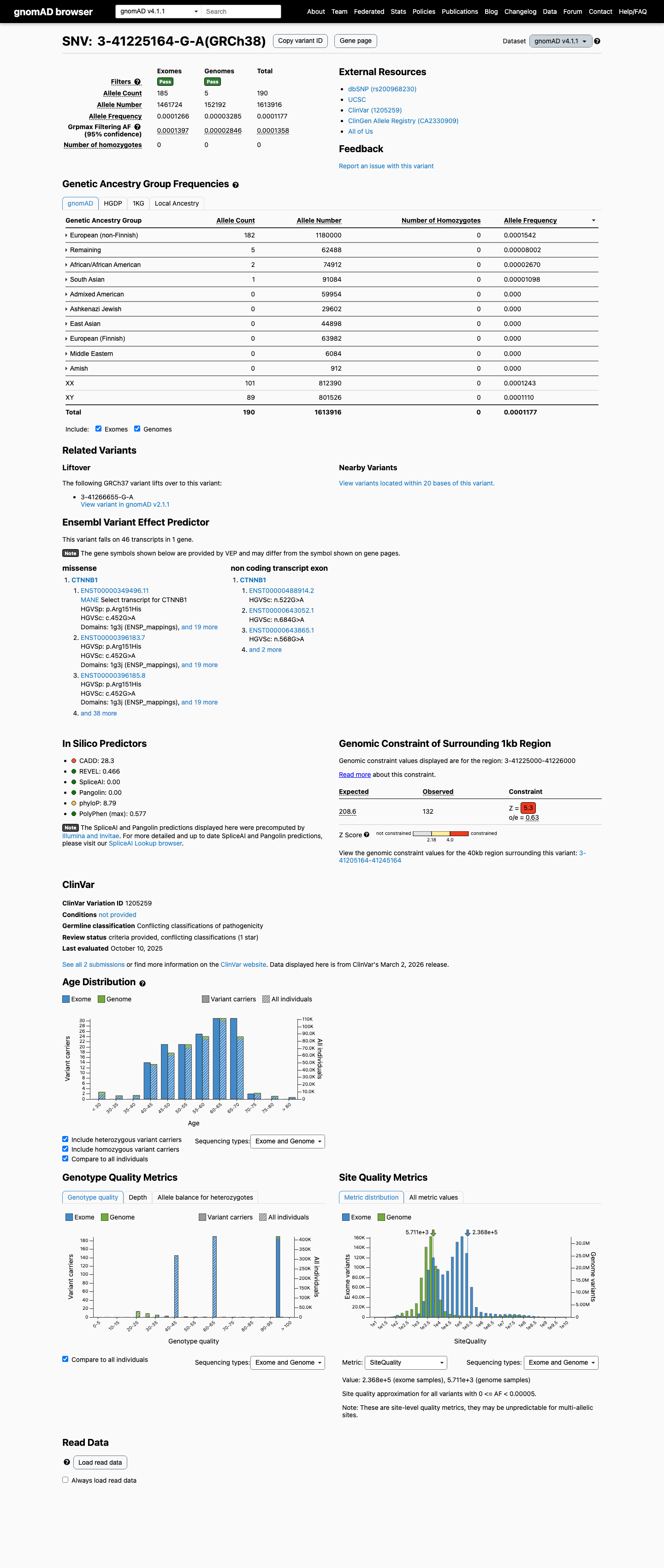
CTNNB1 NM_001904.4:c.452G>A, p.(Arg151His) is present in gnomAD v4.1 at a total allele frequency of 0.000117726 (190/1613916) with a highest observed population frequency of 0.000154237 in European non-Finnish individuals, both below the non-VCEP PM2 rarity threshold of 0.001 (0.1%), supporting rarity but not absence from population databases.
