The PTEN c.434T>G (p.Phe145Cys; p.F145C) variant has been observed once in somatic cancers in COSMIC and has not been reported in ClinVar.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the PTEN PM2 threshold of 0.001% and supports PM2 at Supporting strength.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In the PTEN saturation mutagenesis phosphatase assay, p.Phe145Cys had a cumulative activity score of -0.669, which is above the PTEN PS3_Moderate cutoff of <= -1.11 and below the BS3 threshold of >0, so the current functional evidence does not independently meet PS3 or BS3.
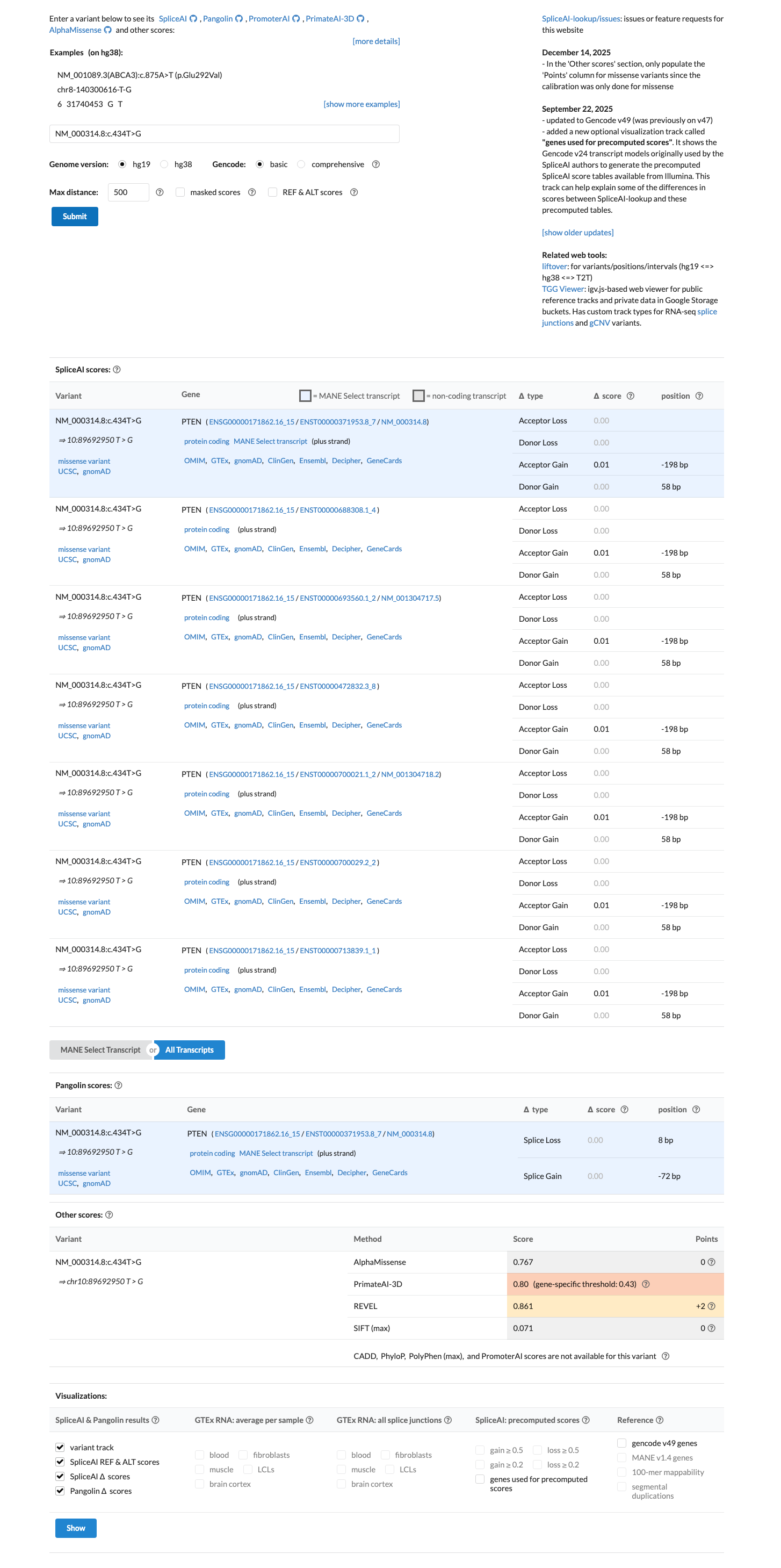
cspec ↗Computational evidence supports a deleterious protein effect because the REVEL score is 0.861, above the PTEN PP3 threshold of >0.7, while SpliceAI predicts no significant splice effect with a maximum delta score of 0.01.
cspec ↗ spliceai ↗