Classification rationale
1
The ABL1 c.908-16G>A (p.?) variant has been reported in ClinVar as likely benign with single-submitter review status.
clinvar ↗2
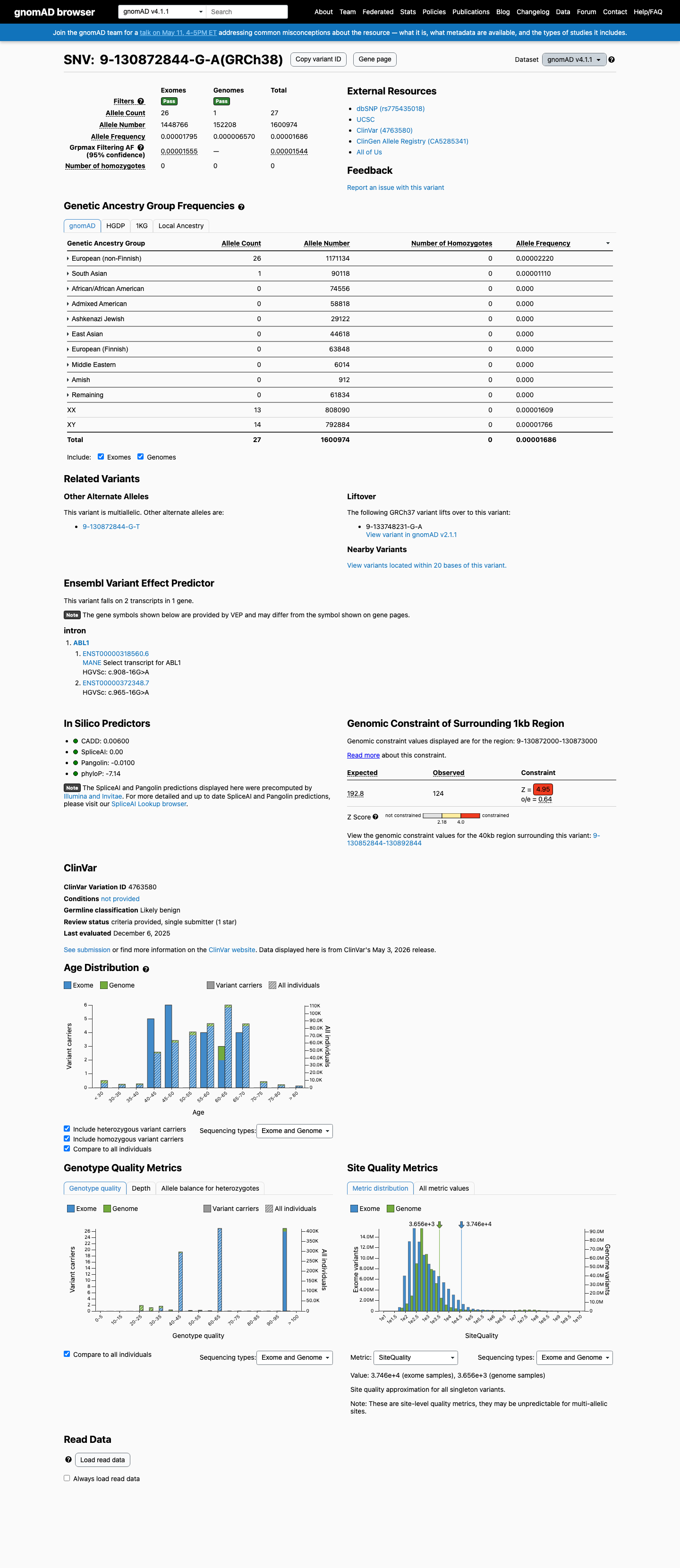
This variant is present at very low frequency in population databases, with allele frequencies of 0.00082% in gnomAD v2.1 and 0.00169% in gnomAD v4.1, both below the 0.1% PM2 threshold used in this workflow.
gnomad_v2 ↗ gnomad_v4 ↗3
SpliceAI predicts no significant splice effect for this intronic change, with a maximum delta score of 0.01, which supports a benign computational interpretation.
spliceai ↗