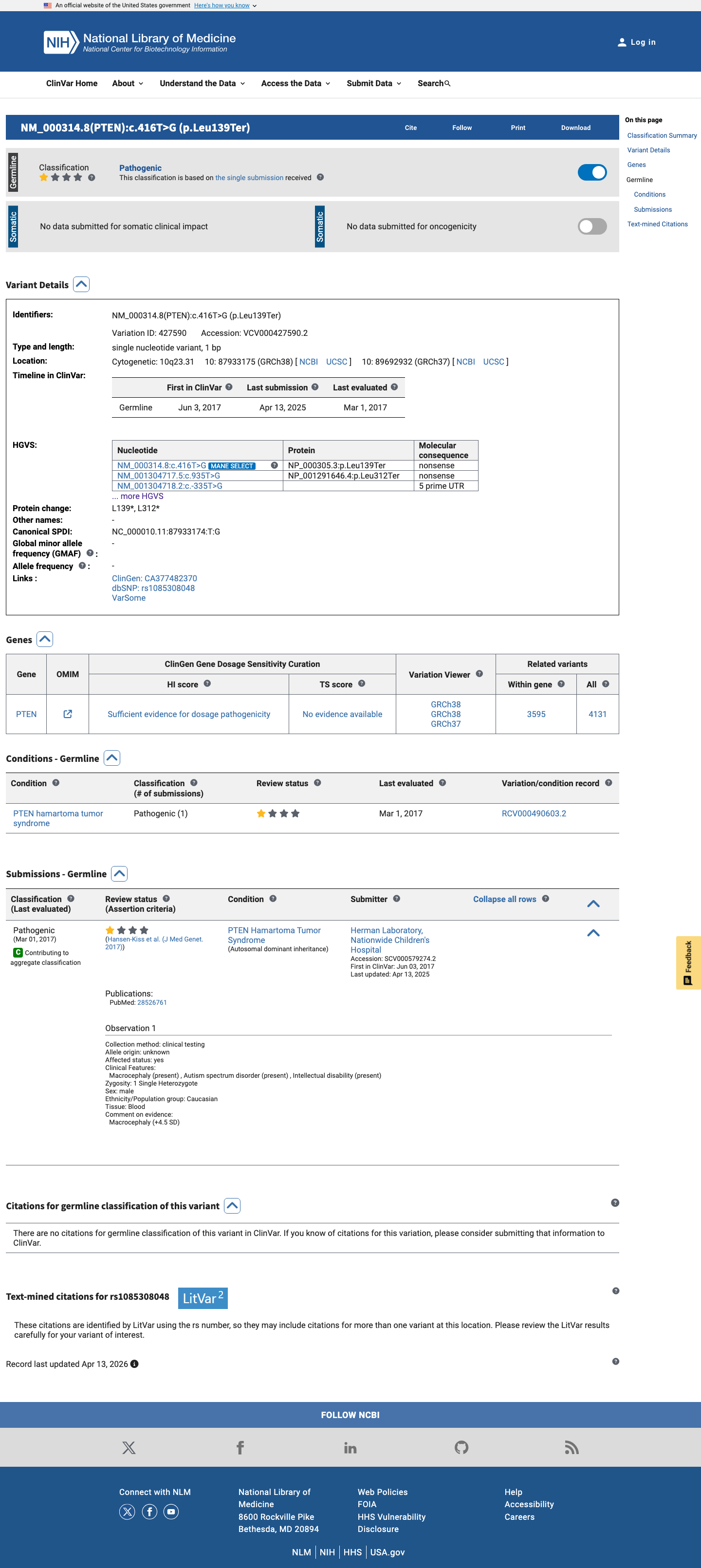
The PTEN c.416T>G (p.(Leu139Ter)) variant has been observed in somatic cancers (COSMIC COSV64297098, n=6) and has been reported in ClinVar as pathogenic by a single submitter.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which meets the PTEN VCEP PM2_Supporting rarity threshold of less than 0.001% allele frequency.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
This variant introduces a premature stop codon at Leu139, and under the PTEN-specific PVS1 decision tree truncating variants at or 5' to p.D375 in transcript NM_000314.8 are assigned PVS1; SpliceAI predicts no significant splice effect with a maximum delta score of 0.01.
cspec ↗ spliceai ↗