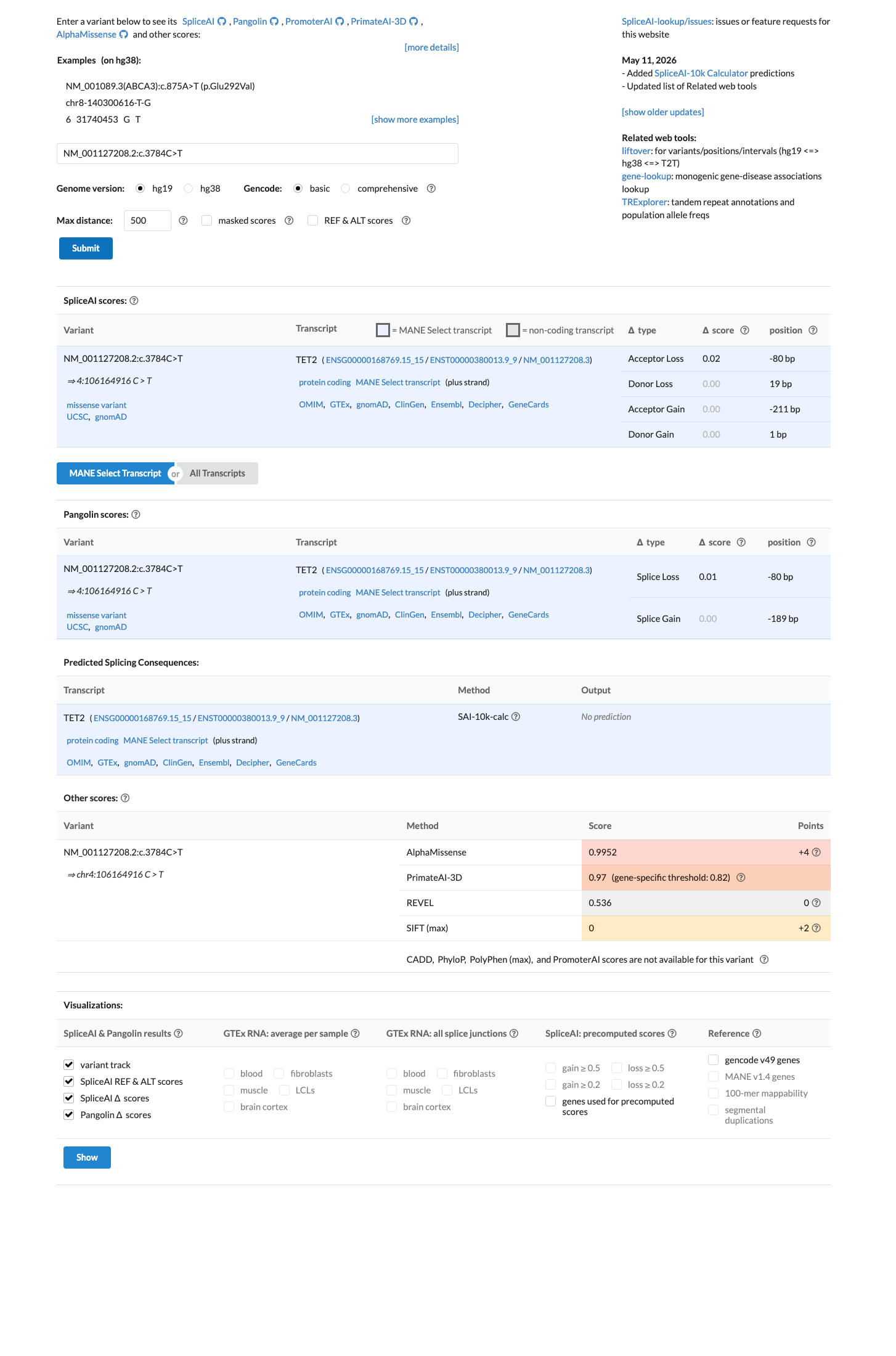
The TET2 c.3784C>T (p.Arg1262Trp; p.R1262W) variant has not been reported in ClinVar.
clinvar ↗This variant is present at low frequency in gnomAD v2.1 and v4.1, with the highest observed frequency 0.00648% (1/15,424 alleles) in European non-Finnish individuals in gnomAD v2.1, which is below the 0.1% PM2 threshold and below the BS1 (>0.3%) and BA1 (>1%) thresholds.
gnomad_v2 ↗ gnomad_v4 ↗No variant-specific functional study establishing either a damaging effect or normal function was identified; the available published TET2 structural study provides protein-domain context but does not assay p.Arg1262Trp directly.
oncokb ↗ PMID:24315485 ↗Computational evidence is mixed overall: SpliceAI predicts no significant splice impact (max delta score 0.02), while REVEL is 0.536 and BayesDel is 0.118577, which does not provide concordant support for either PP3 or BP4.
spliceai ↗