Classification rationale
1
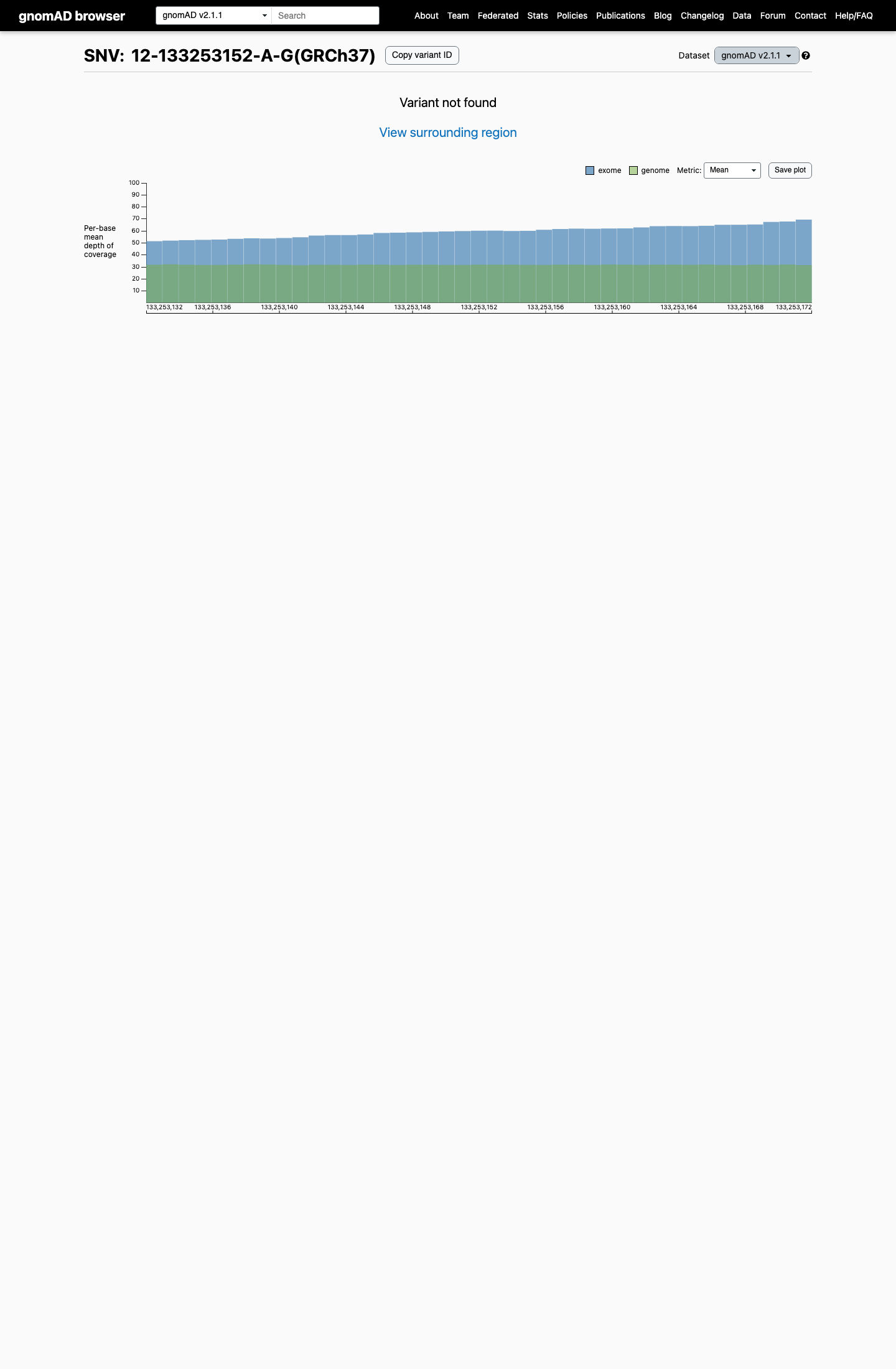
POLE p.(Ser297Pro) is absent from gnomAD v2.1.
