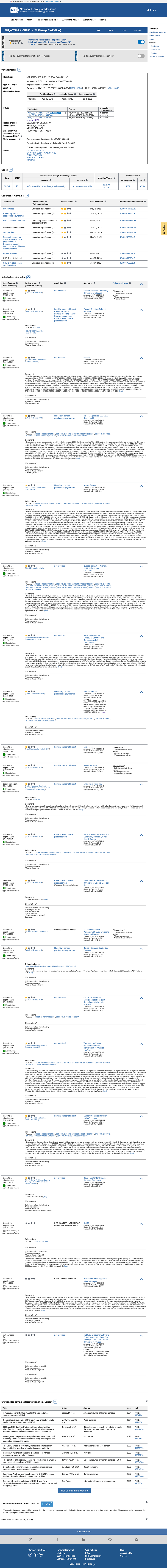
The CHEK2 c.715G>A (p.Glu239Lys, p.E239K) variant has been reported in ClinVar with predominantly uncertain significance submissions and one likely pathogenic submission.
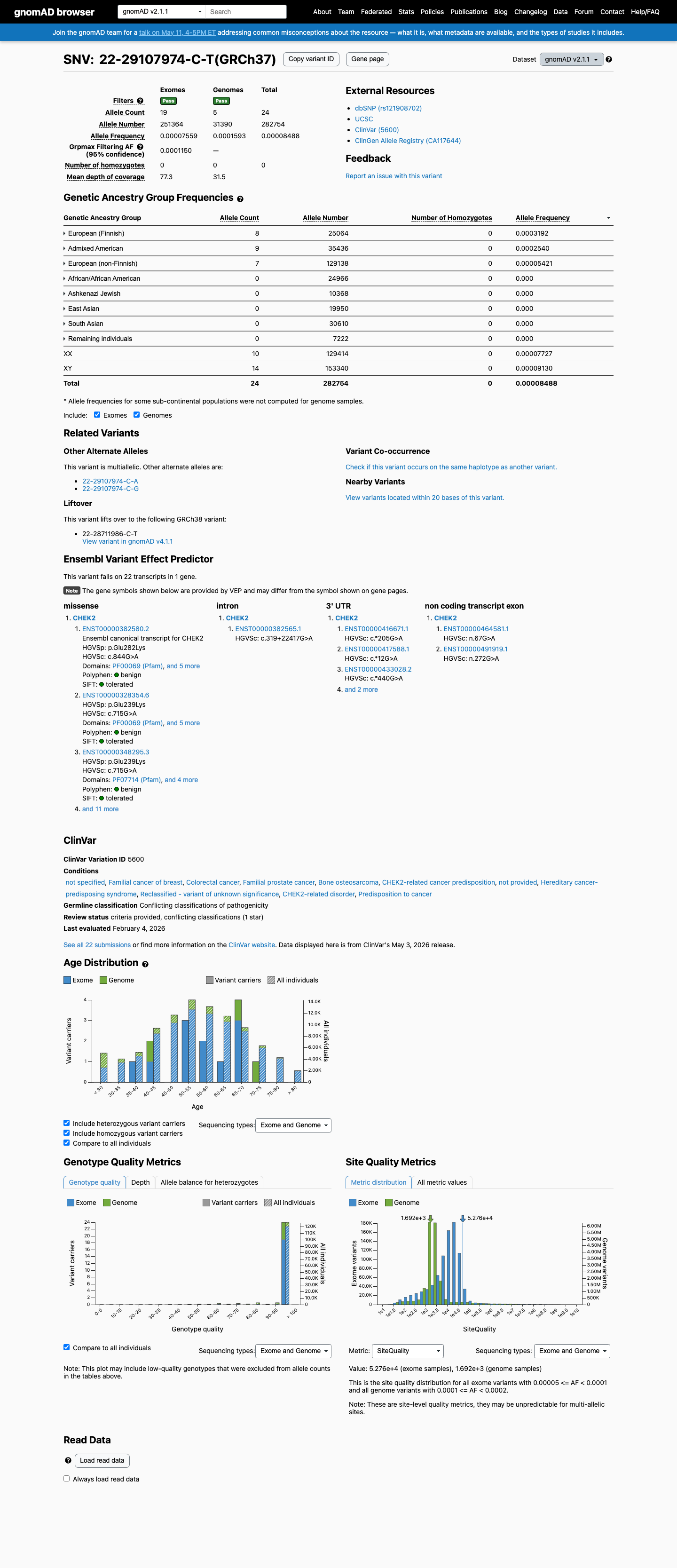
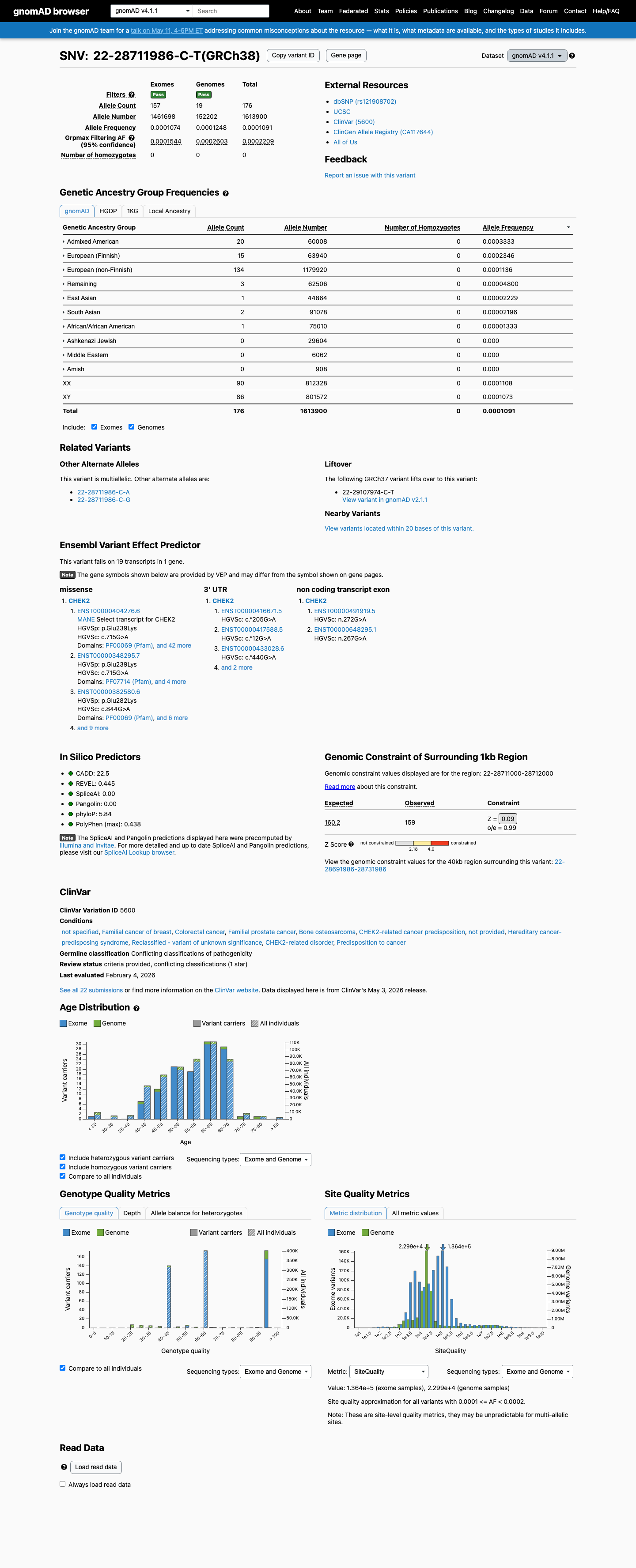
clinvar ↗This variant is present in gnomAD at 0.00849% in v2.1 and 0.01091% in v4.1, with a highest observed subpopulation frequency of 0.03333%, which is below the 0.1% rarity threshold and does not reach BS1 or BA1 population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗In a published yeast-based DNA-damage response assay, p.E239K had an intermediate score of 0.58 relative to 1.00 for wild type and 0.00 for a kinase-dead control, which is consistent with partial functional impairment but does not by itself establish a definitive functional criterion strength.
PMID:22419737 ↗Computational evidence is mixed: SpliceAI predicts no significant splice impact with a max delta score of 0.00, REVEL is 0.445, and BayesDel is 0.263648, so in silico evidence alone does not clearly support either a damaging or benign computational criterion.
spliceai ↗