Classification rationale
1
The ATM c.2207C>T (p.Ala736Val) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as a variant of uncertain significance by 4 clinical laboratories.
clinvar ↗2
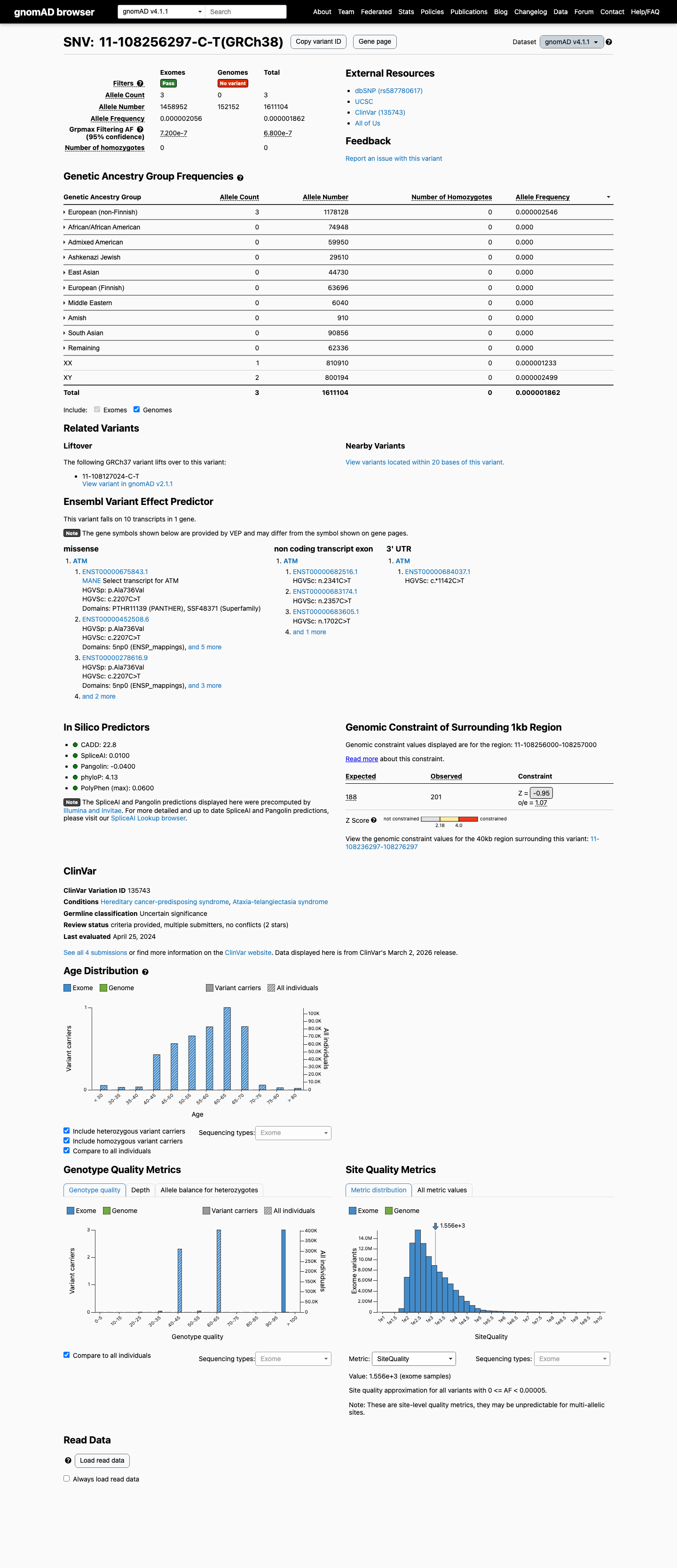
The variant is present at very low frequency in gnomAD, including 3 of 1458952 alleles in v4.1 with no homozygotes, which