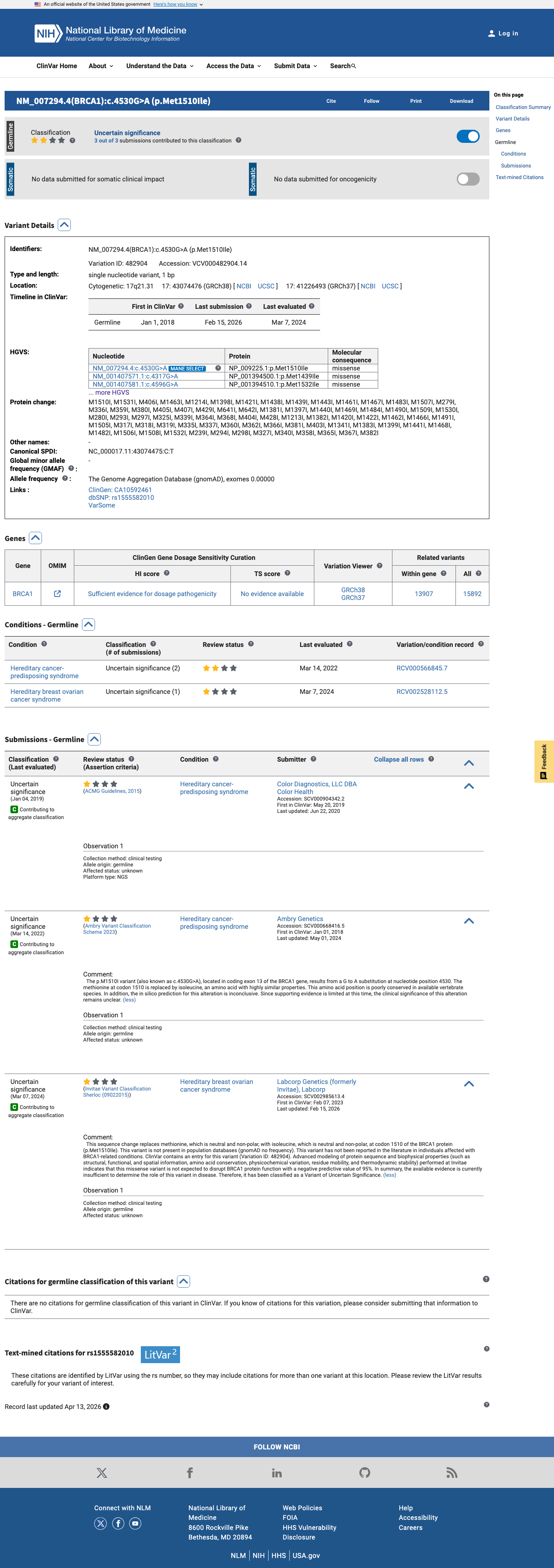
The BRCA1 c.4530G>A (p.Met1510Ile) variant has been reported in ClinVar as a variant of uncertain significance with three clinical laboratory submissions.
clinvar ↗This variant is absent from gnomAD v2.1 and present in gnomAD v4.1 at 1/1,614,132 alleles (AF 6.20e-07), indicating that it is very rare but not absent from population databases.
