Classification rationale
1
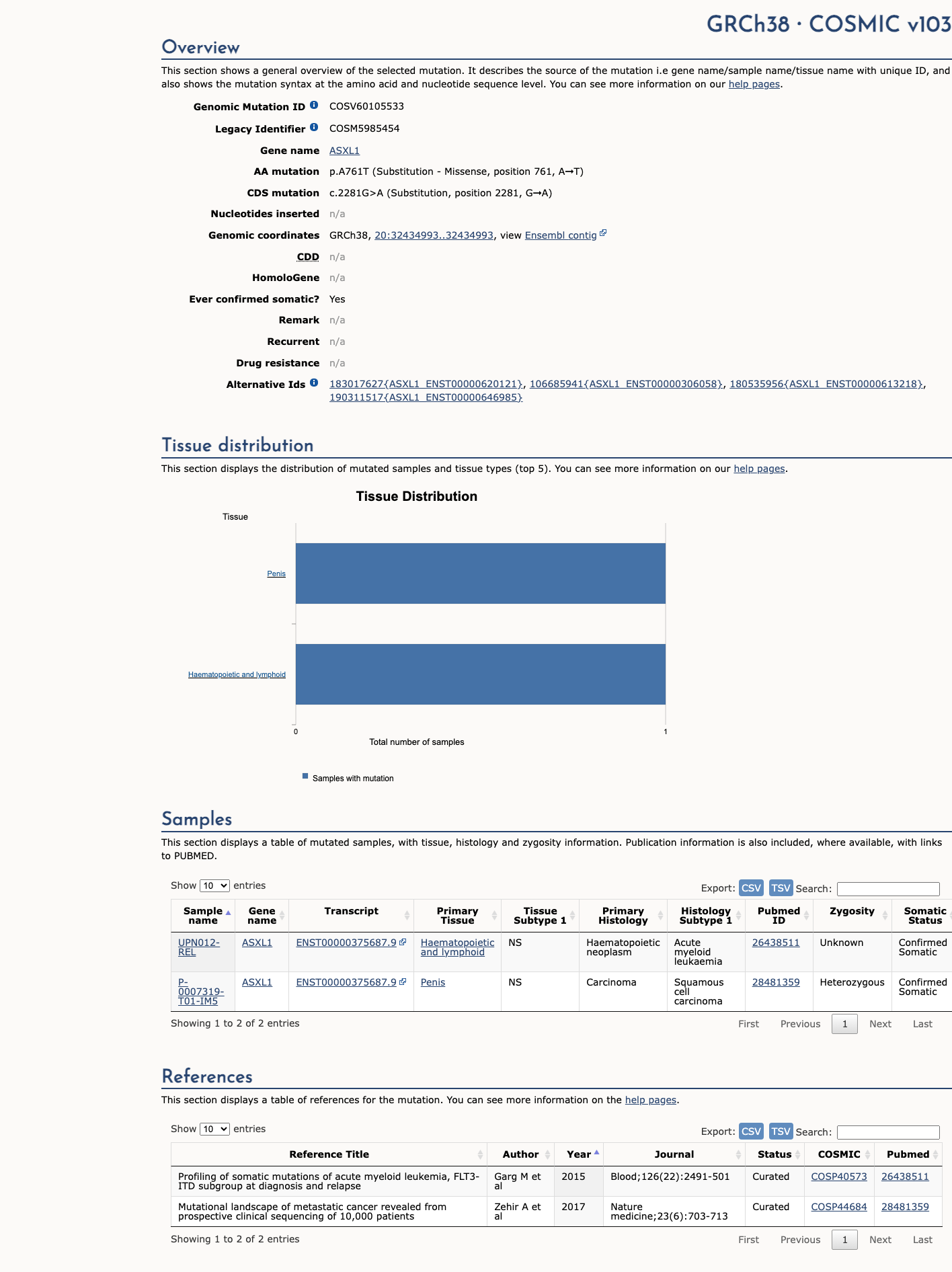
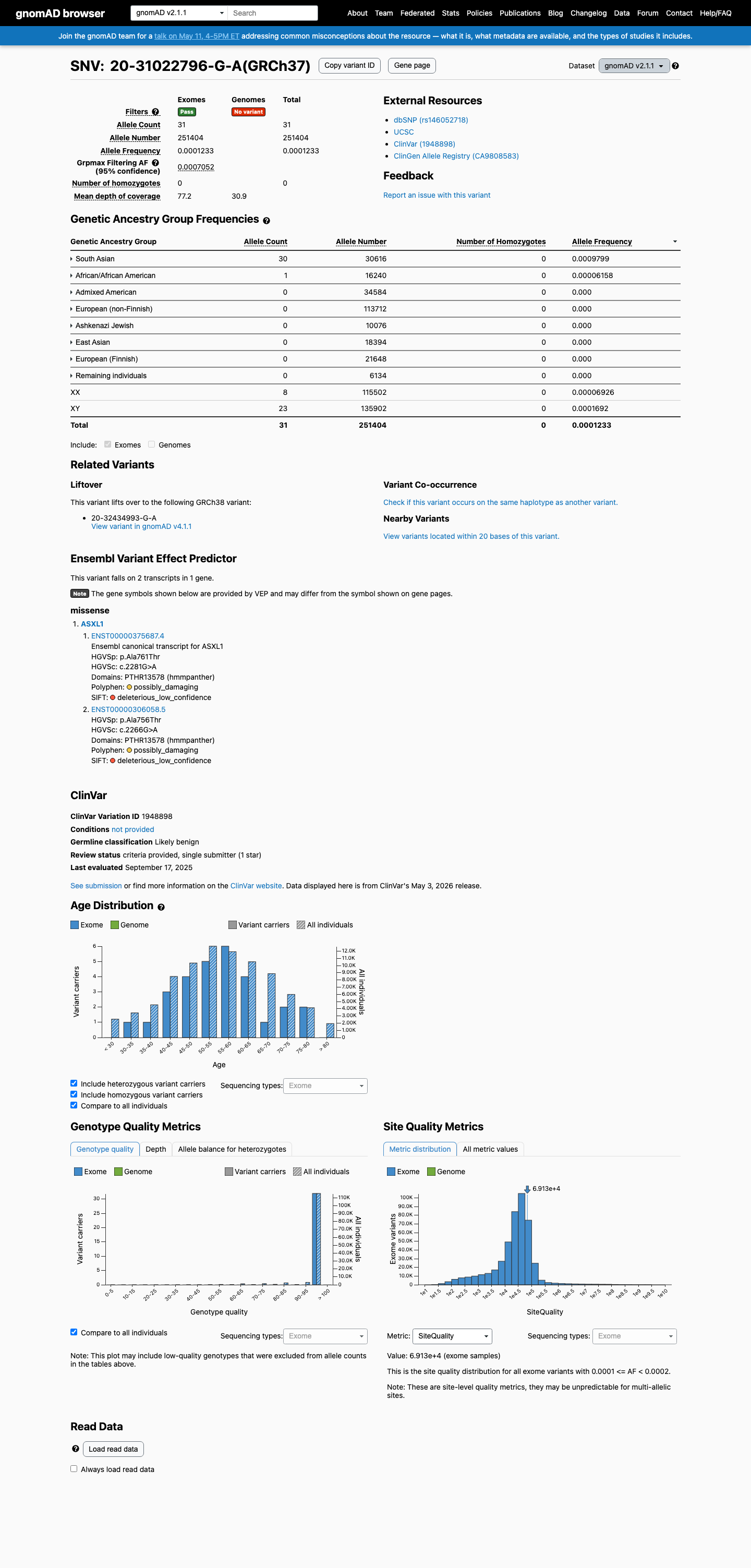
The ASXL1 c.2281G>A (p.Ala761Thr) variant has been reported in ClinVar as Likely benign by a single submitter.
clinvar ↗2
This variant is present in gnomAD, with a highest observed South Asian allele frequency of 0.09799% (30/30,616) in v2.1 and 0.09442% (86/91,078) in v4.1, which is below the default BS1 threshold of 0.3% and BA1 threshold of 1.0%.
