Classification rationale
1
2
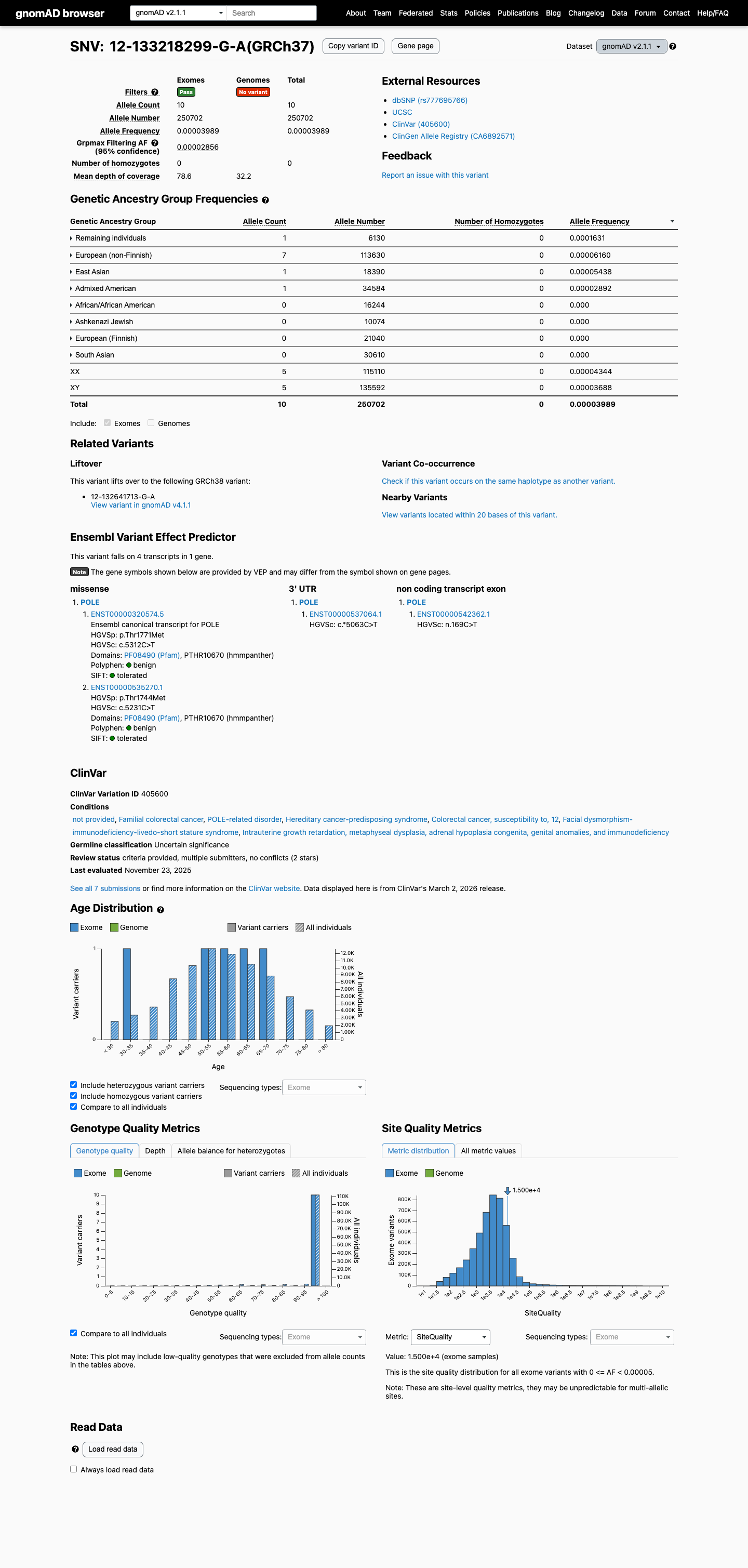
The variant is present at low frequency in population databases, with gnomAD v2.1 AF 3.9888e-05 (10/250702) and gnomAD v4.1 AF 4.09054e-05 (66/1613480), with no homozygotes reported; these values are below the PM2 threshold of 0.1% and below the BS1 and BA1 thresholds of 0.3% and 1%, respectively.
gnomad_v2 ↗ gnomad_v4 ↗3
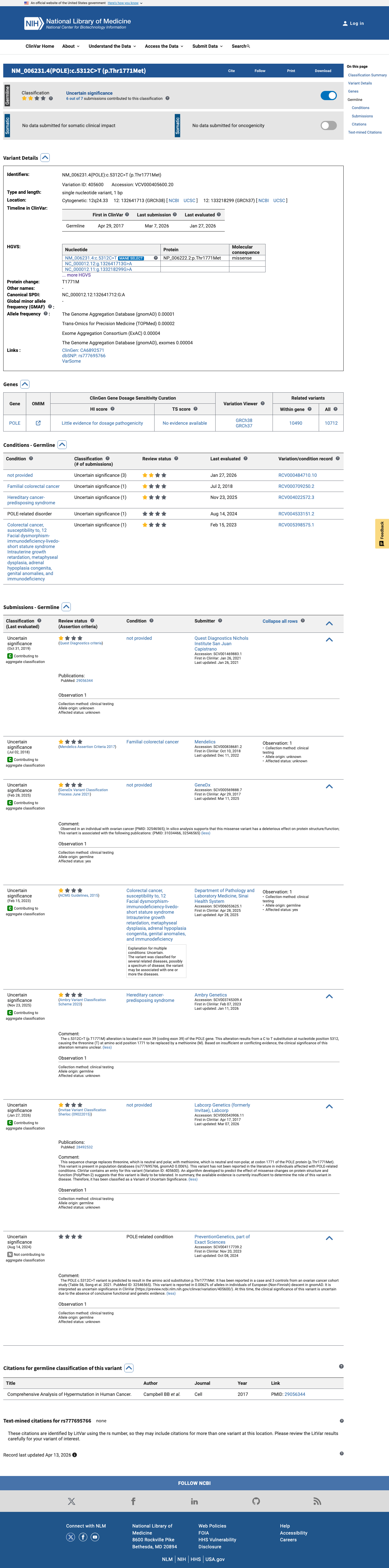
No validated variant-specific functional assay evidence was identified for p.(Thr1771Met).