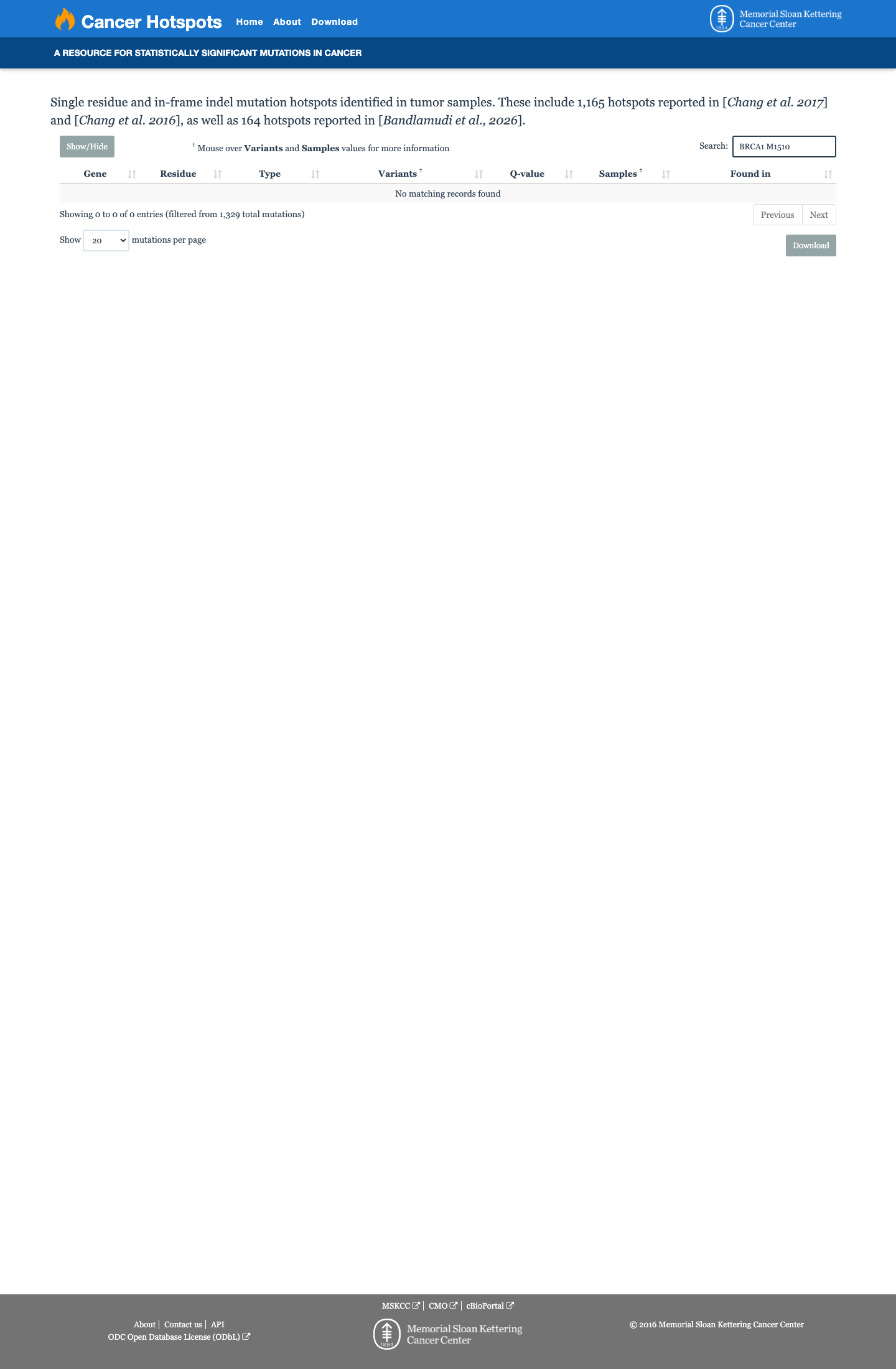
gnomad_v2 ↗ gnomad_v4 ↗No variant-specific calibrated functional assay result for c.4530G>A (p.Met1510Ile) was identified in the reviewed ENIGMA BRCA1 functional tables, so PS3 and BS3 were not applied.
