Classification rationale
1
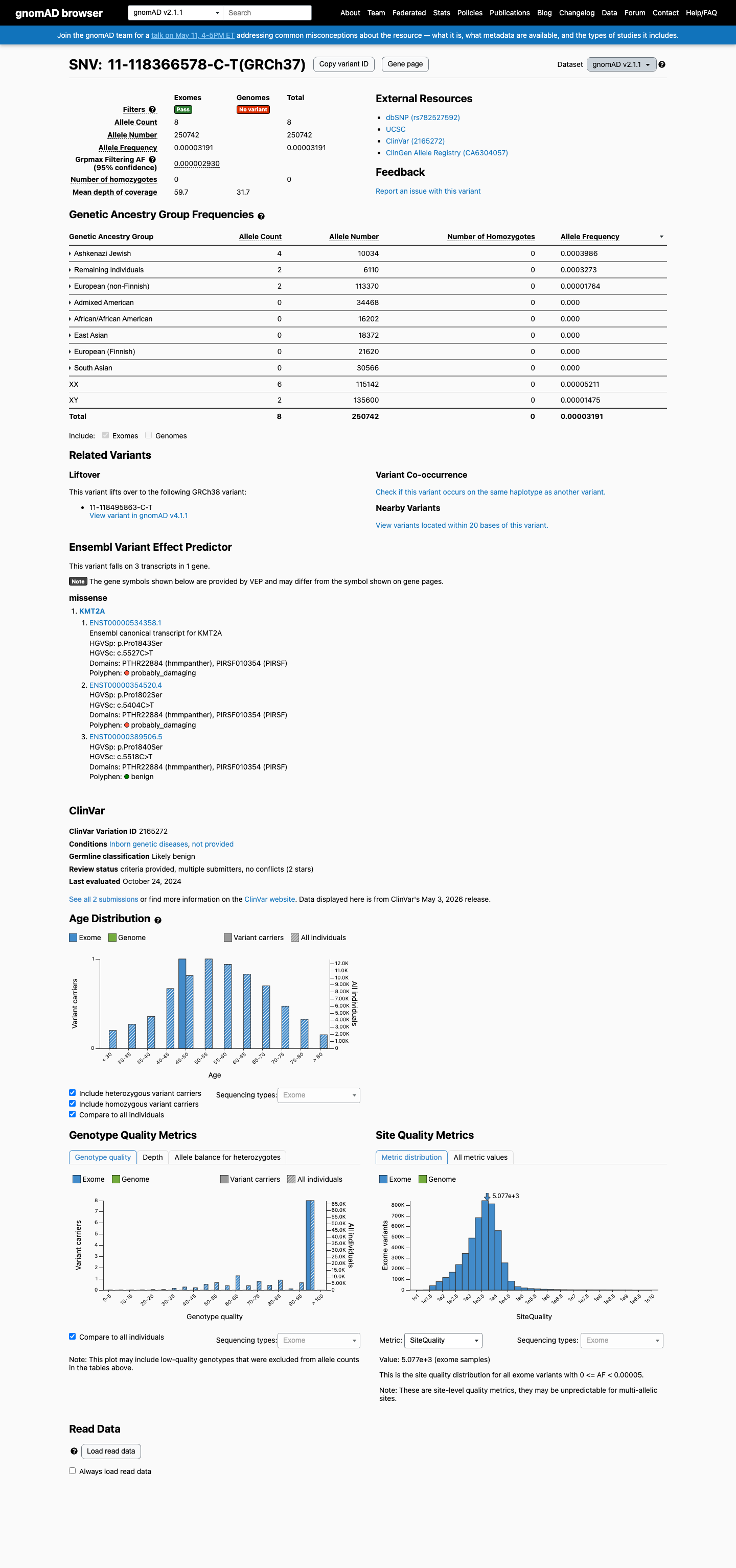
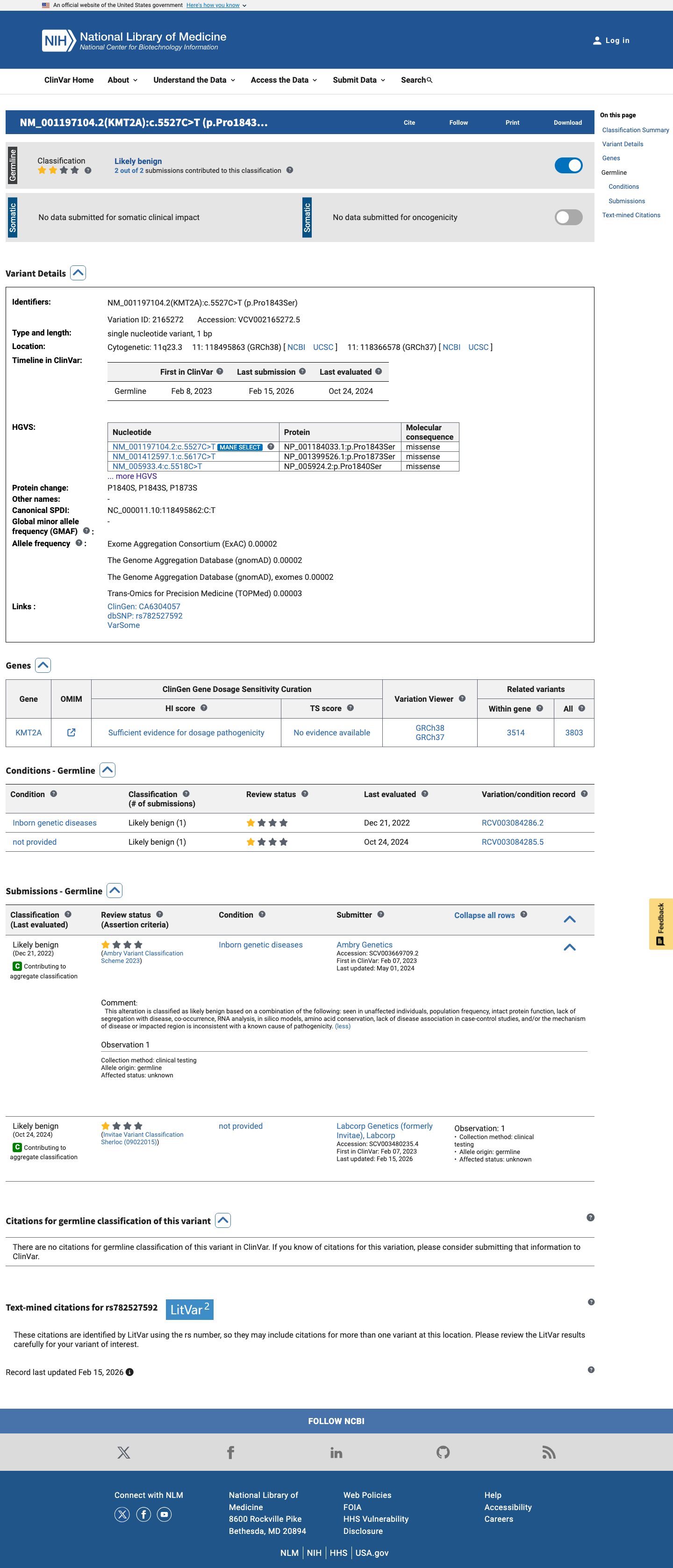
The KMT2A NM_005933.3:c.5518C>T (NP_005924.2:p.(Pro1840Ser)) variant has been reported in ClinVar as likely benign by two clinical laboratories, without expert panel review.