gnomad_v2 ↗ gnomad_v4 ↗3
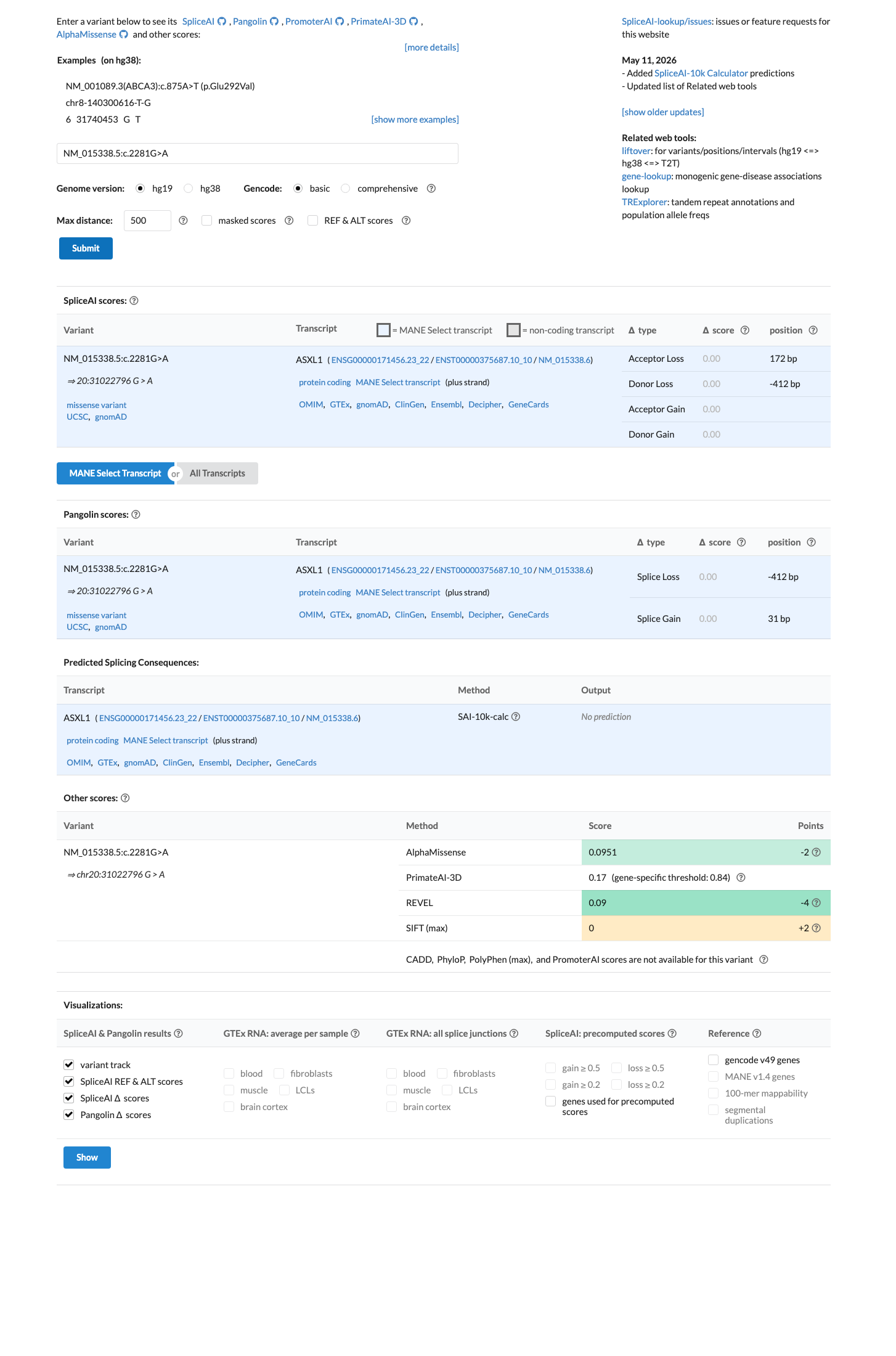
Computational evidence does not support a damaging effect: REVEL is 0.09, BayesDel is -0.340274, and SpliceAI predicts no significant splice impact with a maximum delta score of 0.00.
spliceai ↗