gnomad_v2 ↗2
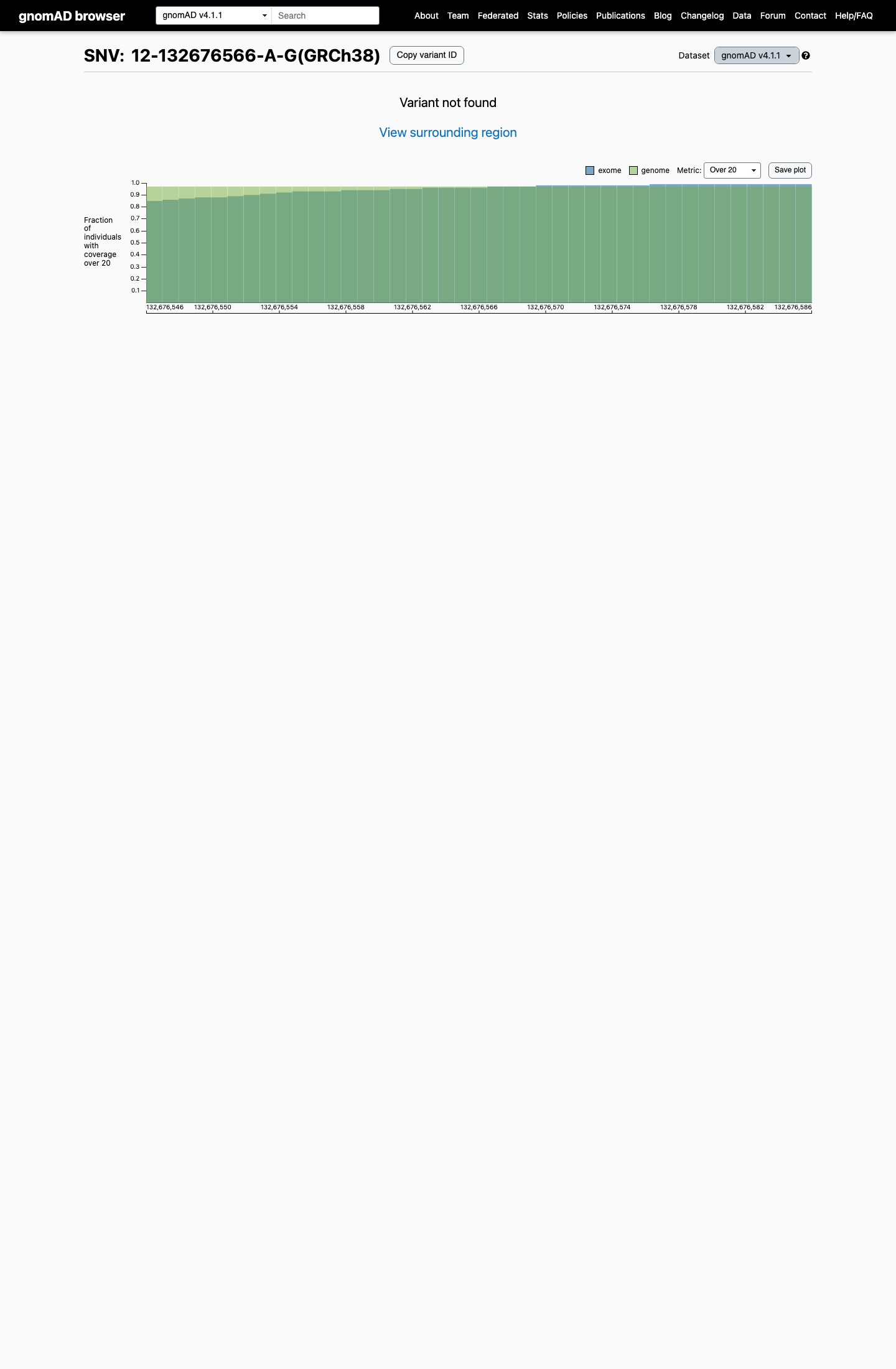
The variant is also absent from gnomAD v4.1, supporting rarity, although no explicit POLE-specific population cutoff was retrieved.