clinvar ↗2
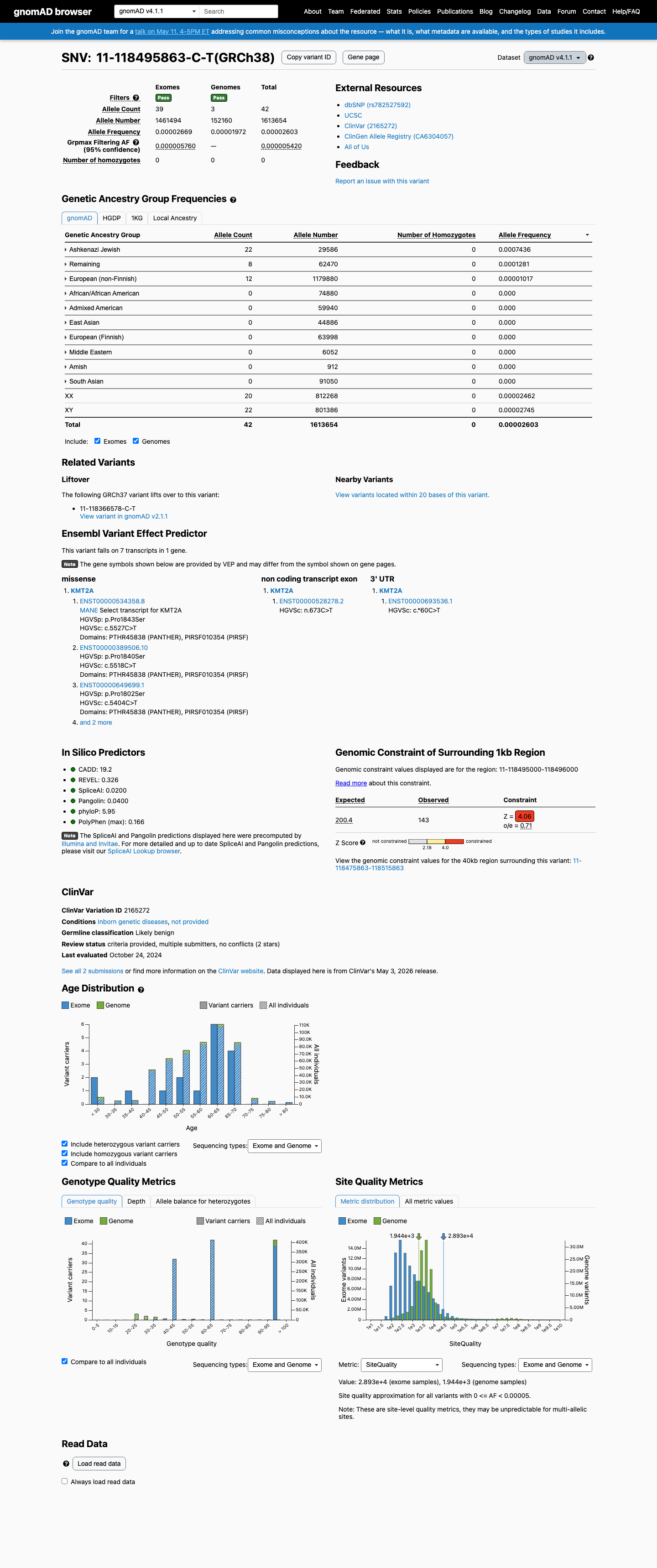
This variant is present in gnomAD v2.1 at 0.00319% (8/250,742 alleles) and in gnomAD v4.1 at 0.00260% (42/1,613,654 alleles), with highest observed Ashkenazi Jewish frequencies of 0.03986% and 0.07436%, respectively; these values are below the default 0.1% PM2 