Classification rationale
1
2
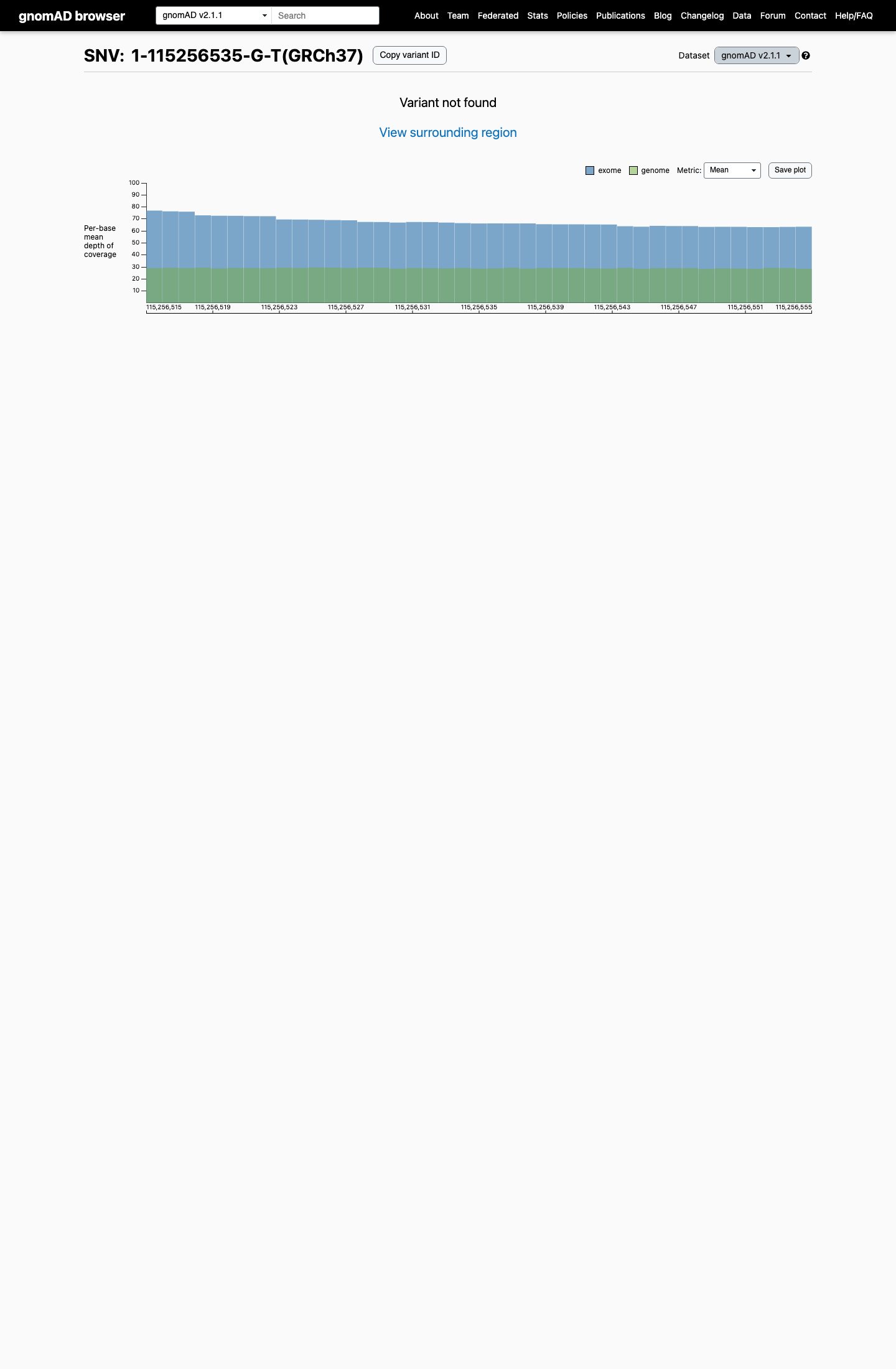
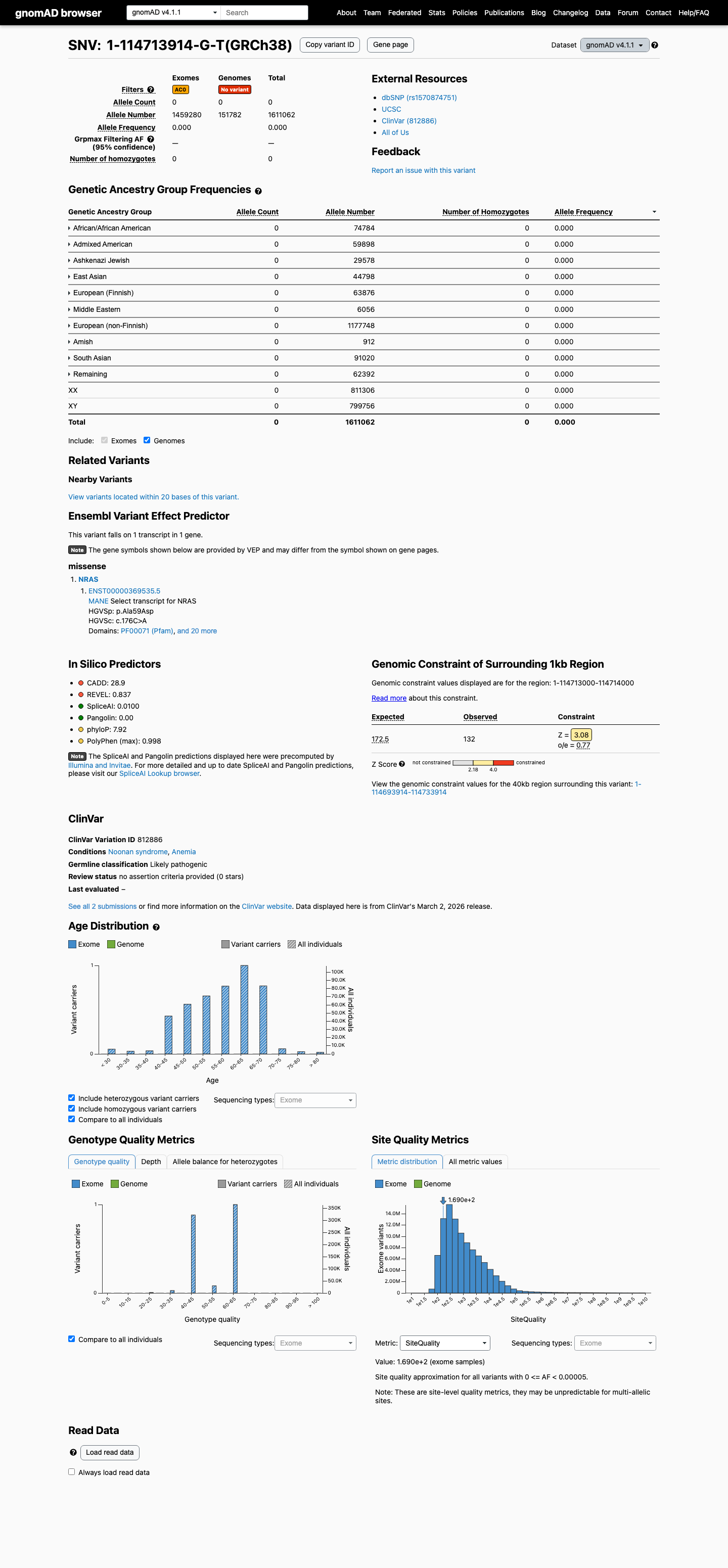
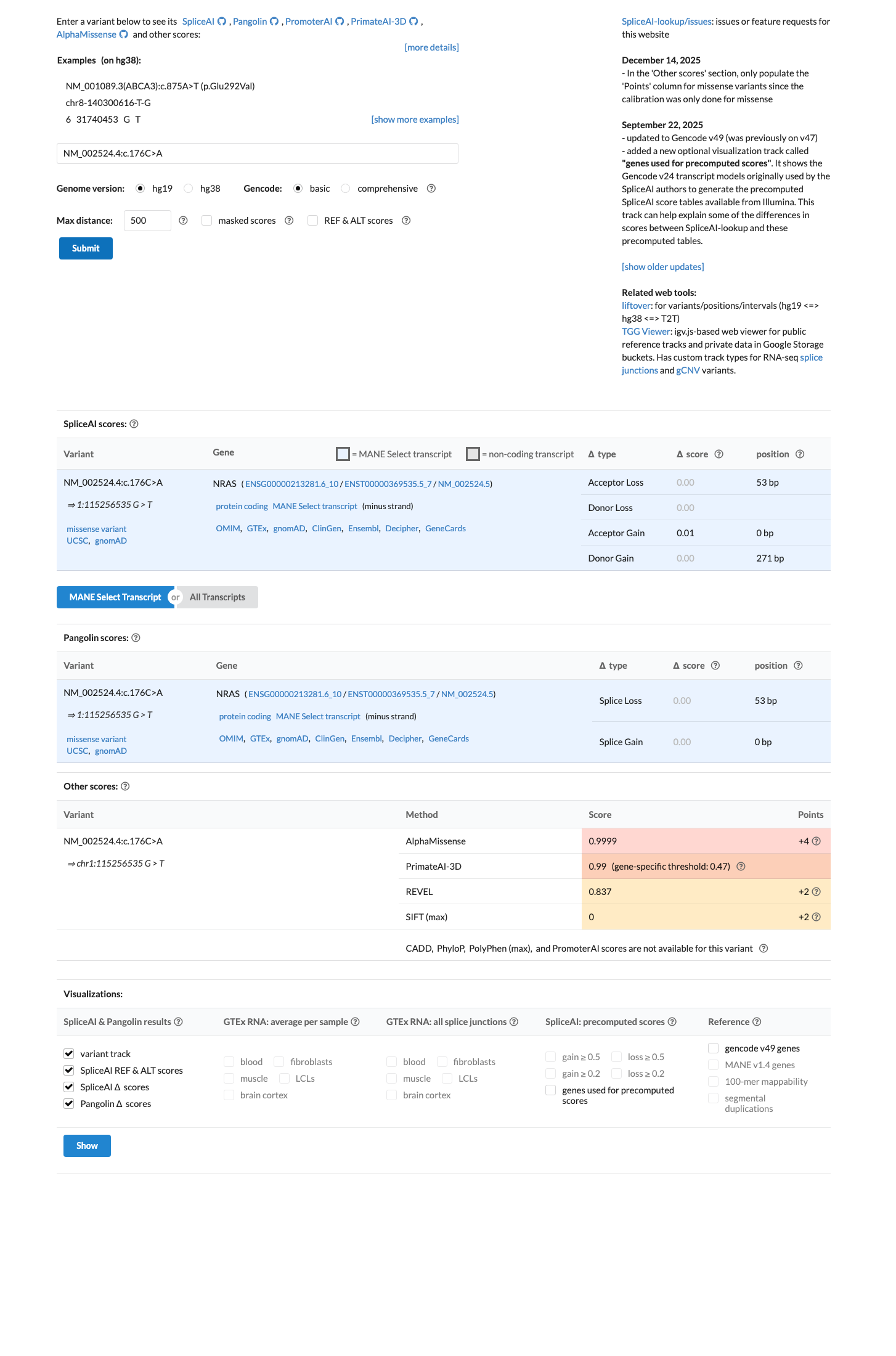
This variant is absent from gnomAD controls, with 