gnomad_v4 ↗3
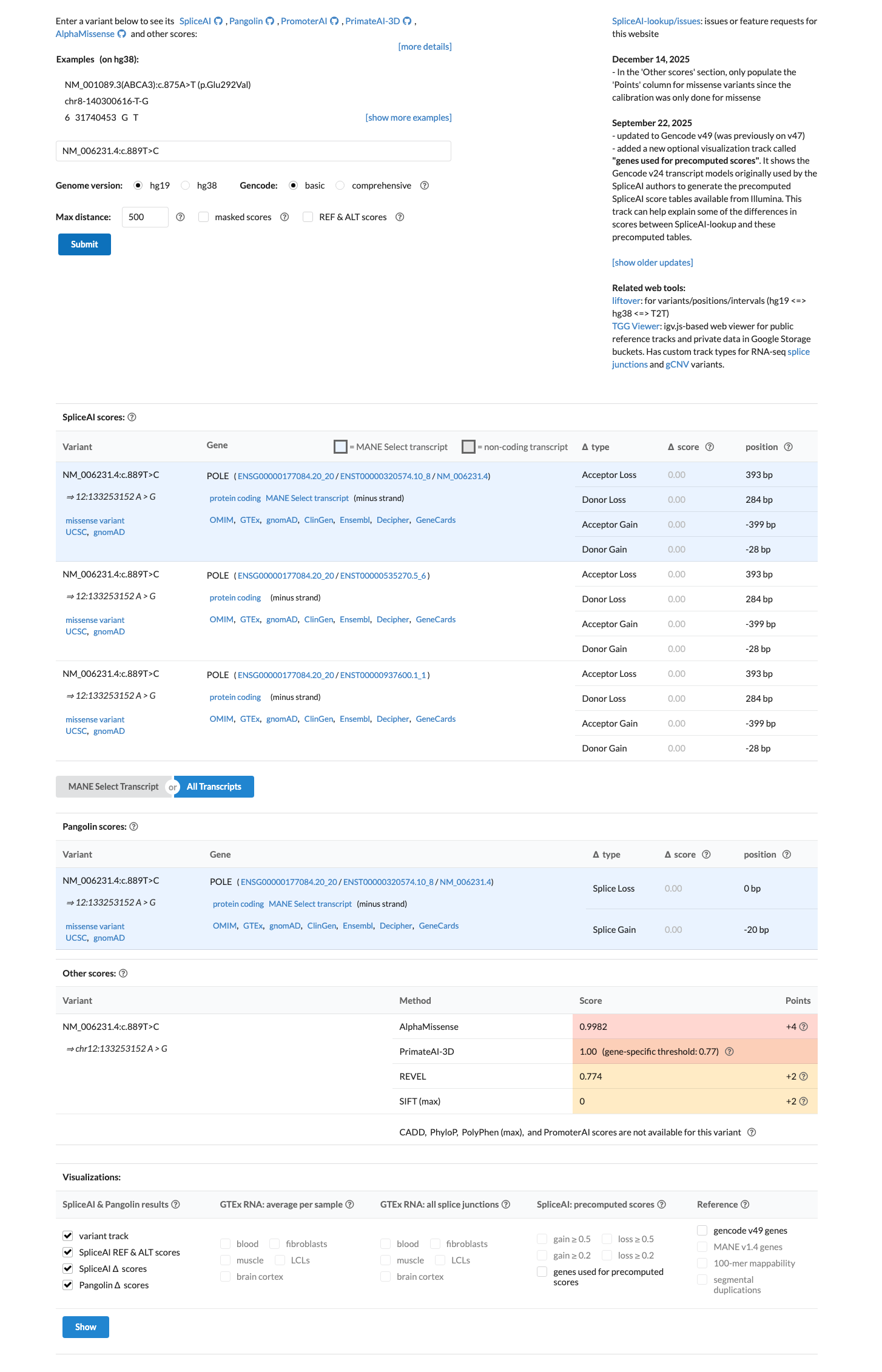
SpliceAI predicts no significant splice effect with a maximum delta score of 0.00, but no explicit computational threshold framework was retrieved to convert the available in silico data into PP3 or BP4.
