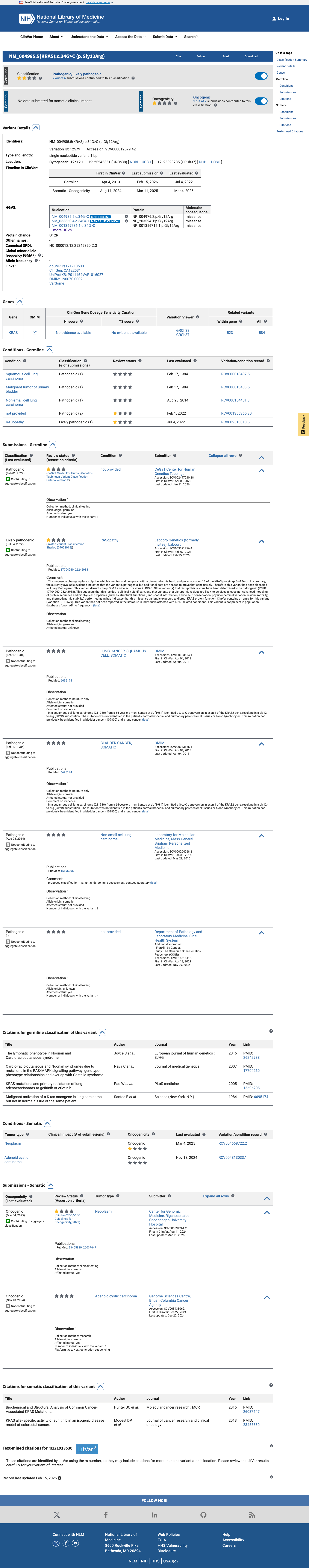
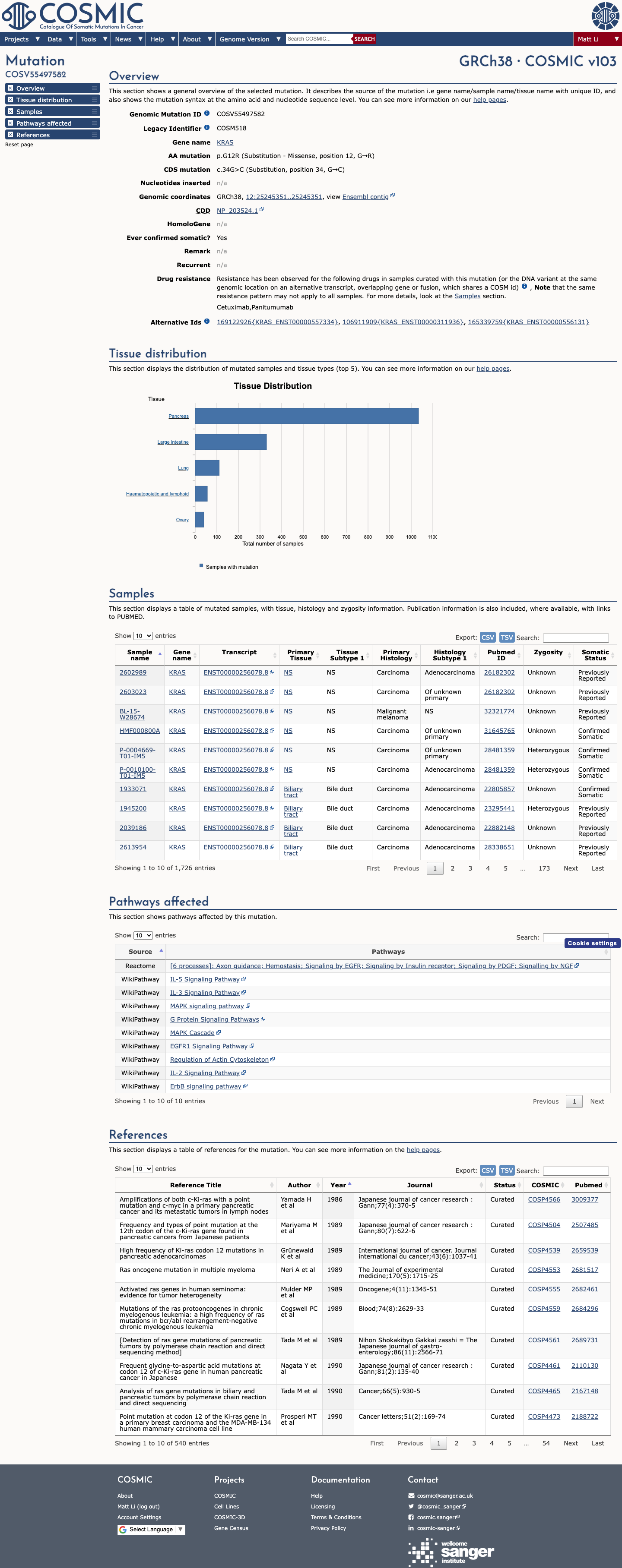
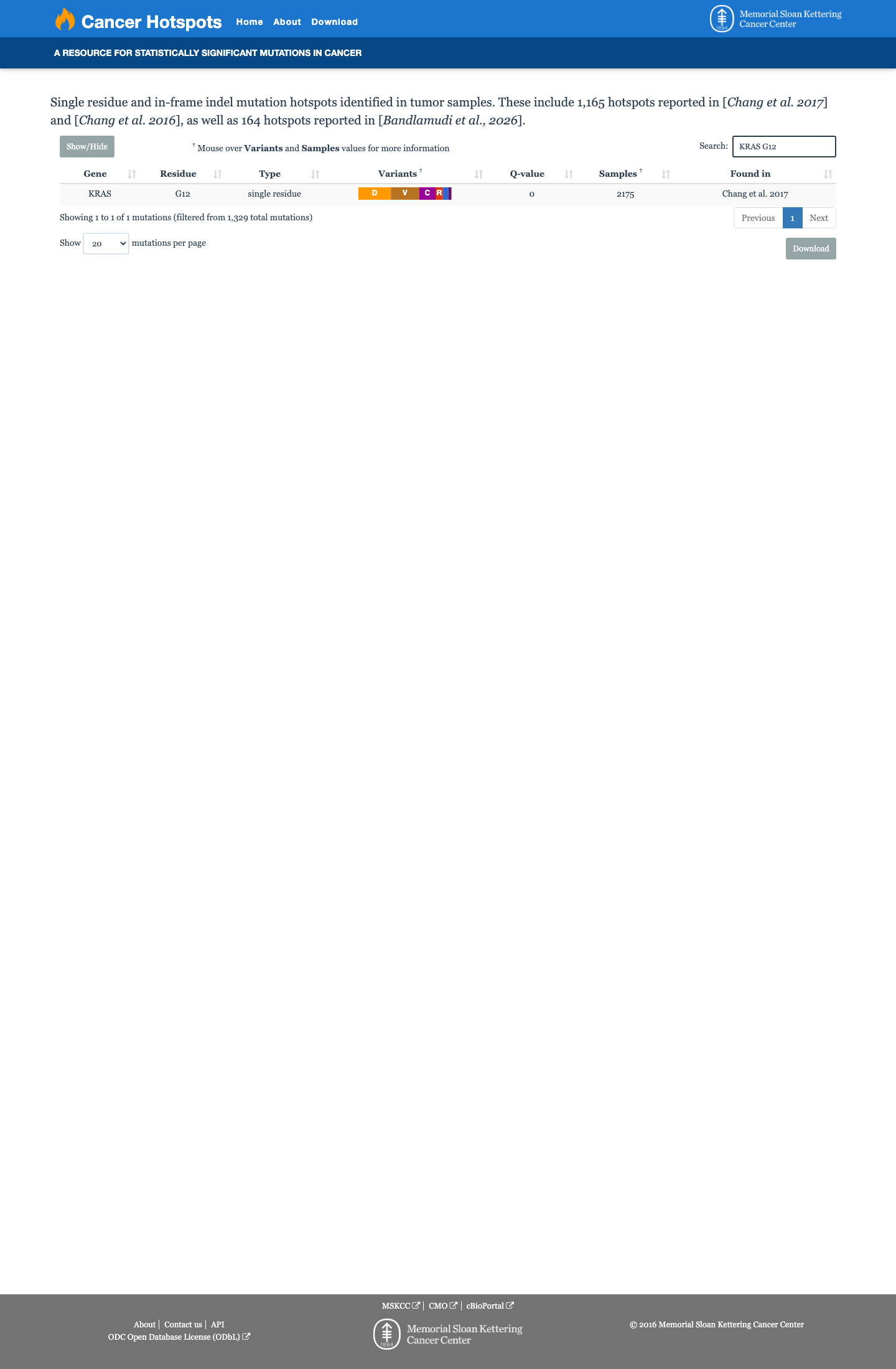
The KRAS c.34G>C (p.Gly12Arg, p.G12R) variant has been observed in somatic cancers in COSMIC (COSV55497582; n=1720) and has been reported in ClinVar with pathogenic and likely pathogenic clinical submissions.