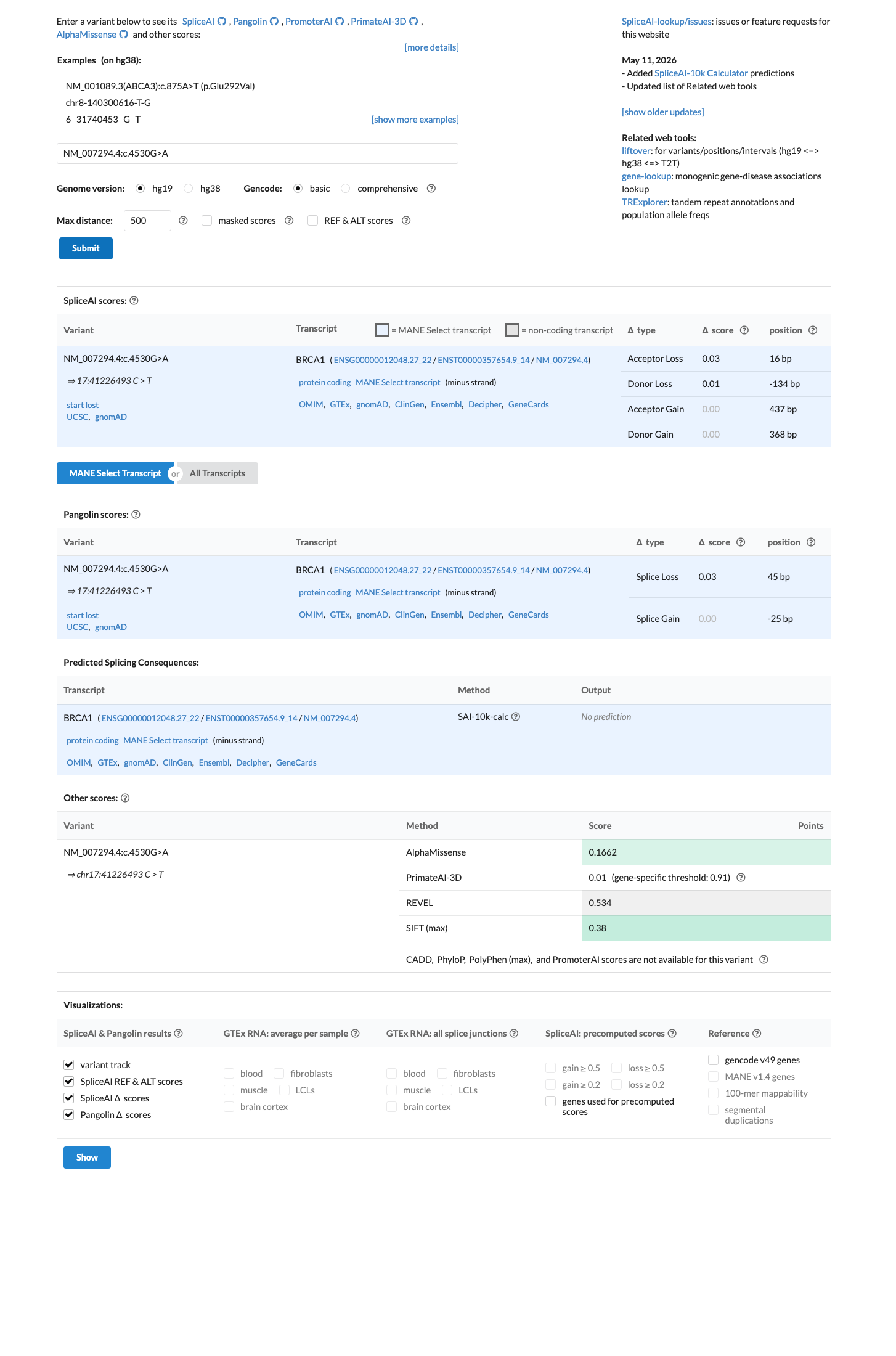
cspec ↗Computational evidence supports a benign direction under the BRCA1 ENIGMA rules because this missense change lies outside the BRCA1 clinically important domains and SpliceAI predicts no significant splice effect (max delta score 0.03), meeting BP1_Strong; BayesDel no-AF is -0.119 and does not meet the PP3 threshold, and REVEL is 0.534.
cspec ↗ spliceai ↗The BRCA1 clinical-history likelihood ratio for this variant is 1.90 in one proband, which falls between the ENIGMA BP5 threshold of 0.48 and the PP4 threshold of 2.08, so neither PP4 nor BP5 was applied.
PMID:31853058 ↗ cspec ↗