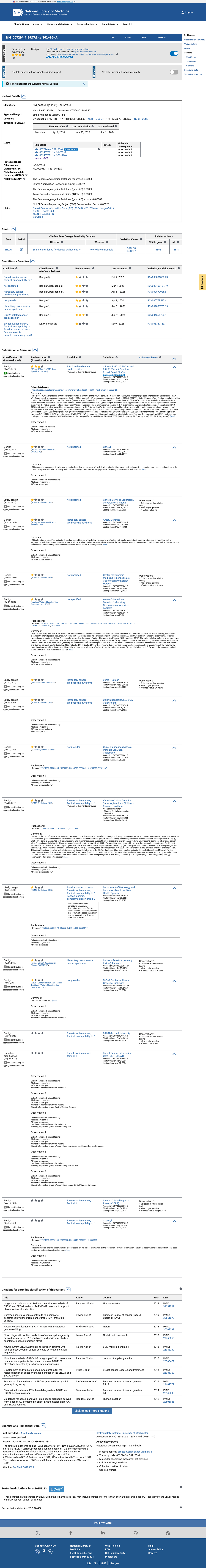
The BRCA1 c.301+7G>A (NP_009225.1:p.?) variant has been reported in ClinVar, where the overall classification is Benign with expert panel review.
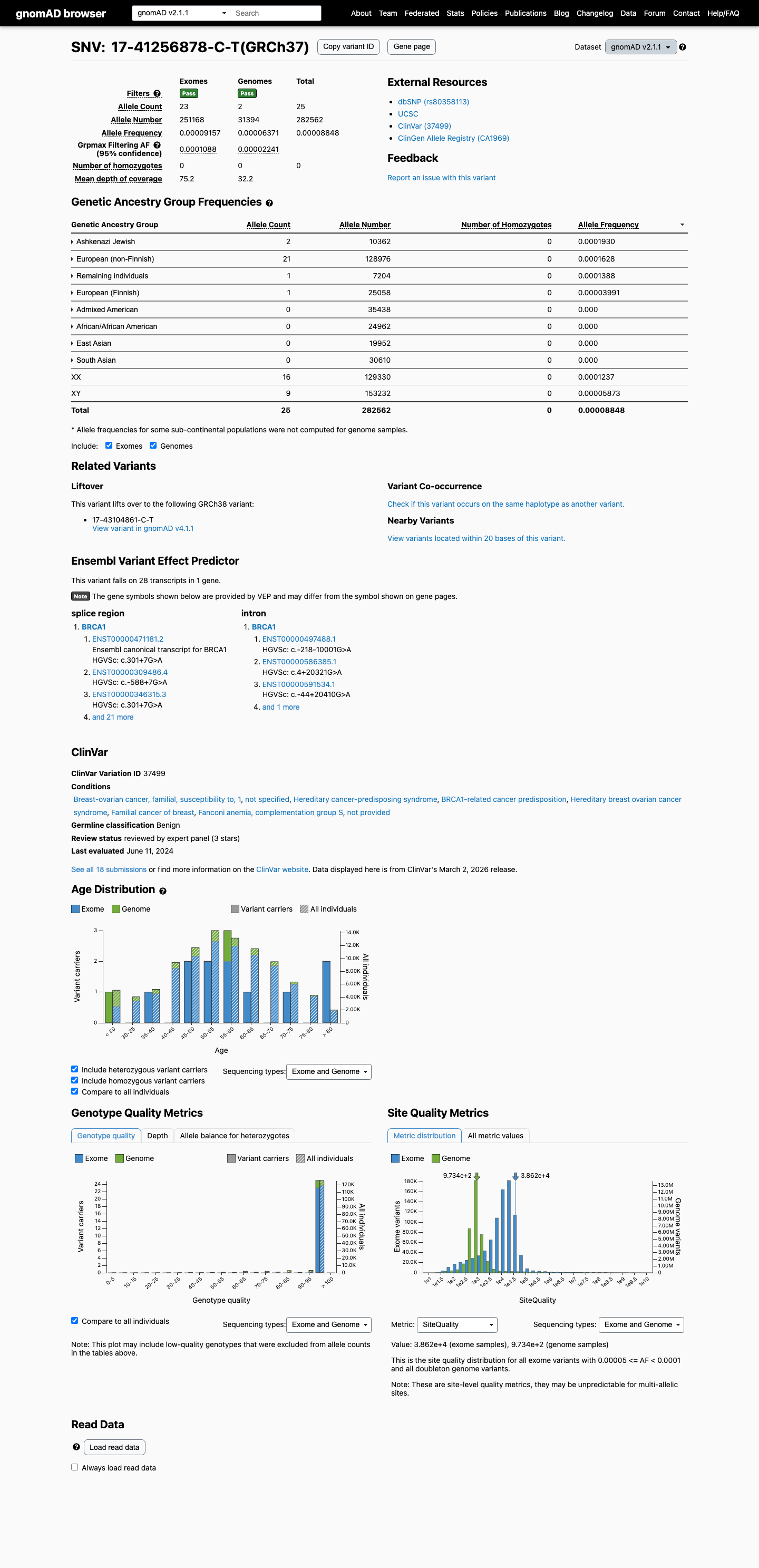
clinvar ↗This variant is present in gnomAD, with AF 0.0000885 in v2.1 (25/282562 alleles; grpmax FAF 0.00010876) and AF 0.0000540 in v4.1 (87/1610834 alleles; grpmax FAF 0.00005311), so it is not absent from controls and does not meet BA1.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Well-established functional evidence supports no damaging effect: ENIGMA Table 9 assigns BS3 Strong based on no aberrant splicing in two studies and a calibrated study showing no functional impact.
PMID:22505045 ↗ PMID:24667779 ↗In silico splicing prediction is conflicting with the functional evidence, because SpliceAI gives a maximum delta score of 0.29, which exceeds the ENIGMA PP3 threshold of 0.20 and argues against BP4.
spliceai ↗ cspec ↗