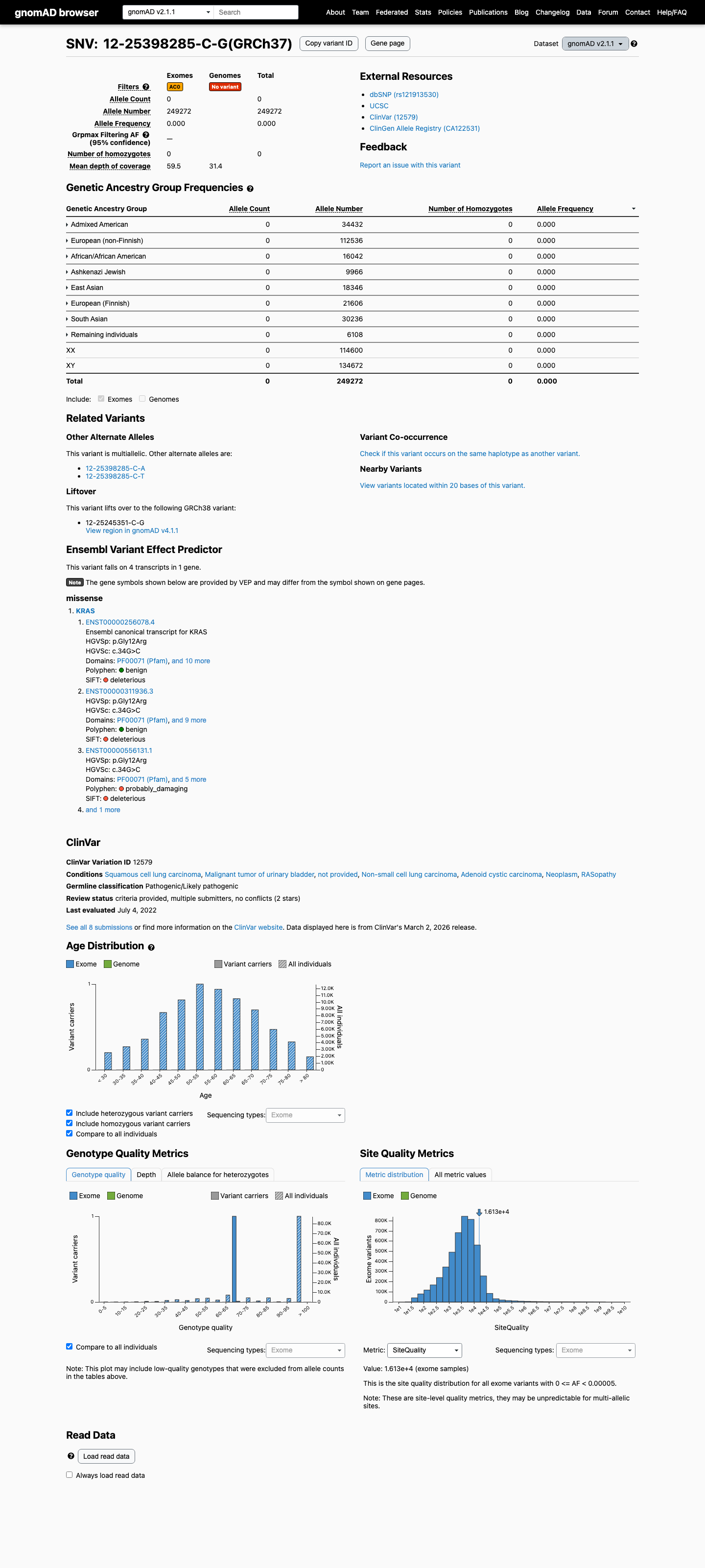
clinvar ↗This variant is absent from population controls, with 0/249272 alleles in gnomAD v2.1 and no observation in gnomAD v4.1, which supports PM2 and argues against BA1 and BS1.