threshold and below the 0.3% BS1 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
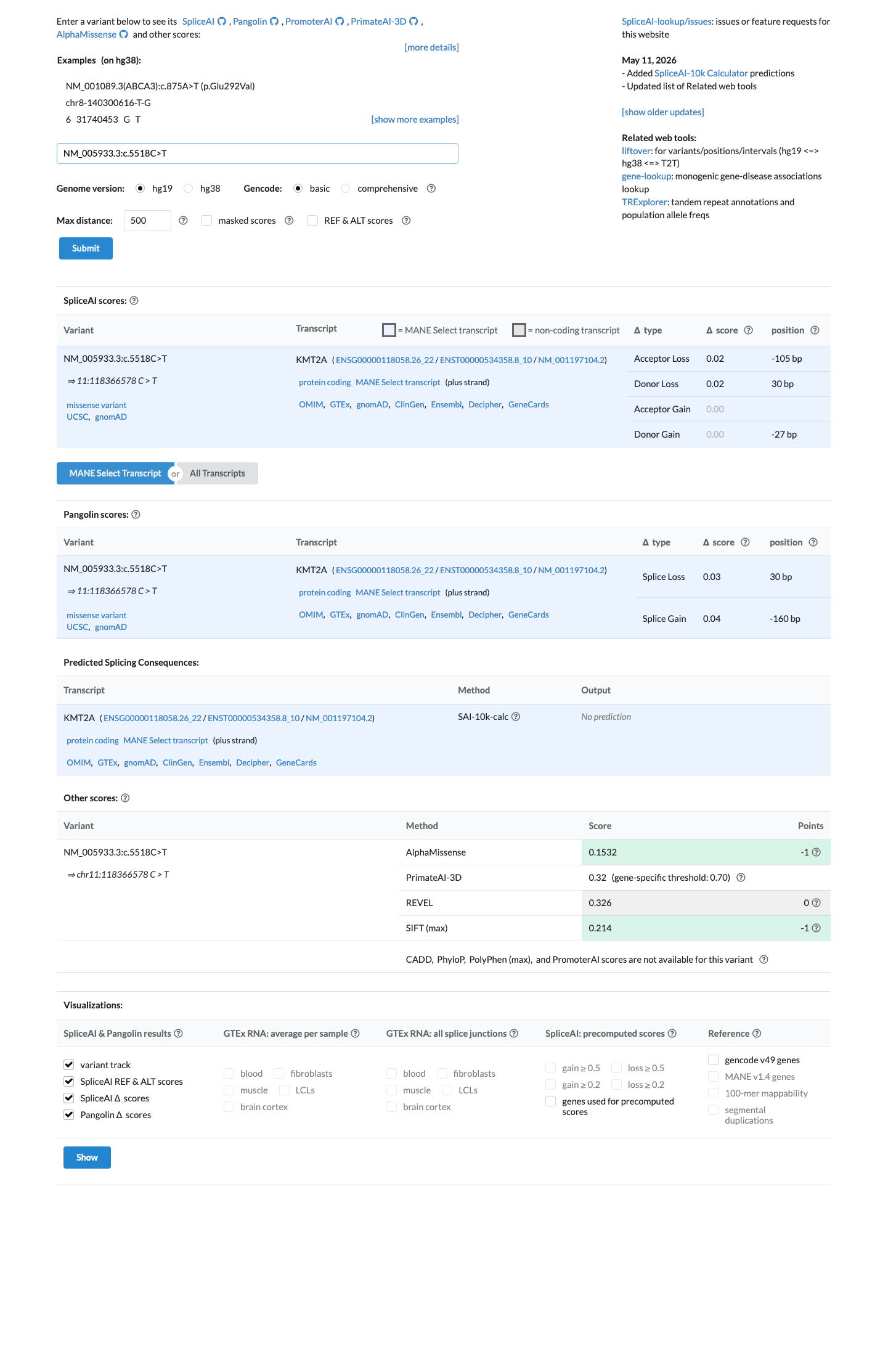
Computational evidence does not support a damaging effect: SpliceAI predicts no significant splice impact with a maximum delta score of 0.02, REVEL is 0.326, and BayesDel is -0.184716.
