Classification rationale
1
The CTNNB1 c.1648C>T (p.Arg550Cys; p.R550C) variant has been reported in ClinVar as a variant of uncertain significance by a single submitter.
clinvar ↗2
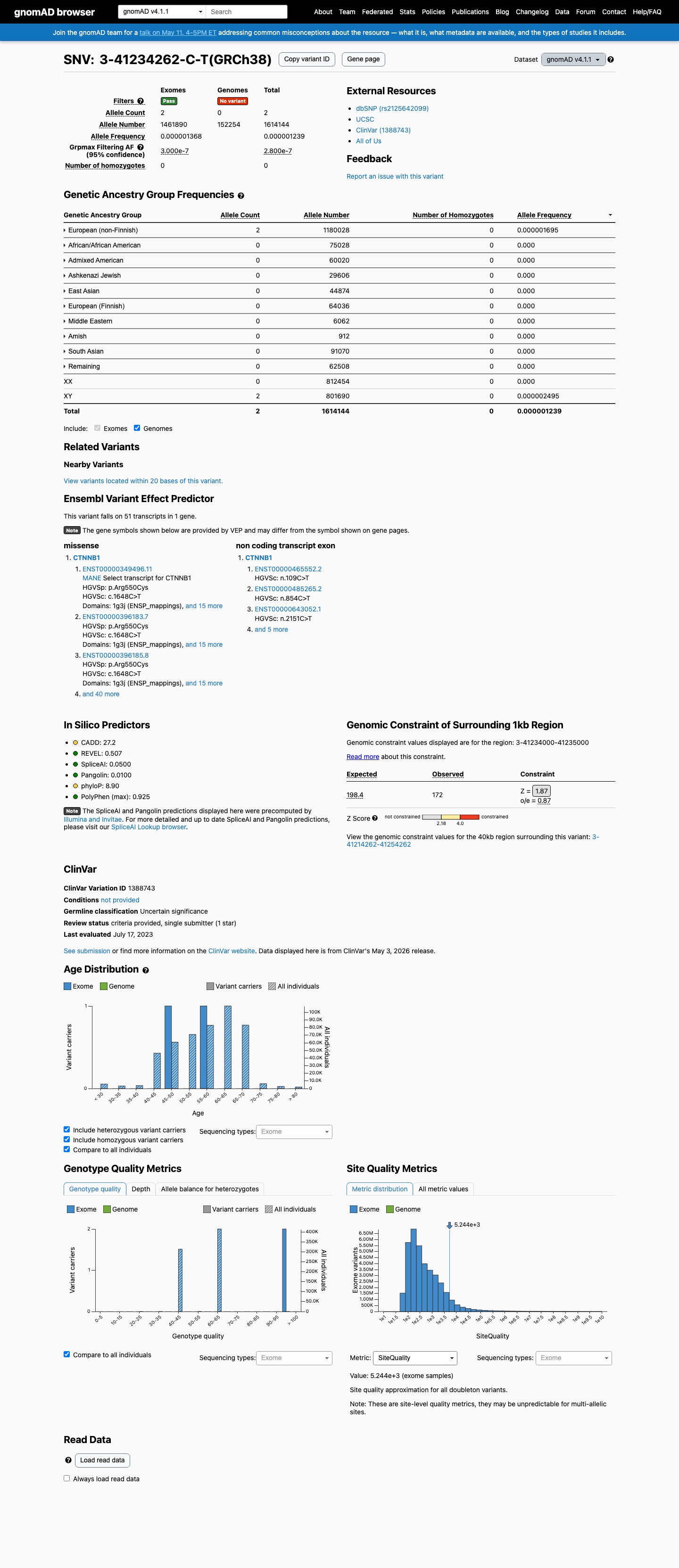
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in the general population and meeting PM2 under the generic ACMG framework.
gnomad_v2 ↗ gnomad_v4 ↗3
Available computational evidence is mixed: SpliceAI predicts no significant splice effect with a maximum delta score of 0.05, while REVEL is 0.507 and BayesDel is 0.164625, so the in silico data support neither PP3 nor BP4.
spliceai ↗