Classification rationale
1
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, so the observed allele frequency is 0 in the queried population datasets.
gnomad_v2 ↗ gnomad_v4 ↗3
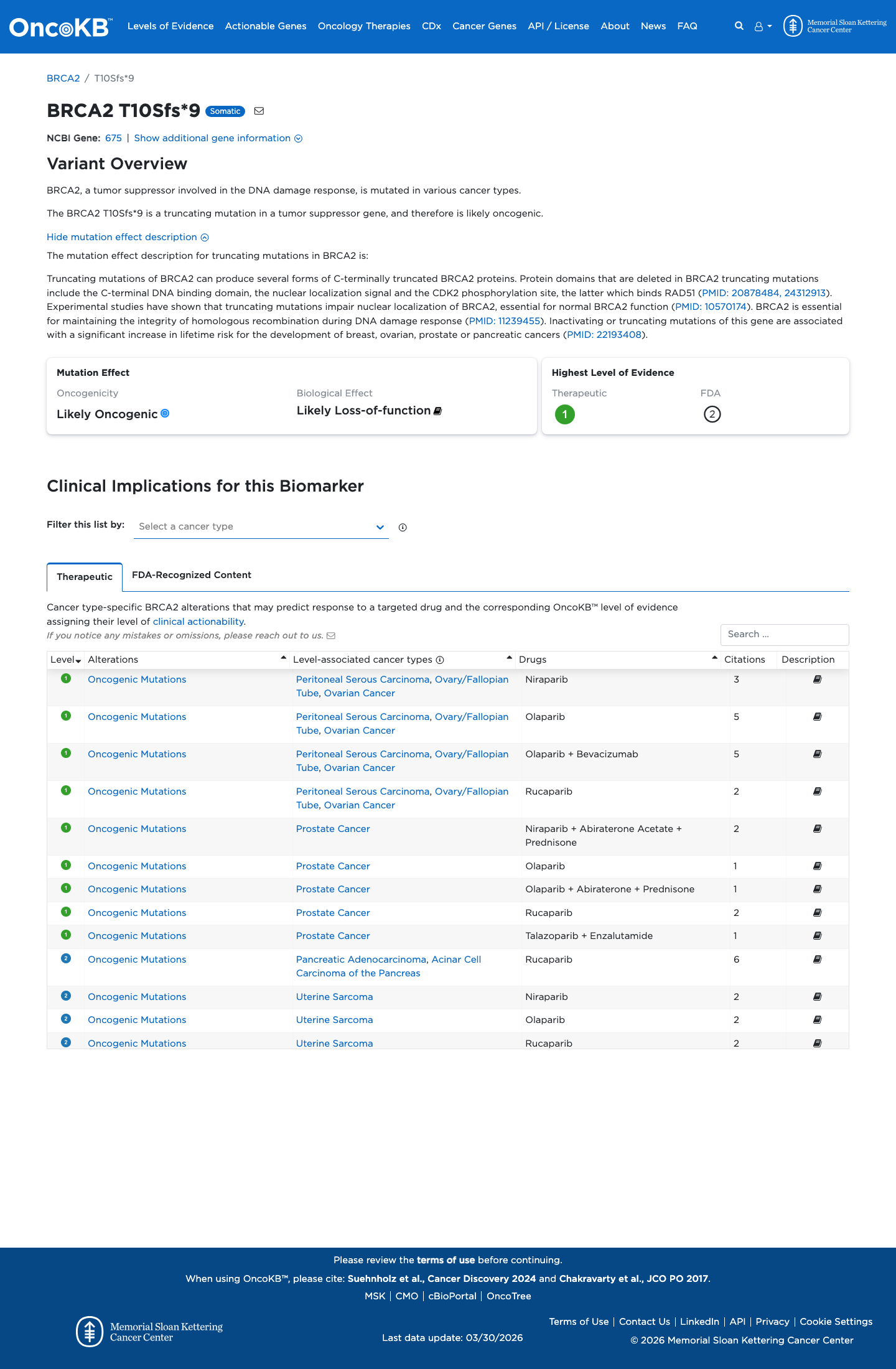
The deletion causes an early frameshift with premature termination in exon 2, and ENIGMA BRCA2 Table 4 assigns exon 2 protein-truncating variants PVS1 and PM5_Strong (PTC), which supports a loss-of-function disease mechanism for this variant.
cspec ↗4
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.04.
spliceai ↗