spliceai ↗4
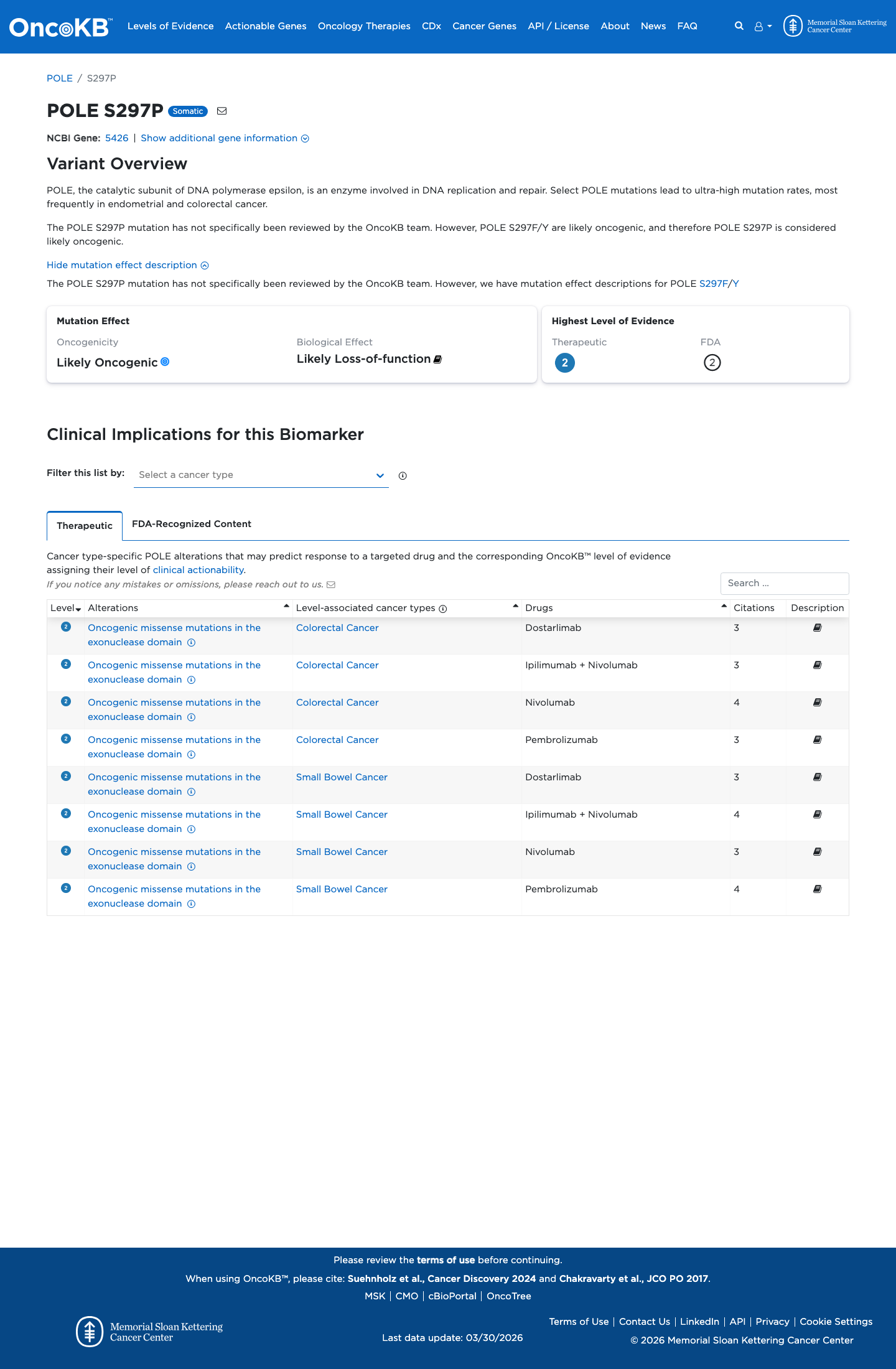
No ClinVar classification has been reported for this variant.
clinvar ↗5
With PM2 applied and no additional pathogenic or benign criteria established, this variant is classified as a variant of uncertain significance under the retrieved generic ACMG/AMP combination rules.