oncokb ↗4
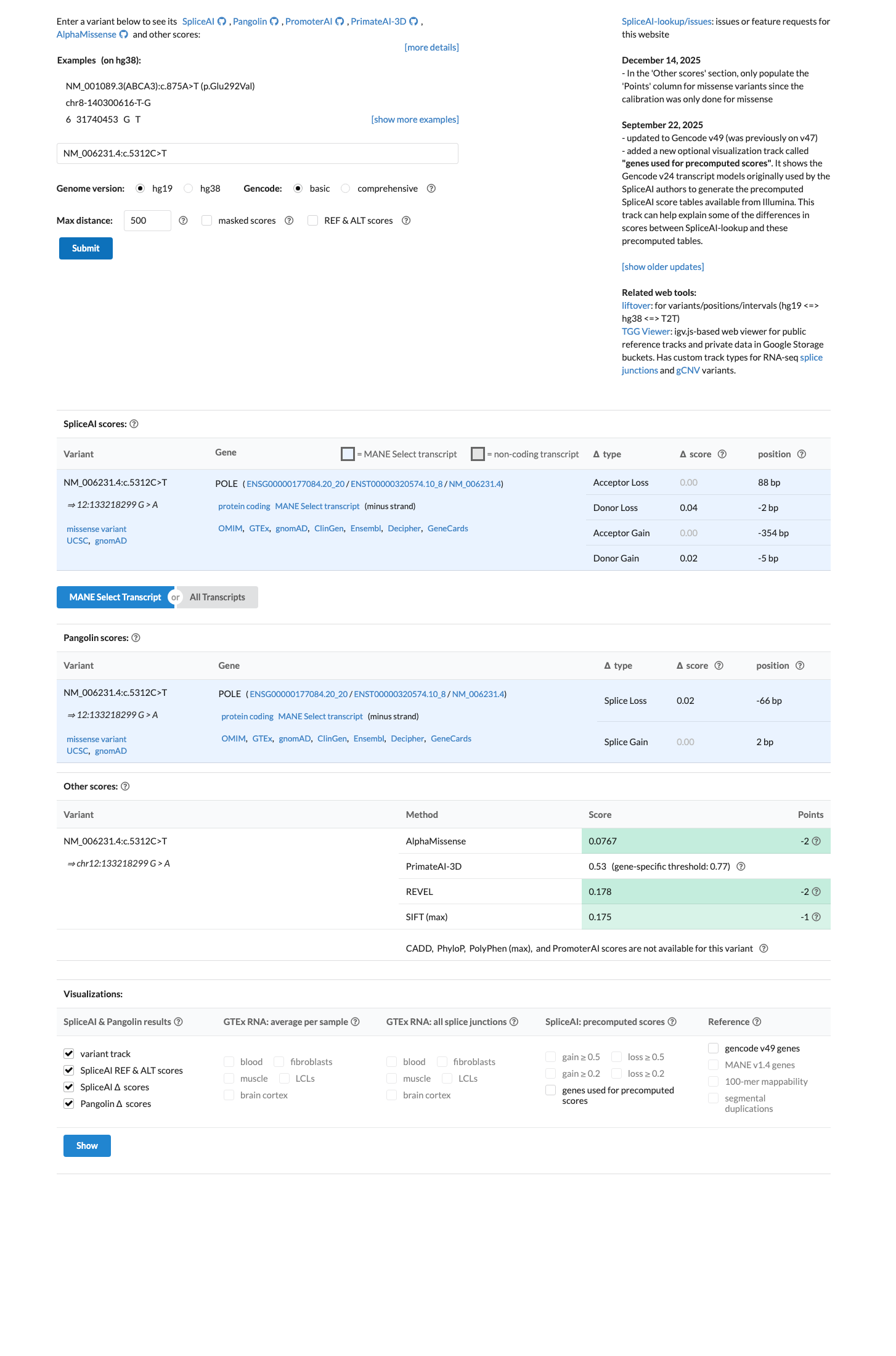
SpliceAI predicts no significant splice impact (maximum delta score 0.04), and the local REVEL score is 0.178; however, the variant is not represented in the León-Castillo POLE supplementary computational tables, so the custom POLE PP3 and BP4 rules were not applied.
spliceai ↗