Classification rationale
1
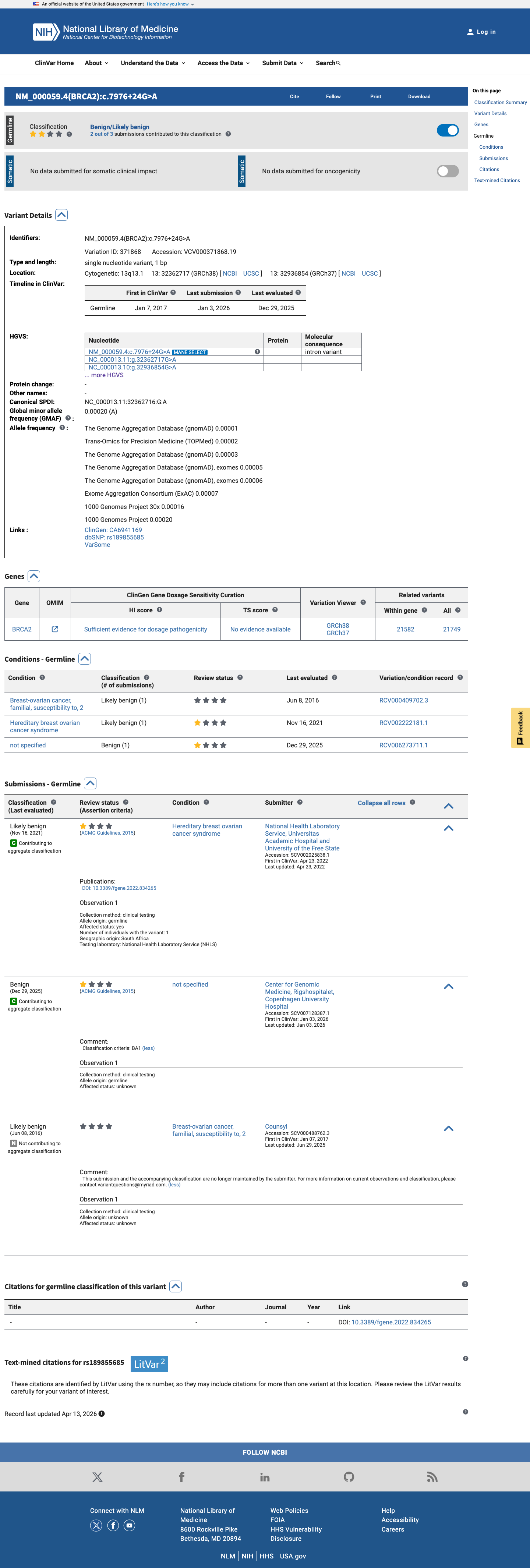
The BRCA2 NM_000059.4:c.7976+24G>A (NP_000050.3:p.?) variant has been reported in ClinVar (Variation ID 371868), although submission-level classification details were not available in the reviewed record.
clinvar ↗2
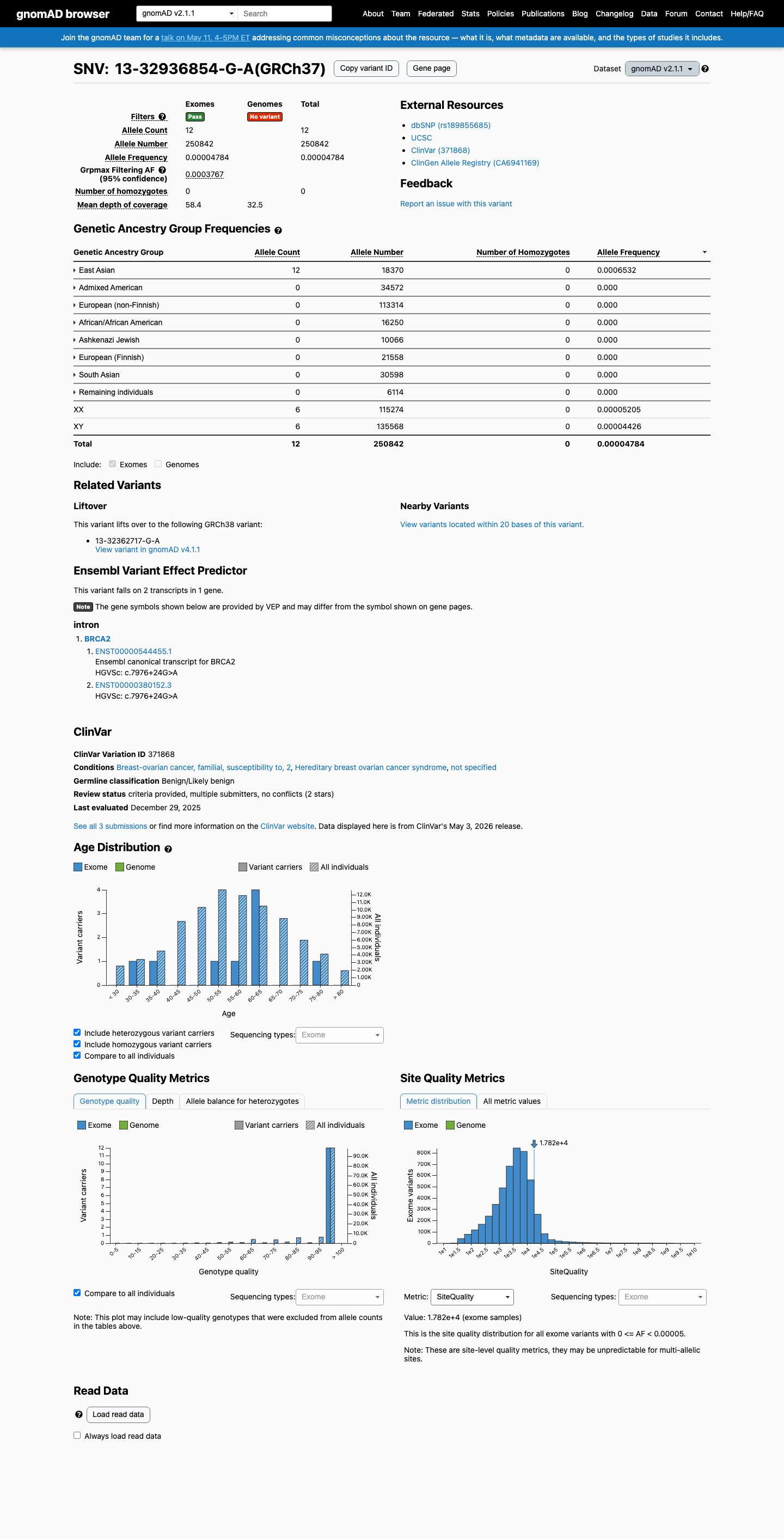