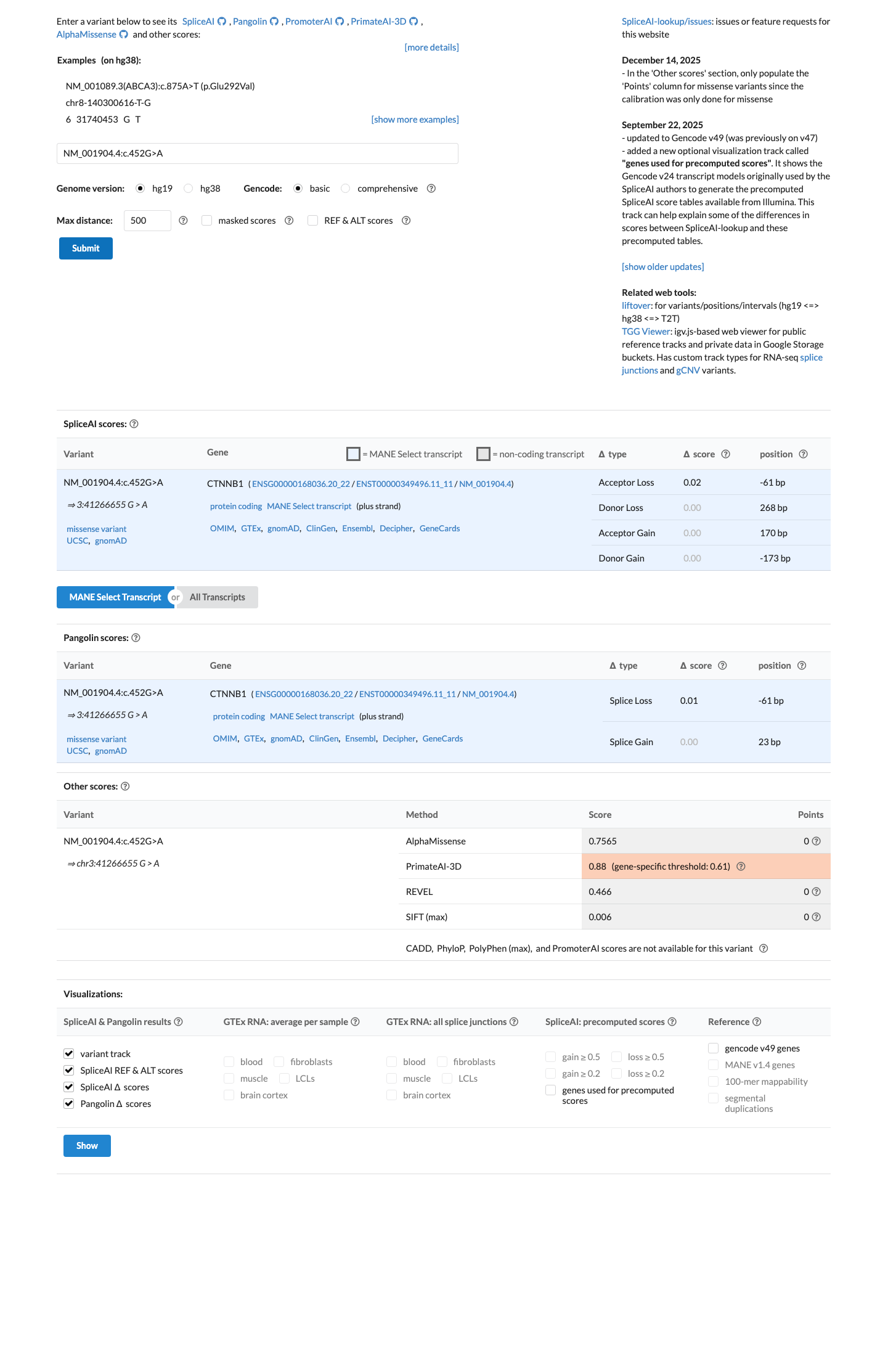
gnomad_v4 ↗SpliceAI predicts a maximum delta score of 0.02, which is below the splice impact threshold of 0.2 and argues against a splice-disrupting effect.
spliceai ↗ClinVar contains conflicting single-submitter classifications of likely benign and uncertain significance, so this assertion set does not provide decisive classification evidence.
clinvar ↗In the absence of gene-specific criteria, robust case-level enrichment, segregation, de novo, or functional evidence, CTNNB1 p.(Arg151His) is classified as a variant of uncertain significance.