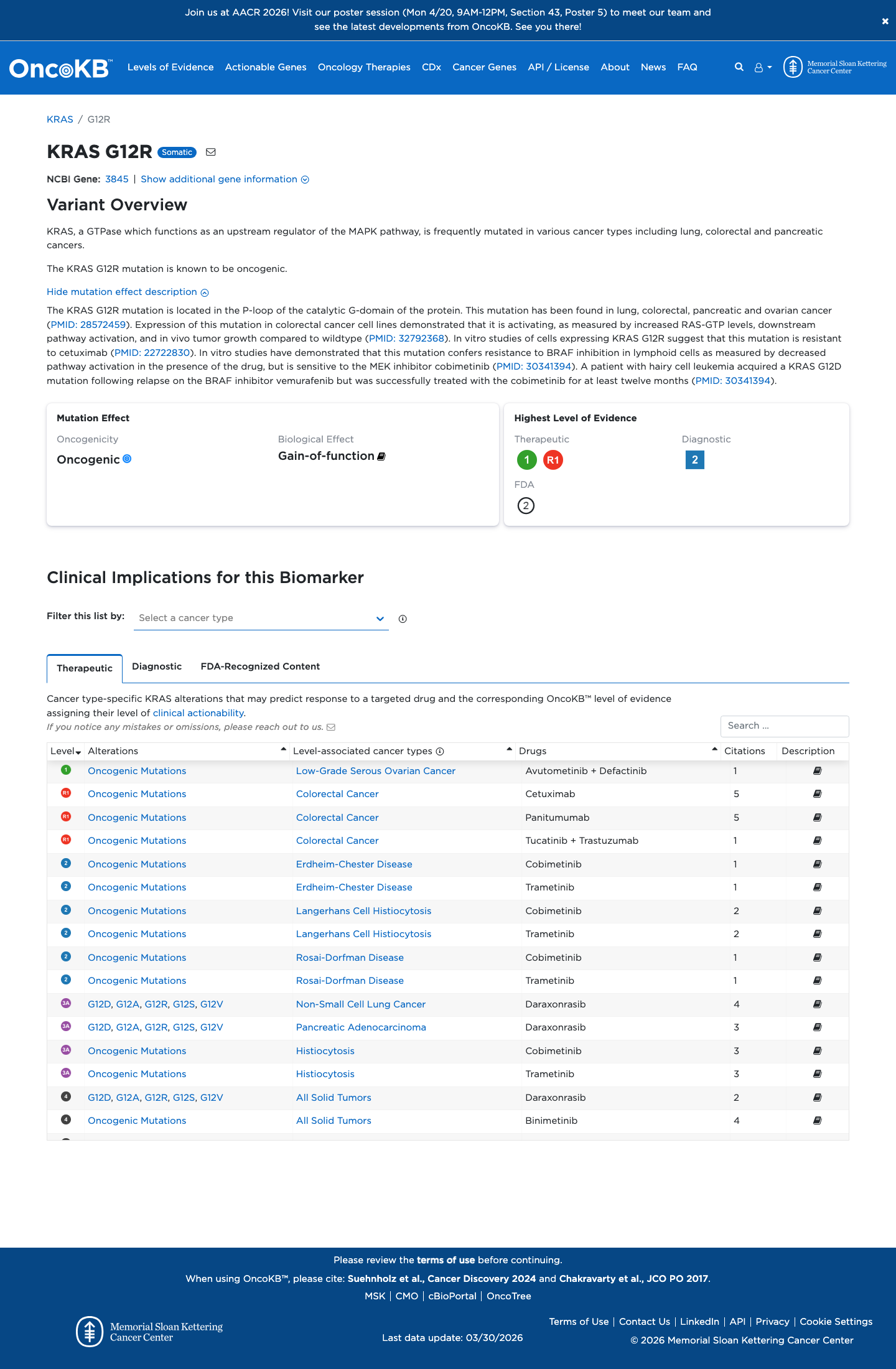
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Available functional evidence is consistent with an activating KRAS effect, and OncoKB classifies this variant as oncogenic with gain-of-function, but the curated RASopathy VCEP materials did not provide sufficient approved variant-specific assay evidence to apply PS3.