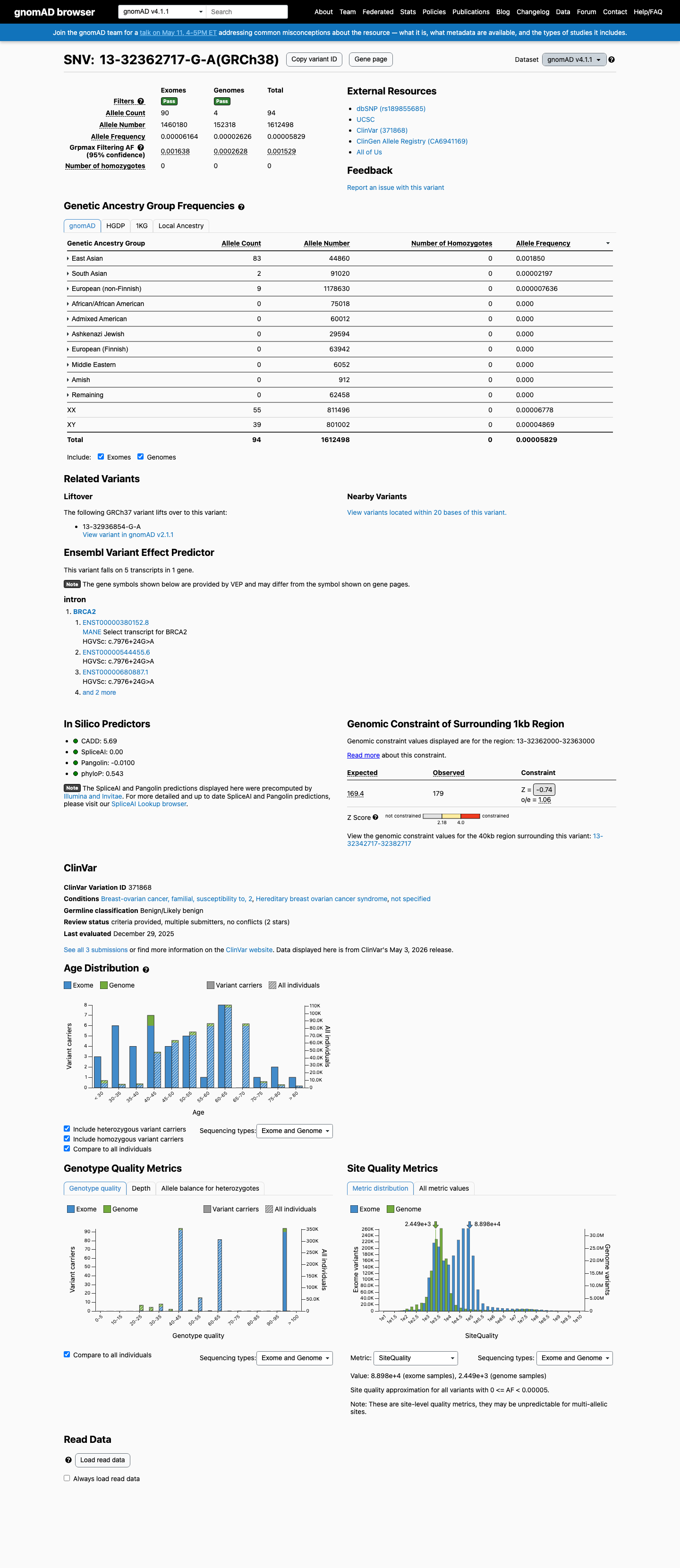
This variant is present in population databases, including gnomAD v2.1 at 12/250842 alleles with an East Asian grpmax filter allele frequency of 0.00037674 and gnomAD v4.1 at 94/1612498 alleles; the gnomAD v2.1 frequency exceeds the ENIGMA BRCA2 BS1 