Classification rationale
1
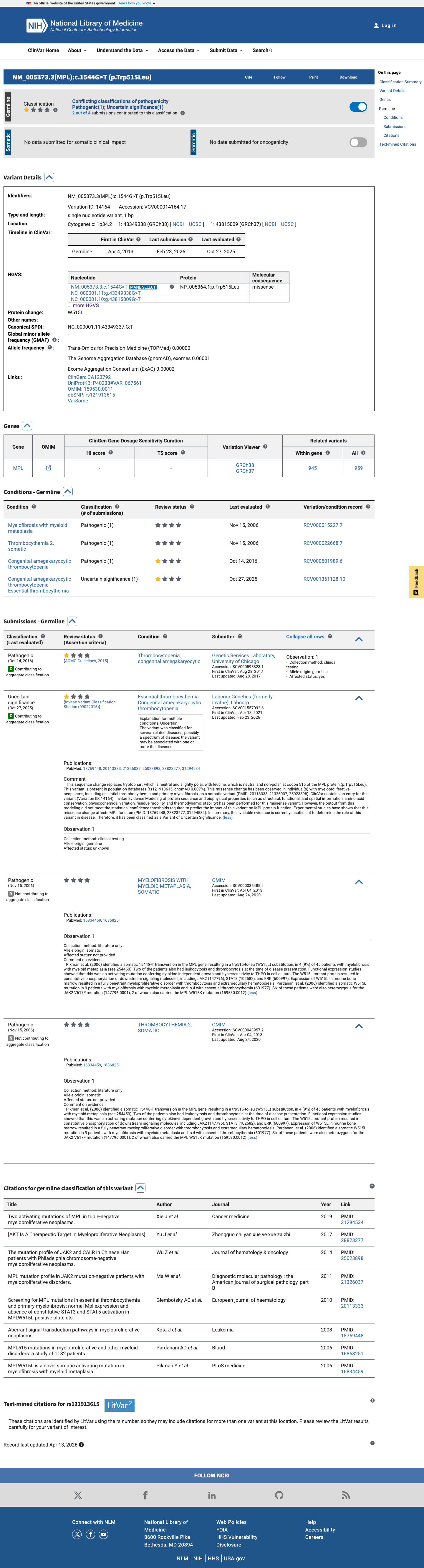
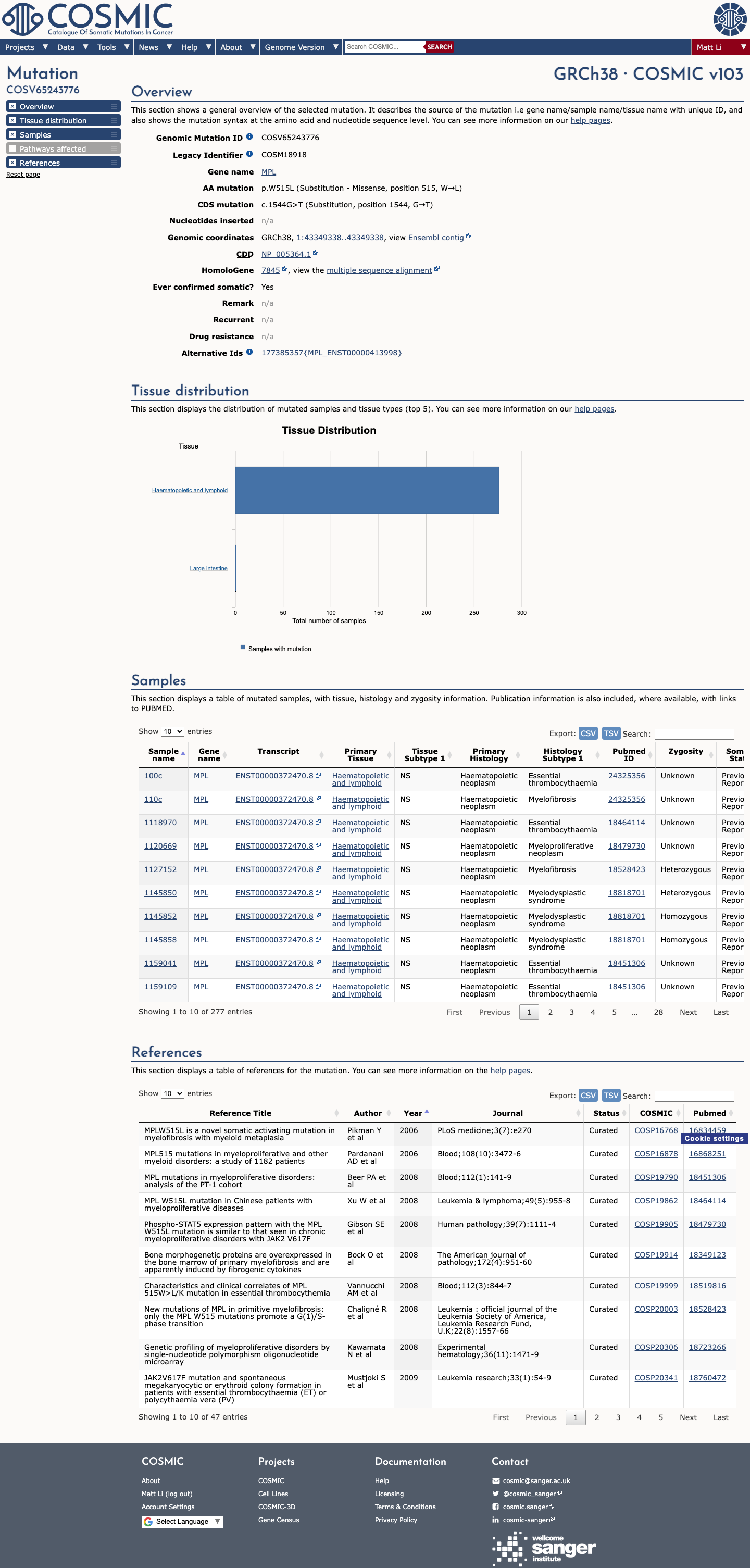
The MPL c.1544G>T (p.Trp515Leu; p.W515L) variant has been reported in somatic myeloproliferative neoplasms and is listed in ClinVar with one Pathogenic submission and one Uncertain significance submission.
