Classification rationale
1
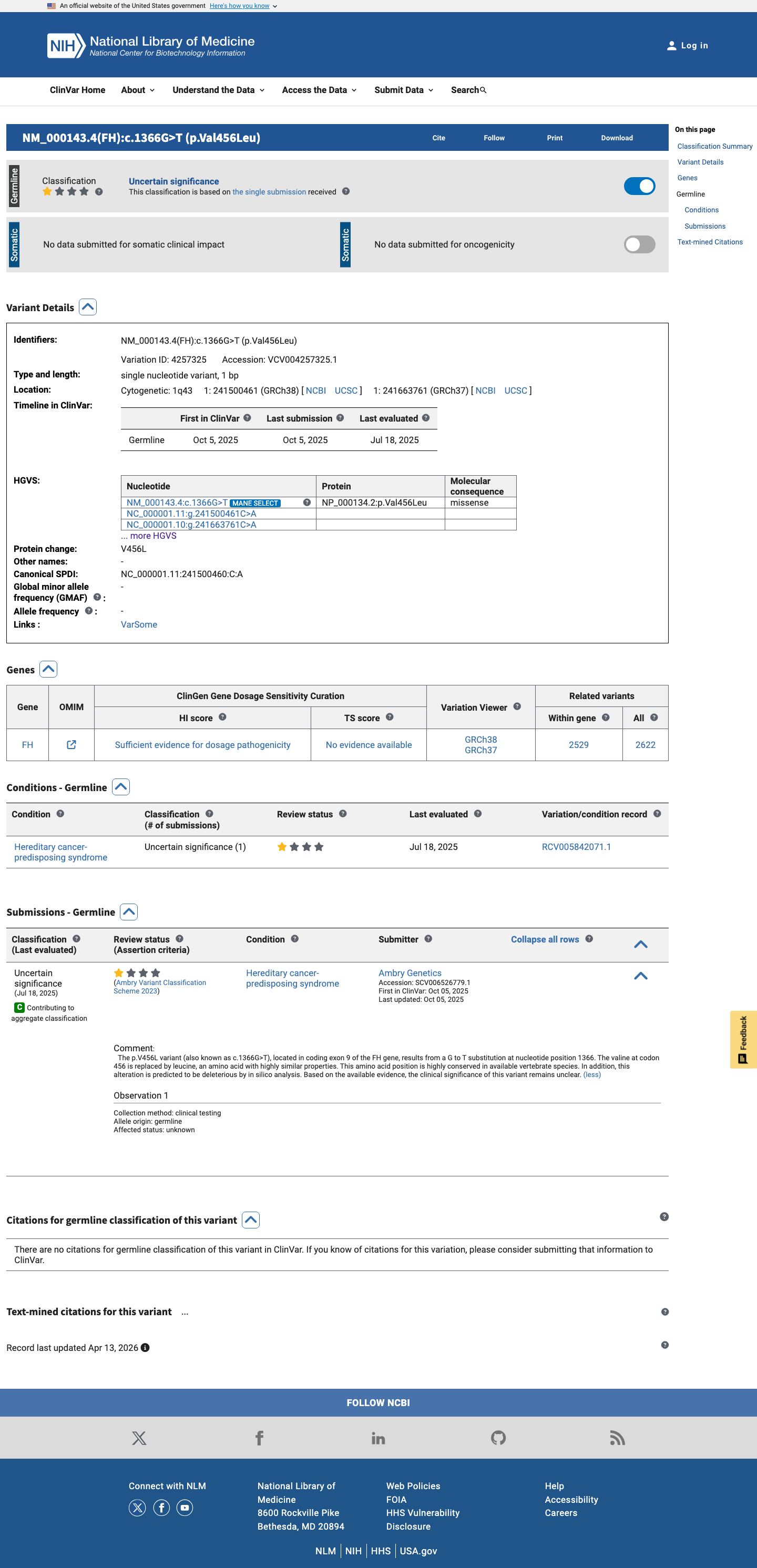
The FH c.1366G>T (p.Val456Leu) variant has been reported in ClinVar as a variant of uncertain significance by one clinical laboratory.