strong threshold of 0.0001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
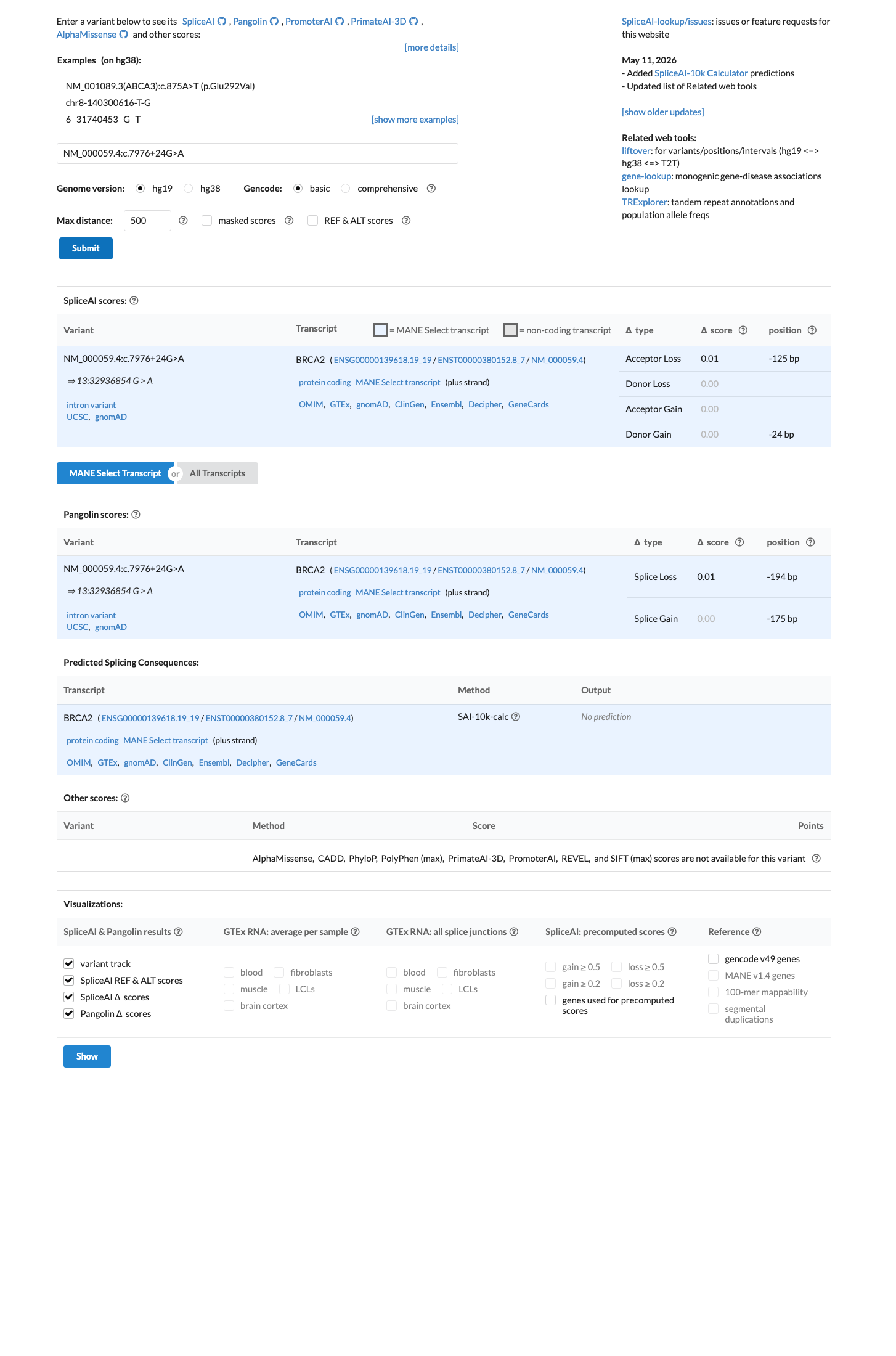
In silico splicing prediction does not support an abnormal splicing effect, with a SpliceAI maximum delta score of 0.00, consistent with BP4 and BP7 and arguing against PP3.
spliceai ↗ cspec ↗