supports PM2_Supporting and is well below the ATM BS1 and BA1 frequency thresholds.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗3
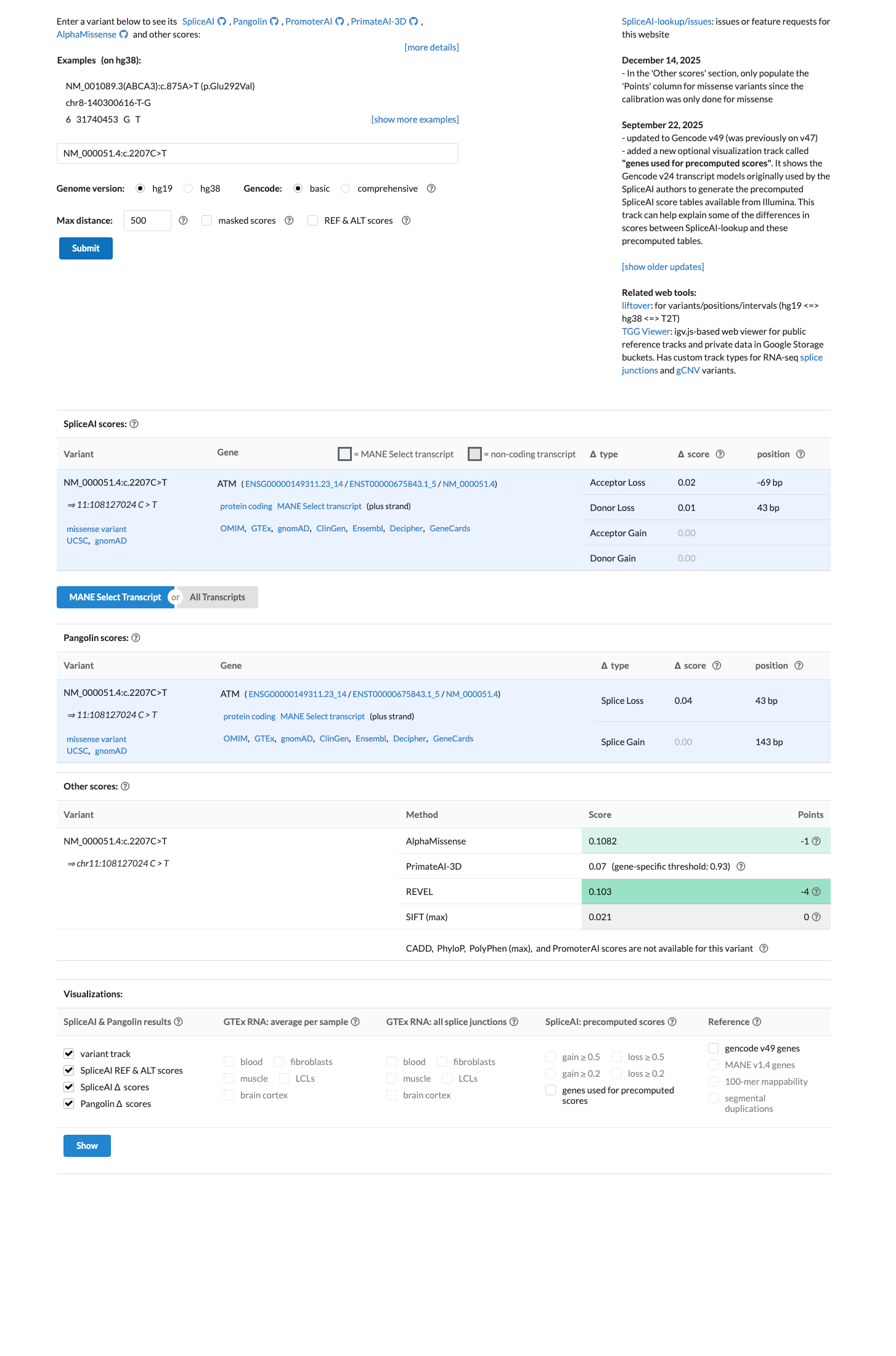
In silico evidence argues against a deleterious effect: REVEL is 0.103 and SpliceAI shows a maximum delta score of 0.02, supporting BP4 and arguing against PP3.
spliceai ↗ cspec ↗