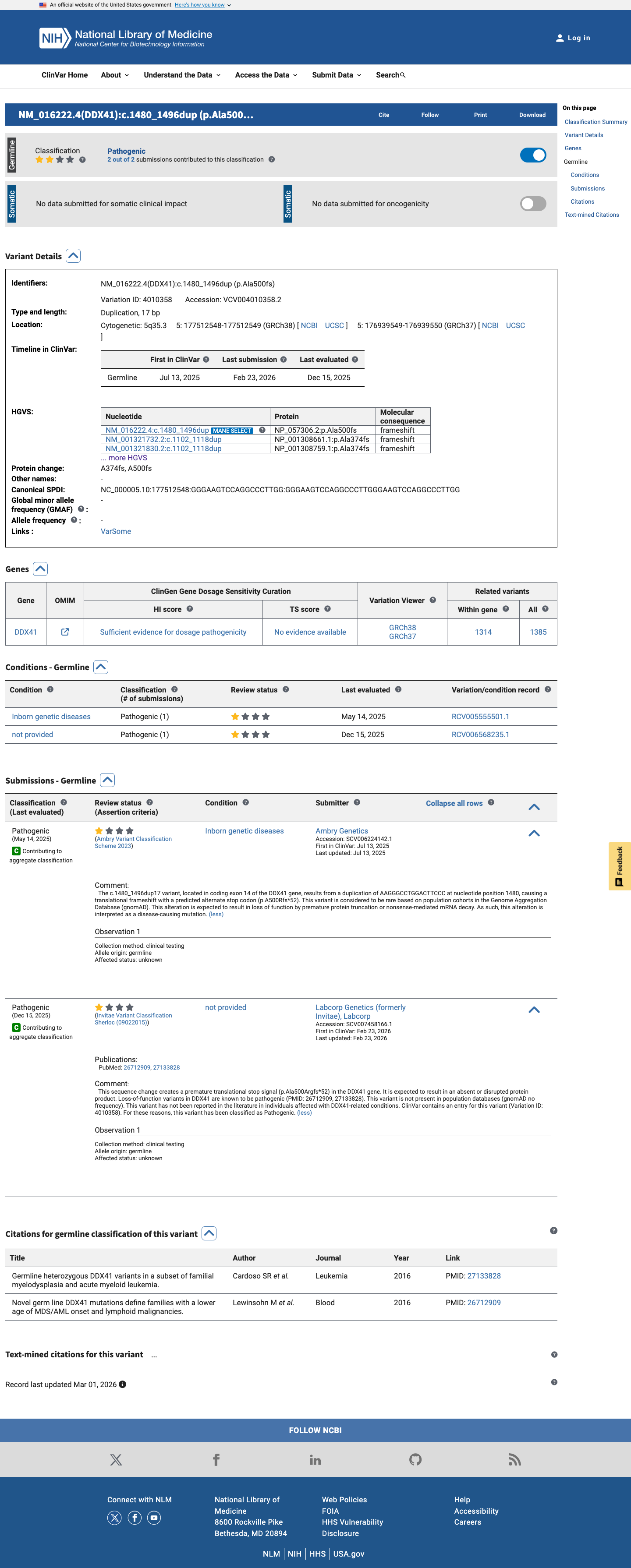
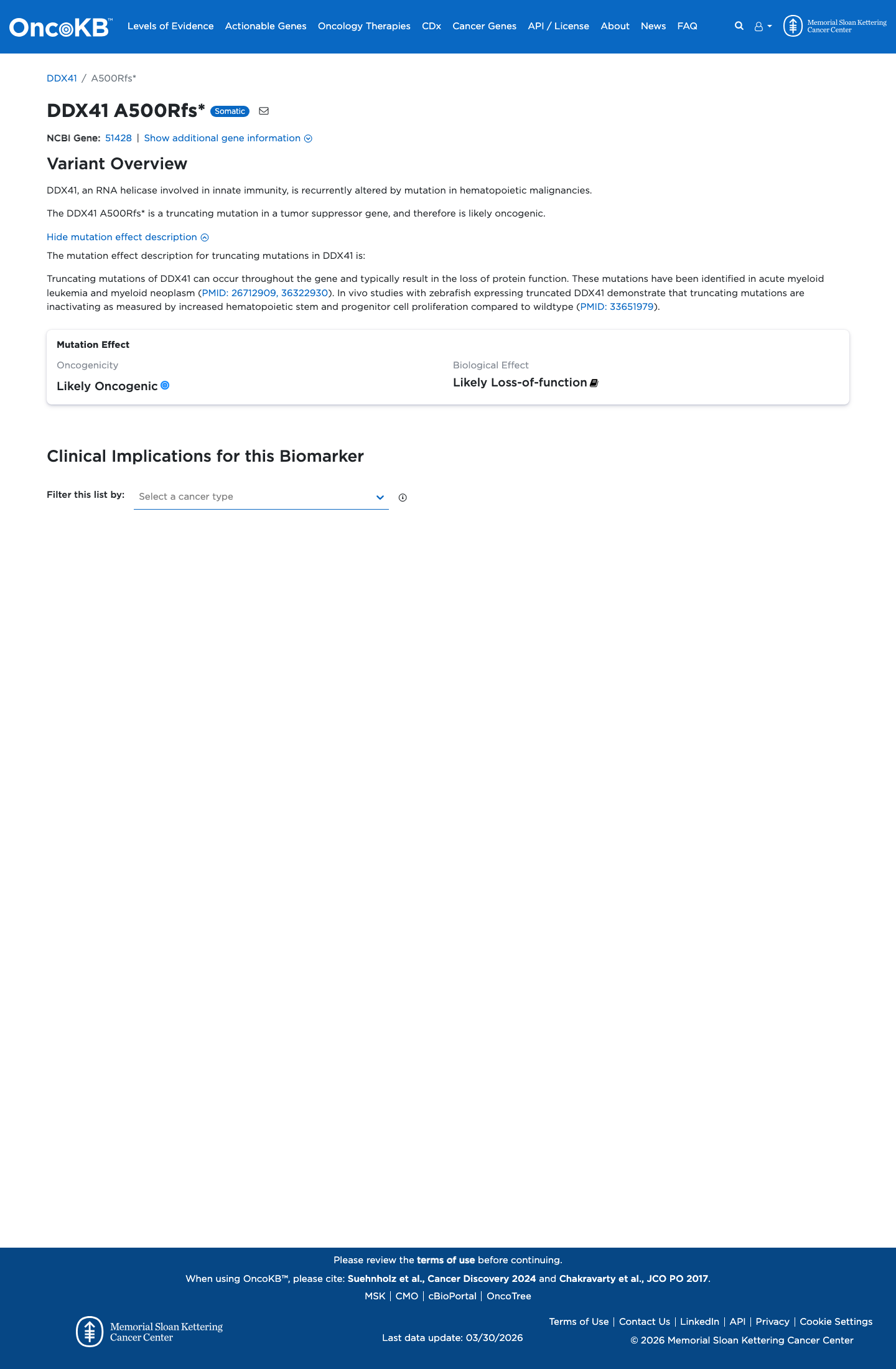
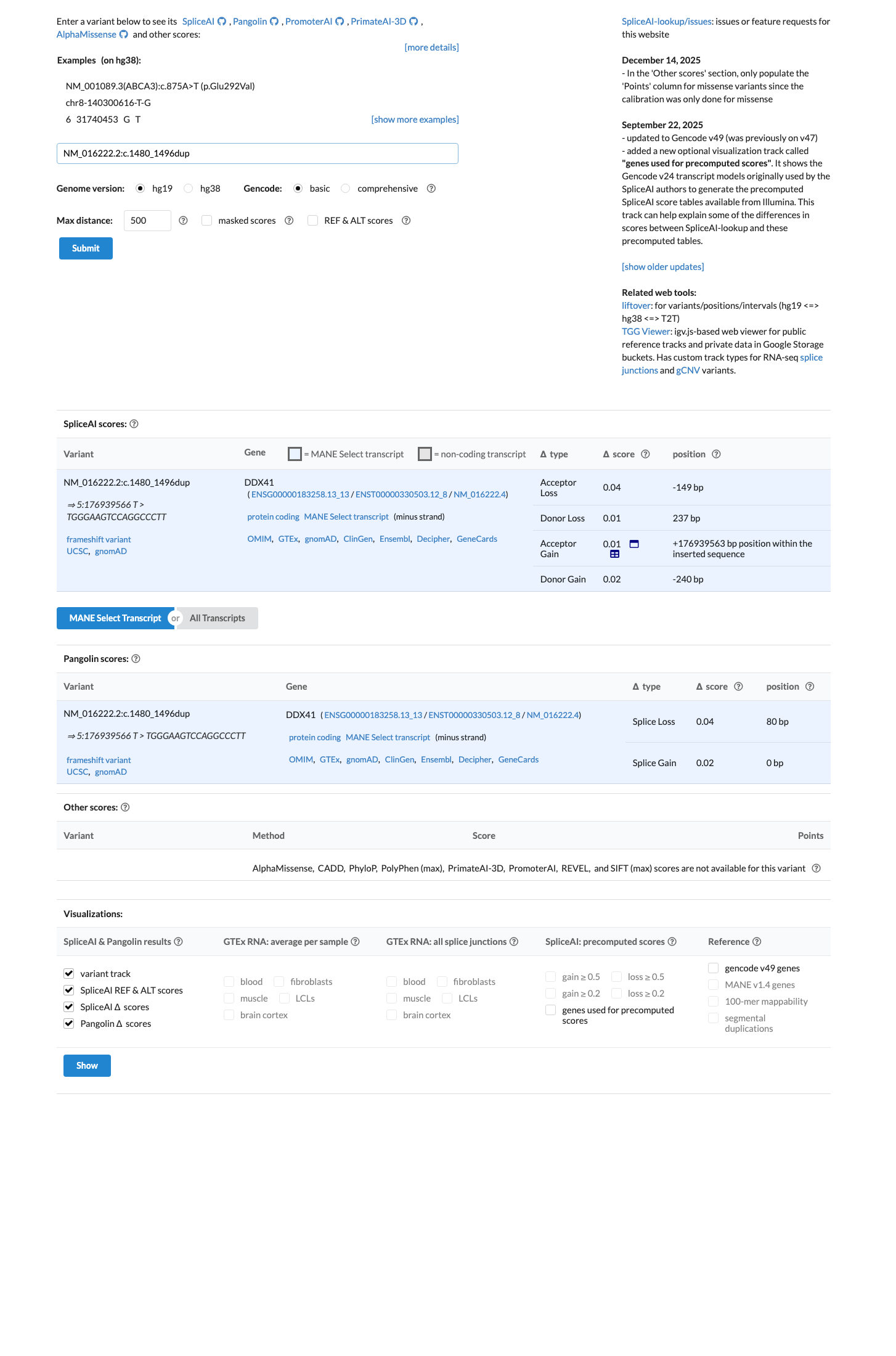
NM_016222.2:c.1480_1496dup is a frameshifting duplication predicted to produce p.(Ala500ArgfsTer52) [p.(A500Rfs*52)], altering the reading frame from codon 500 and truncating the normal C-terminal portion of DDX41; this is consistent with a loss-of-function effect in a gene for which loss of function has been implicated in DDX41-related hematologic malignancy predisposition syndrome.
oncokb ↗The variant is absent from gnomAD v2.1 and is seen once in gnomAD v4.1 at AF 6.84053e-07 (1/1,461,874 alleles), with highest observed population AF 8.99276e-07 in non-Finnish Europeans, supporting marked rarity in population databases.
gnomad_v4 ↗Because the predicted premature termination occurs in the terminal exon, the available loss-of-function evidence is most appropriately weighted below full very-strong strength, and no additional independently curated case-level or functional evidence in the supplied materials upgrades the variant beyond the likely pathogenic range.
Overall, the available evidence supports classification of NM_016222.2:c.1480_1496dup (DDX41 p.(Ala500ArgfsTer52)) as Likely Pathogenic.
clinvar ↗