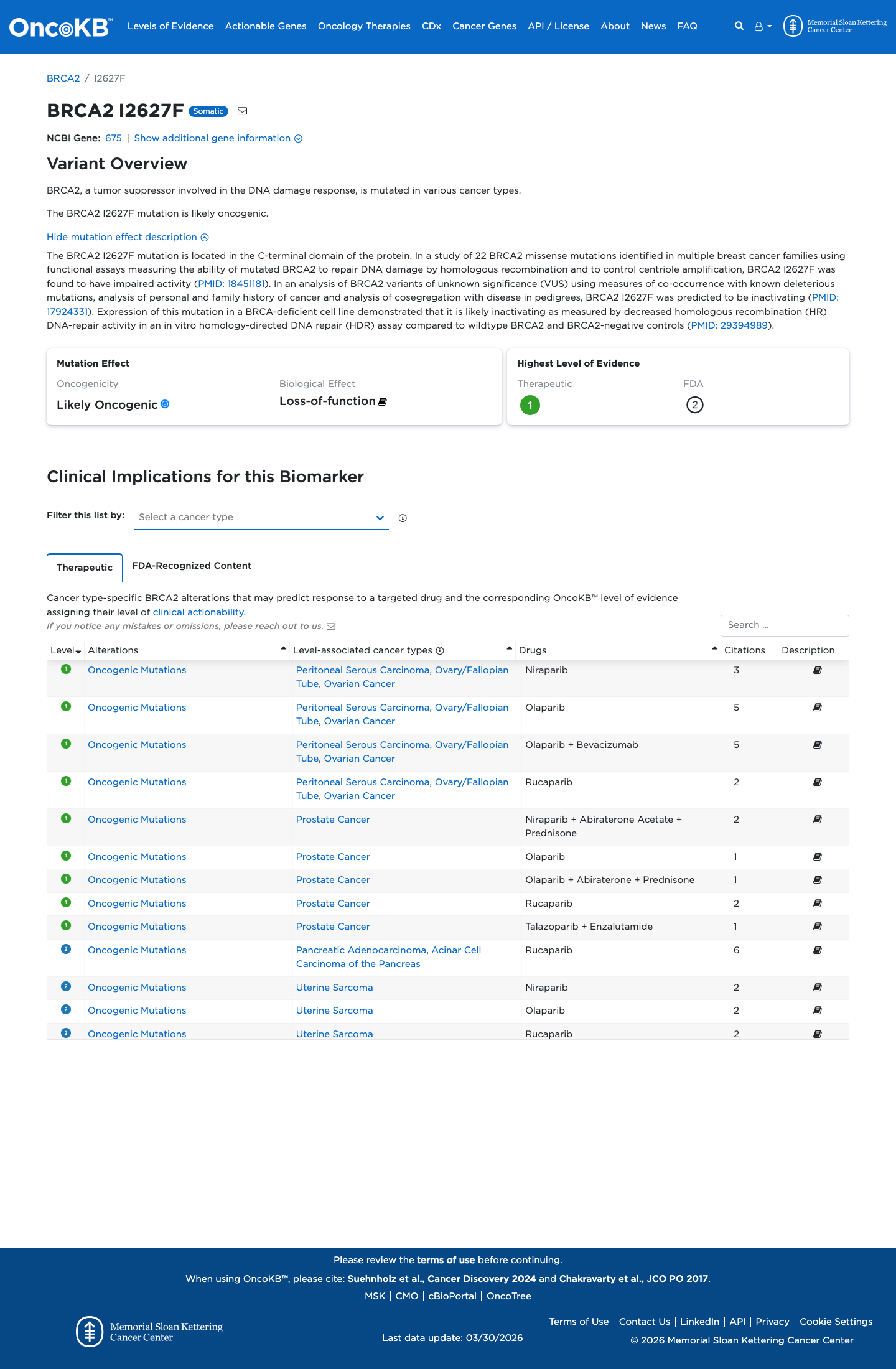
The BRCA2 c.7879A>T (p.Ile2627Phe) variant has been observed in somatic cancer once in COSMIC and has been reported in ClinVar as Pathogenic, including review by the ClinGen ENIGMA BRCA1/2 expert panel.
clinvar ↗This variant is absent from gnomAD v2.1 and present at very low frequency in gnomAD v4.1 at 4/1614168 alleles (AF 2.47806e-06; 0 homozygotes; grpmax FAF 7.9e-07), which is well below benign population thresholds but does not satisfy ENIGMA absence-based PM2.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Calibrated functional evidence supports a damaging effect on BRCA2 protein function, and ENIGMA Table 9 assigns PS3 Strong for c.7879A>T (p.Ile2627Phe).
The variant lies within the BRCA2 DNA-binding domain, while SpliceAI predicts no significant splice effect with a maximum delta score of 0.07.
cspec ↗ spliceai ↗