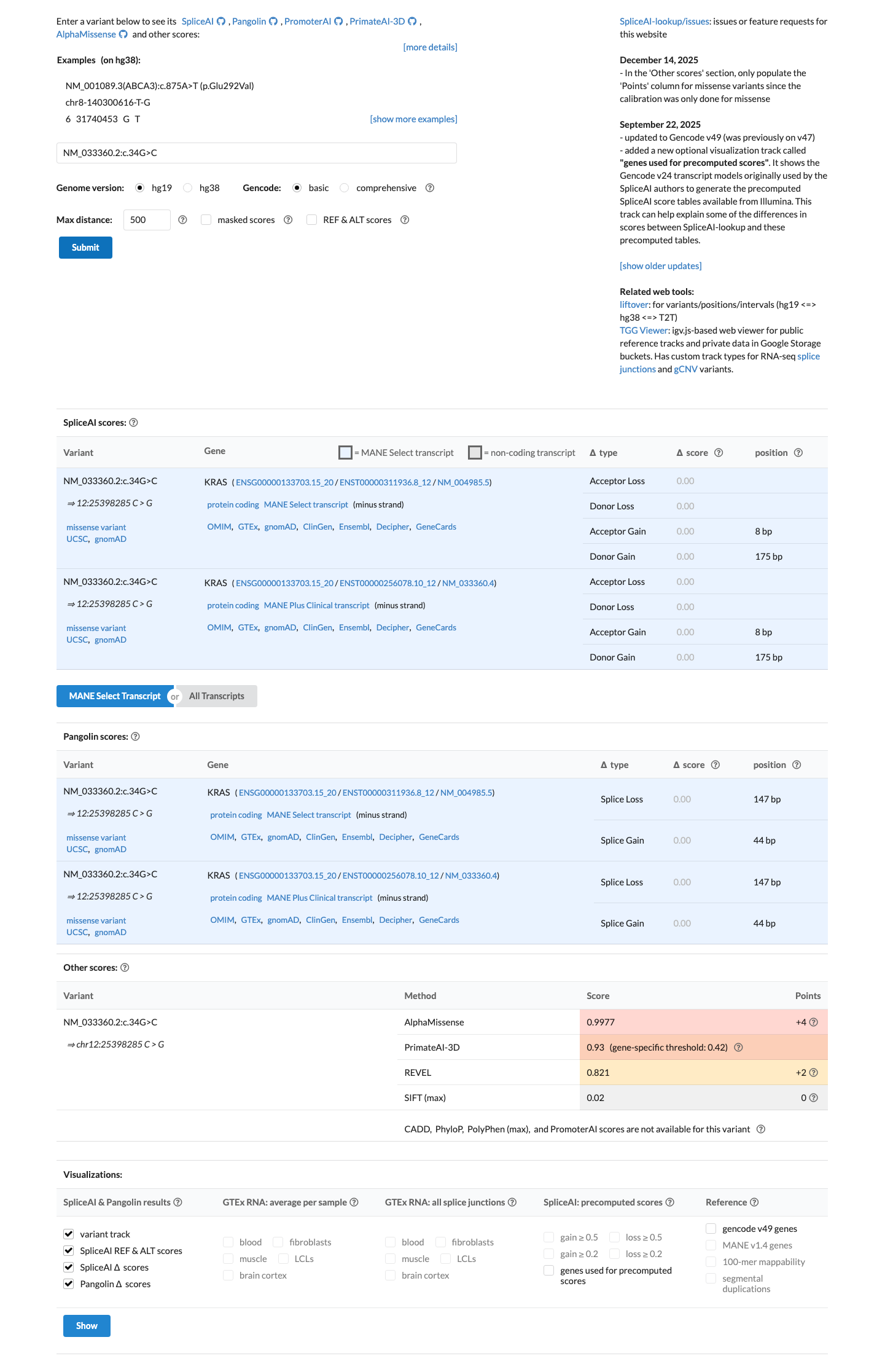
oncokb ↗ PMID:23455880 ↗ PMID:26037647 ↗ PMID:32792368 ↗This missense change affects codon 12 within the KRAS P-loop domain, and computational data are consistent with a damaging missense effect, with REVEL 0.821 and no predicted splice disruption by SpliceAI (maximum delta score 0.00).
cspec ↗ spliceai ↗