clinvar ↗ PMID:16834459 ↗ PMID:16868251 ↗ PMID:20113333 ↗ PMID:21326037 ↗2
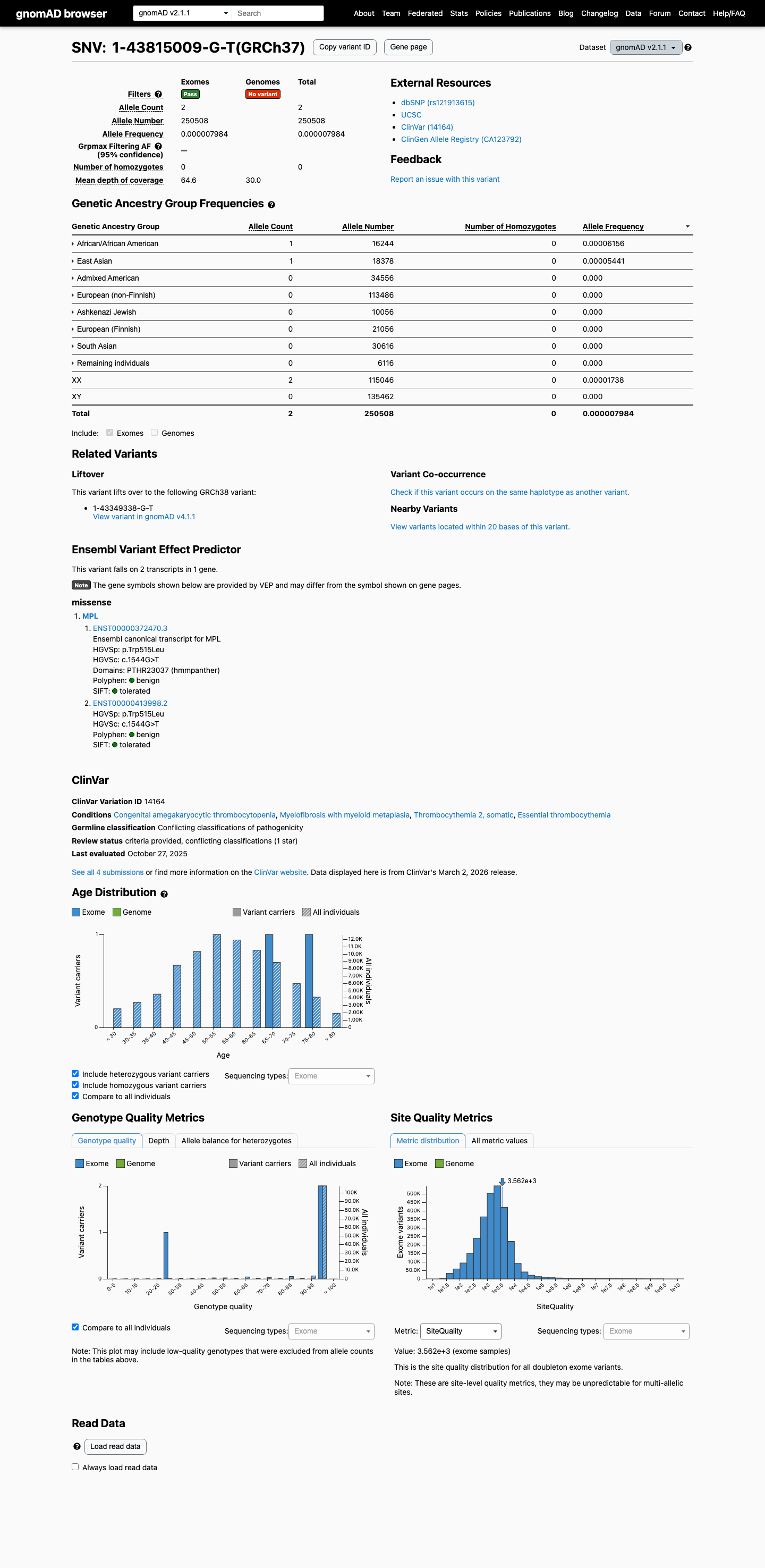
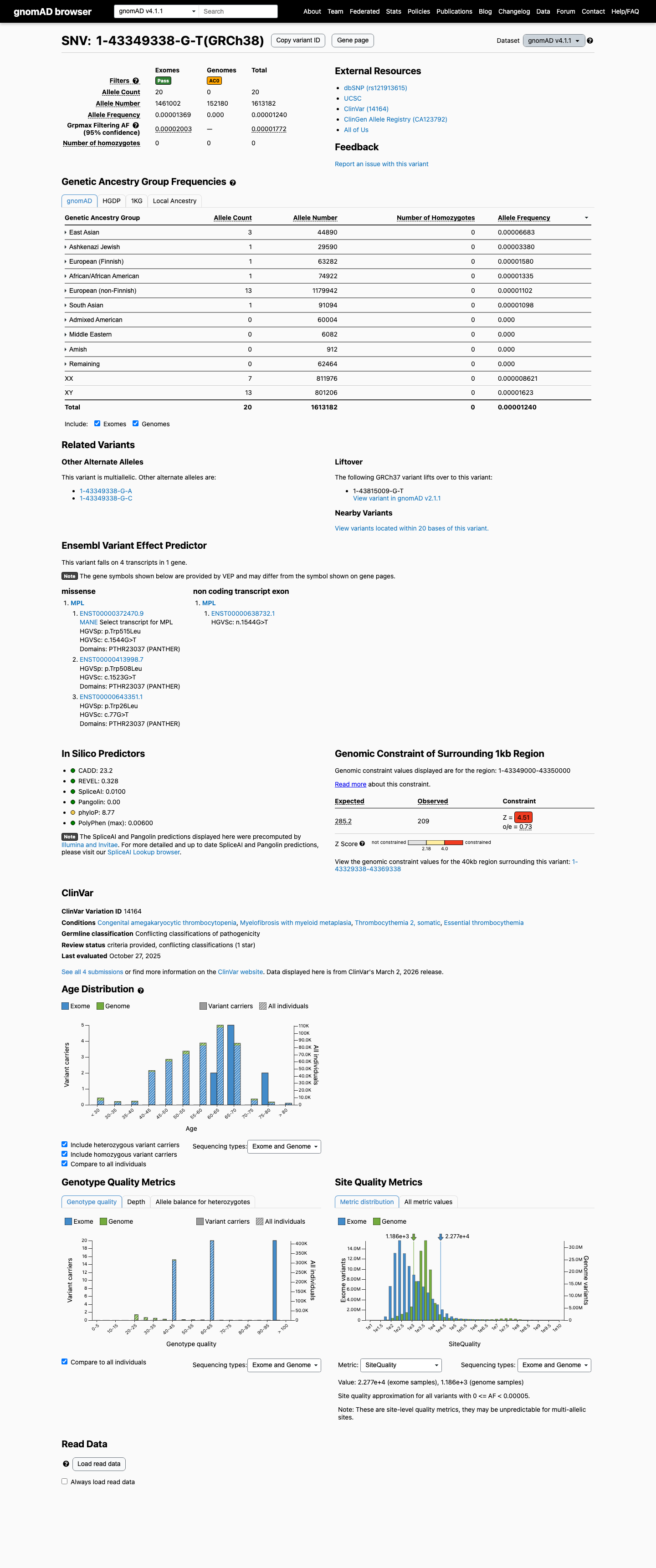
This variant is rare in population databases, with gnomAD v2.1 allele frequency 7.98378e-06 (0.00080%, 2/250508) and gnomAD v4.1 allele frequency 1.23979e-05 (0.00124%, 20/1613182), both below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
Available functional evidence supports an abnormal activating effect, as p.W515L is described as an activating MPL variant and OncoKB classifies it as oncogenic with gain-of-function activity.
oncokb ↗ PMID:16834459 ↗ PMID:28823277 ↗4
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.03.
spliceai ↗