spliceai ↗4
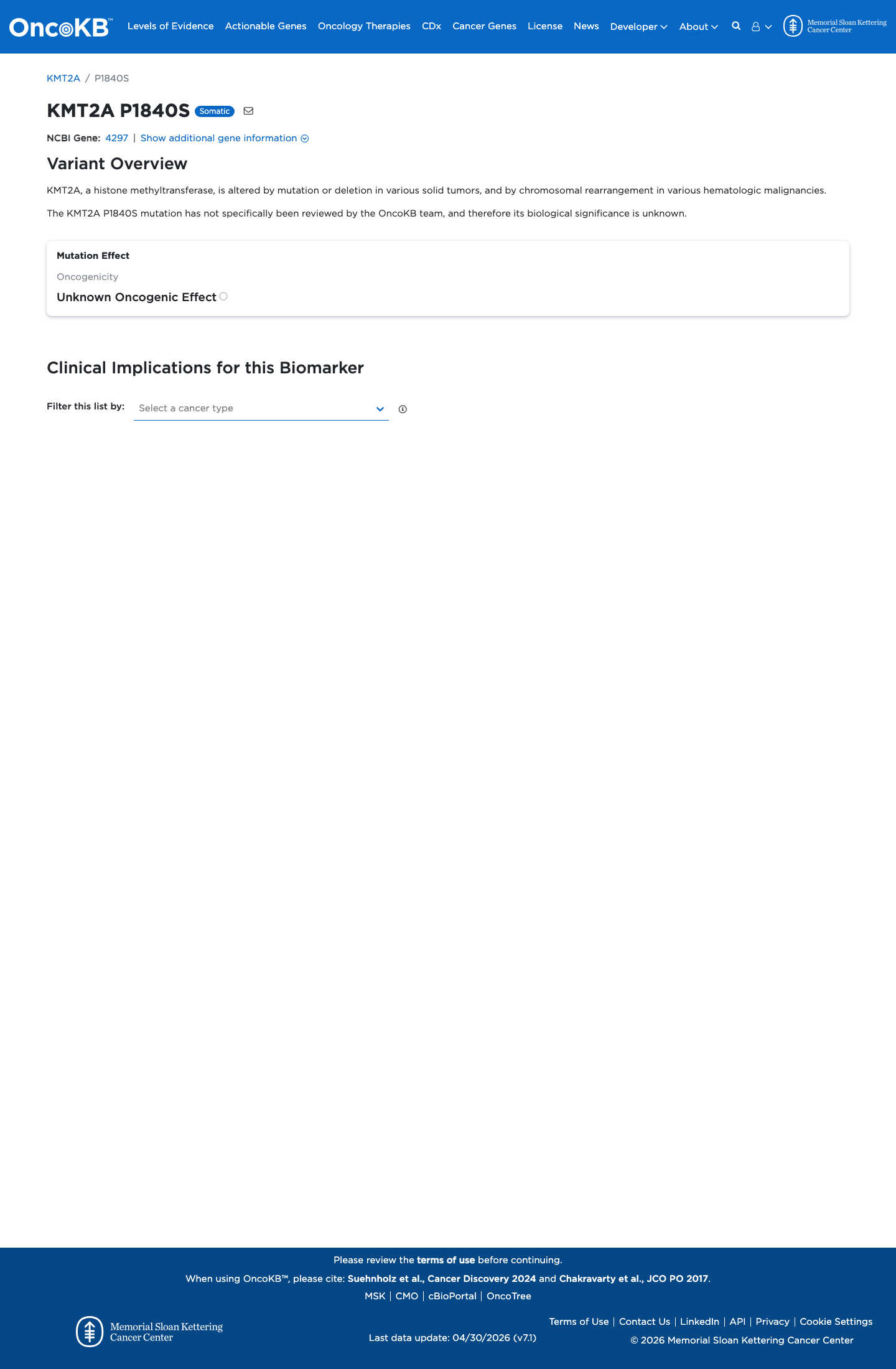
This variant has not been shown to lie in a statistically significant hotspot.
hotspots ↗