Classification rationale
1
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, so the observed allele frequency is 0, which is below the RUNX1 MM-VCEP PM2_Supporting threshold of less than or equal to 0.00005.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
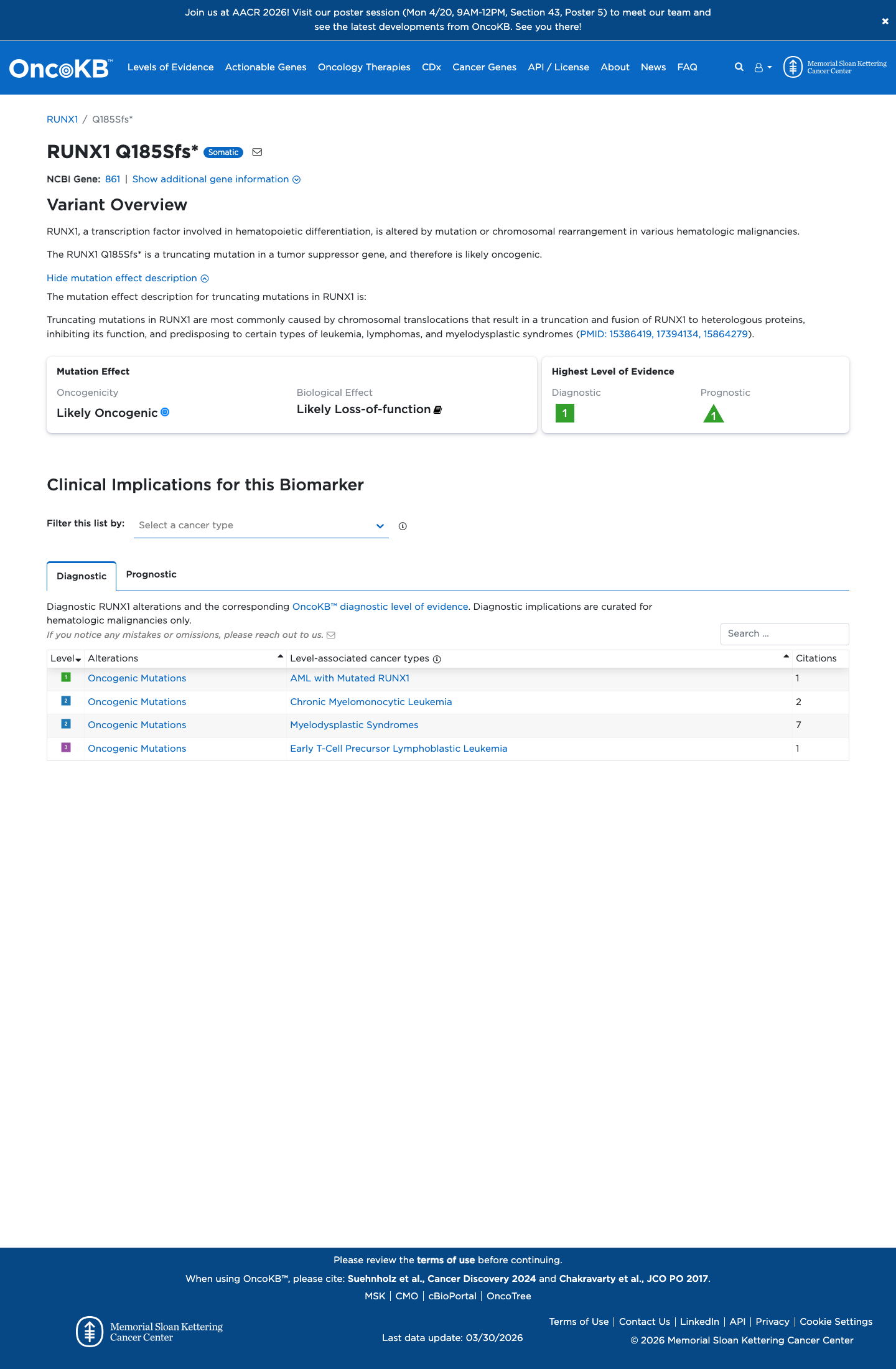