clinvar ↗2
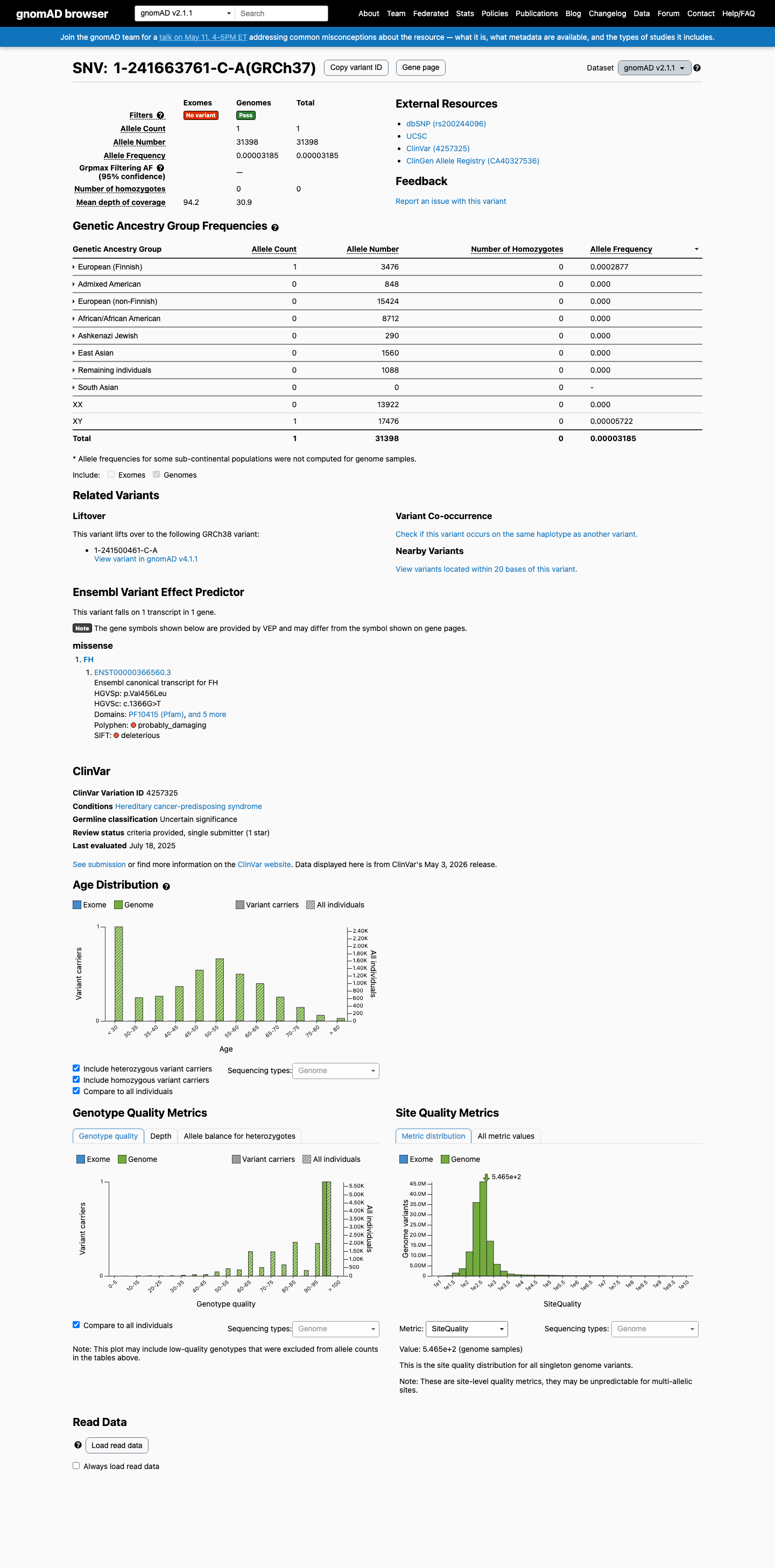
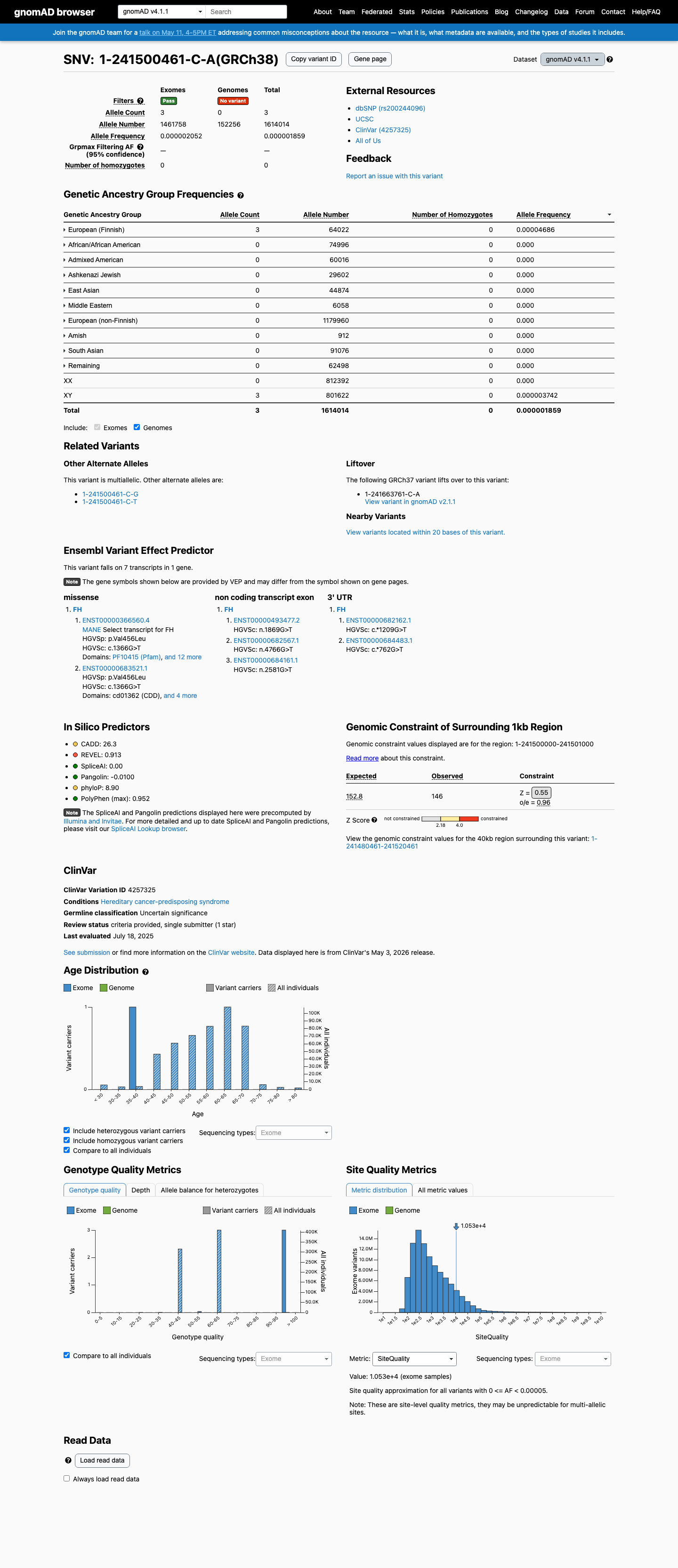
This variant is rare in population databases, with gnomAD v2.1 AF 0.00318% (1/31,398 alleles) and gnomAD v4.1 AF 0.00019% (3/1,614,014 alleles), which are both below the 0.1% PM2 threshold and well below the BS1 and BA1 frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗3
Computational evidence supports a deleterious missense effect, with REVEL 0.913 and BayesDel 0.286106, while SpliceAI predicts no significant splice impact for this variant (max delta score 0.00).
spliceai ↗