0/1,611,062 alleles in gnomAD v4.1 and no observation in gnomAD v2.1, supporting PM2_Supporting and arguing against a benign population frequency threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
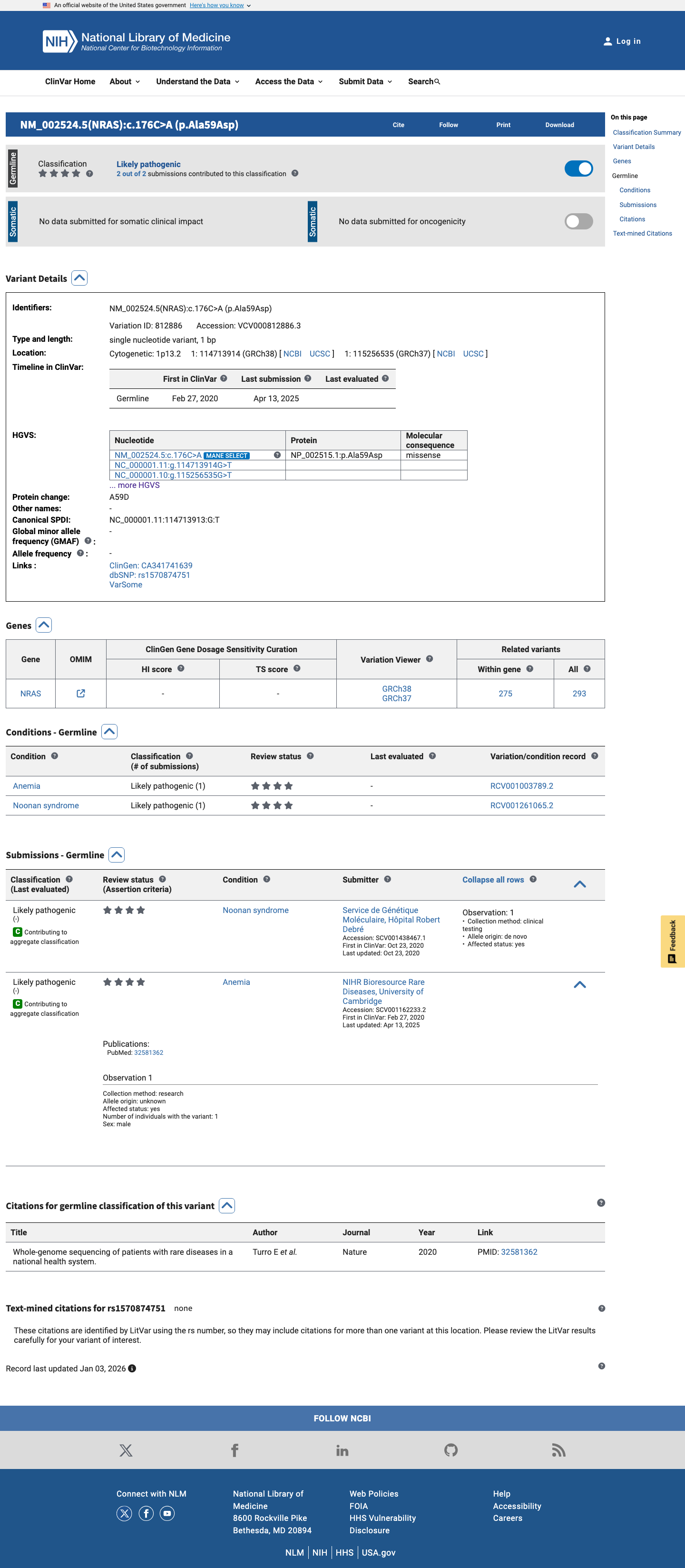
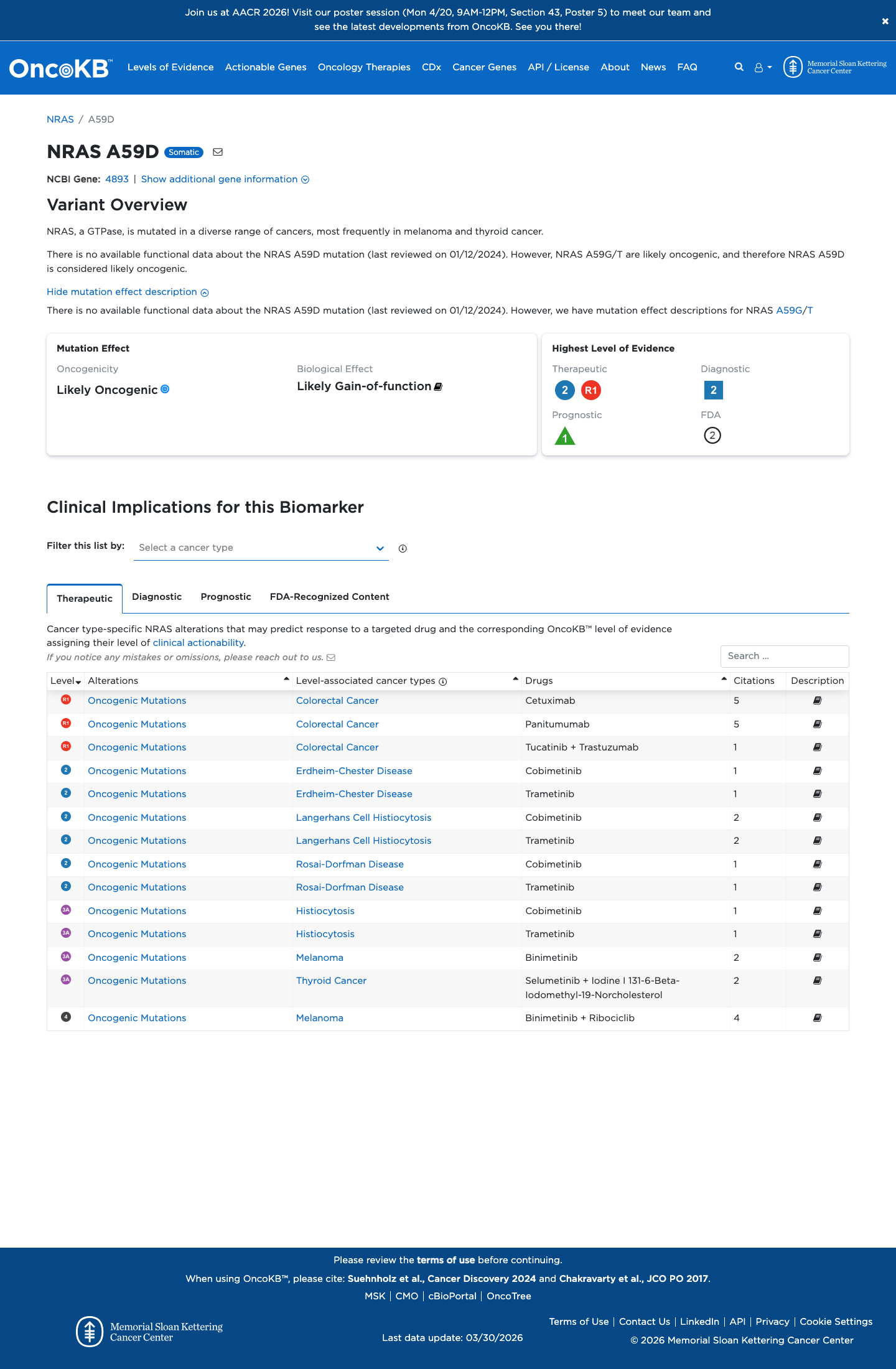
The variant is located in the NRAS Switch II domain (amino acids 57-64), a critical and well-established functional region specified for PM1 in the NRAS RASopathy criteria, but no ClinGen-approved variant-specific functional assay result was identified to apply PS3.
cspec ↗4
Computational evidence supports a deleterious missense effect because the REVEL score is 0.837, above the PP3 threshold of 0.7, while SpliceAI predicts no significant splice effect with a maximum delta score of 0.01.
spliceai ↗