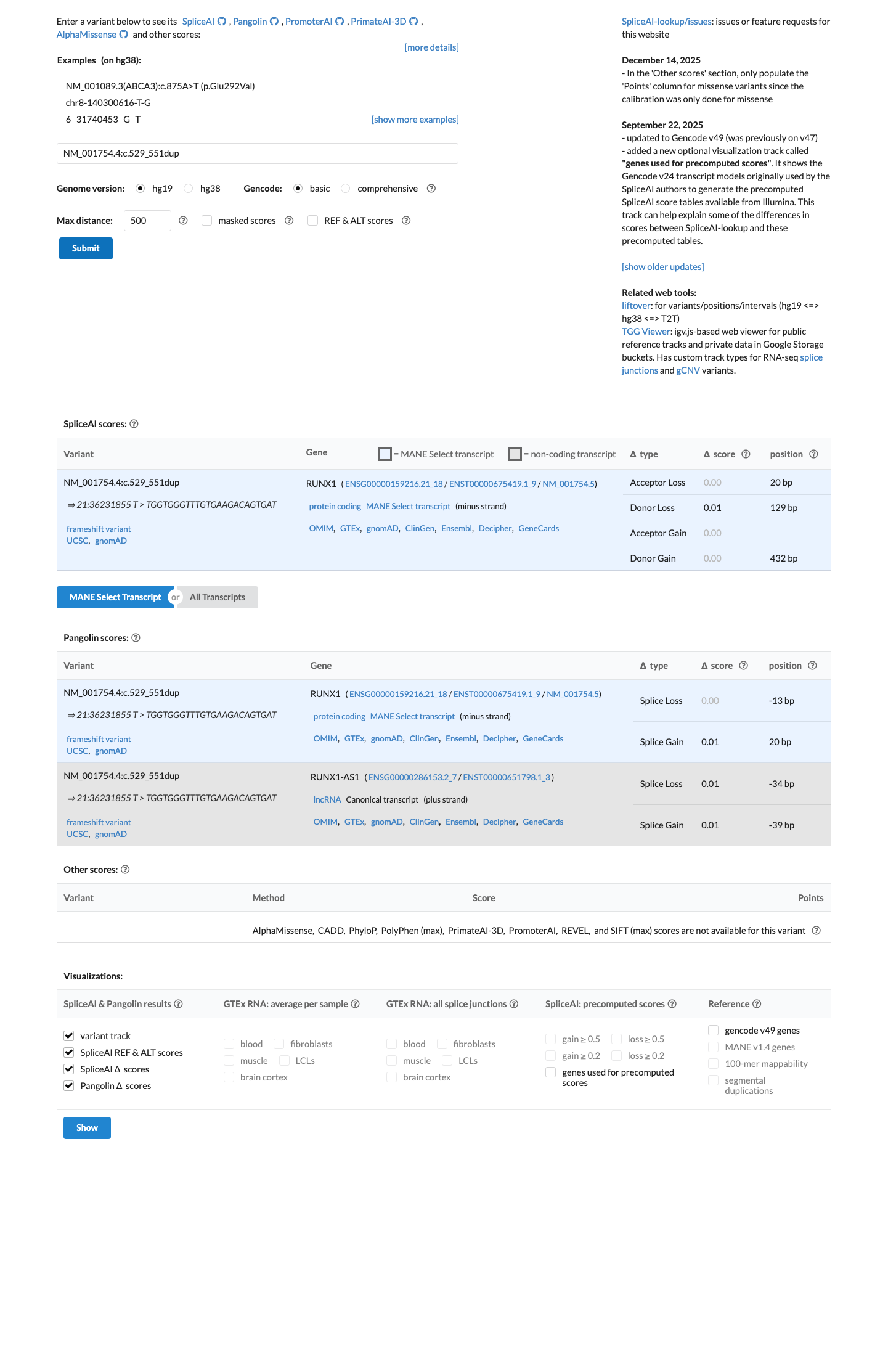
This duplication is predicted to cause a frameshift, p.(Gln185SerfsTer34), and SpliceAI predicts no significant splice impact with a maximum delta score of 0.01, which is consistent with applying PVS1 and PM5_Supporting under the RUNX1 MM-VCEP framework.
spliceai ↗ cspec ↗