Classification rationale
1
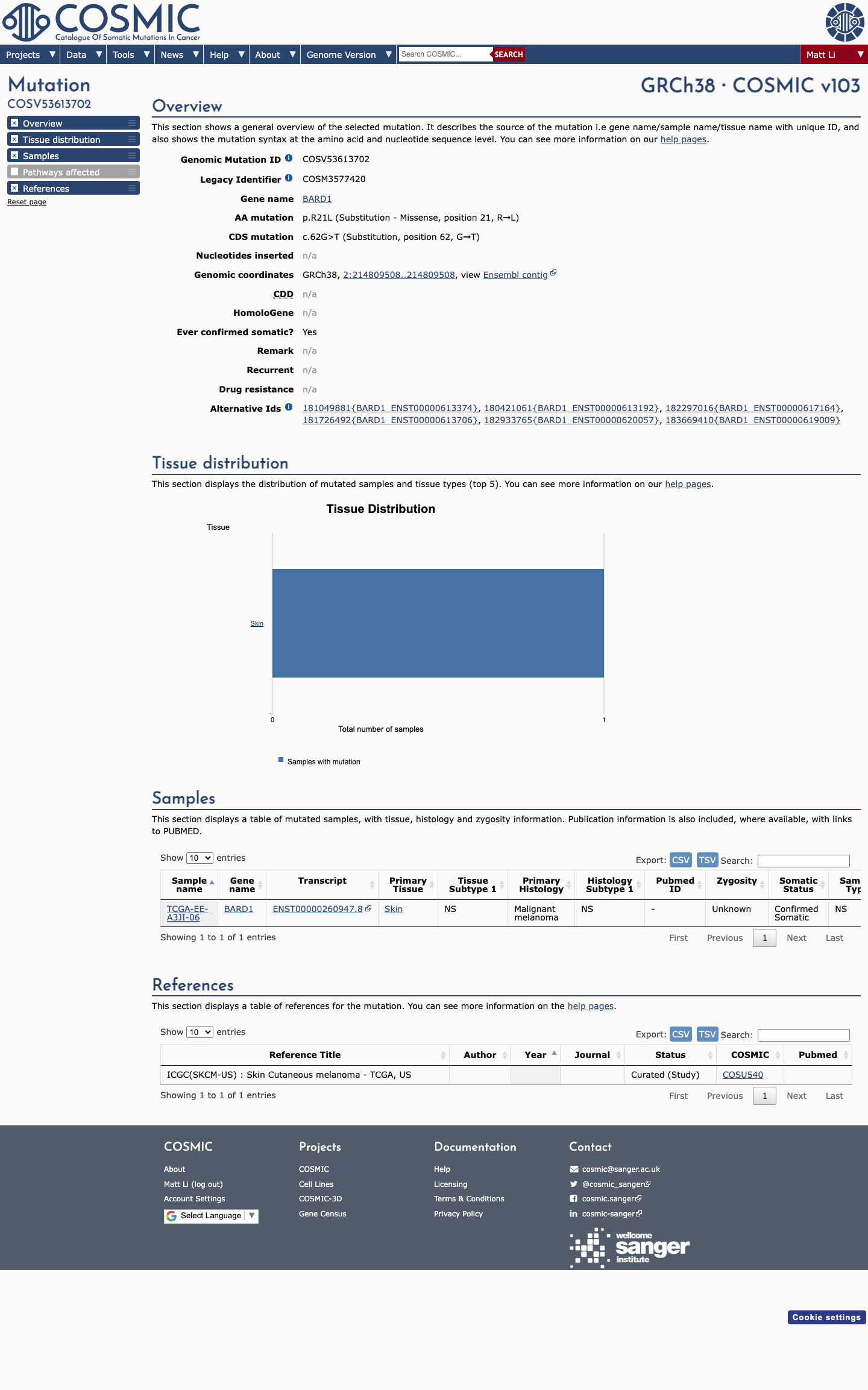
The BARD1 c.62G>T (p.Arg21Leu) variant has not been identified in a statistically significant cancer hotspot and has been reported in ClinVar as a variant of uncertain significance by a single submitter.
hotspots ↗ clinvar ↗2
This variant is absent from gnomAD v2.1 and has 0/1,601,354 alleles in gnomAD v4.1 (AF 0.0%), which is below the 0.1% threshold used for PM2 and supports rarity in population databases.
gnomad_v2 ↗ gnomad_v4 ↗3
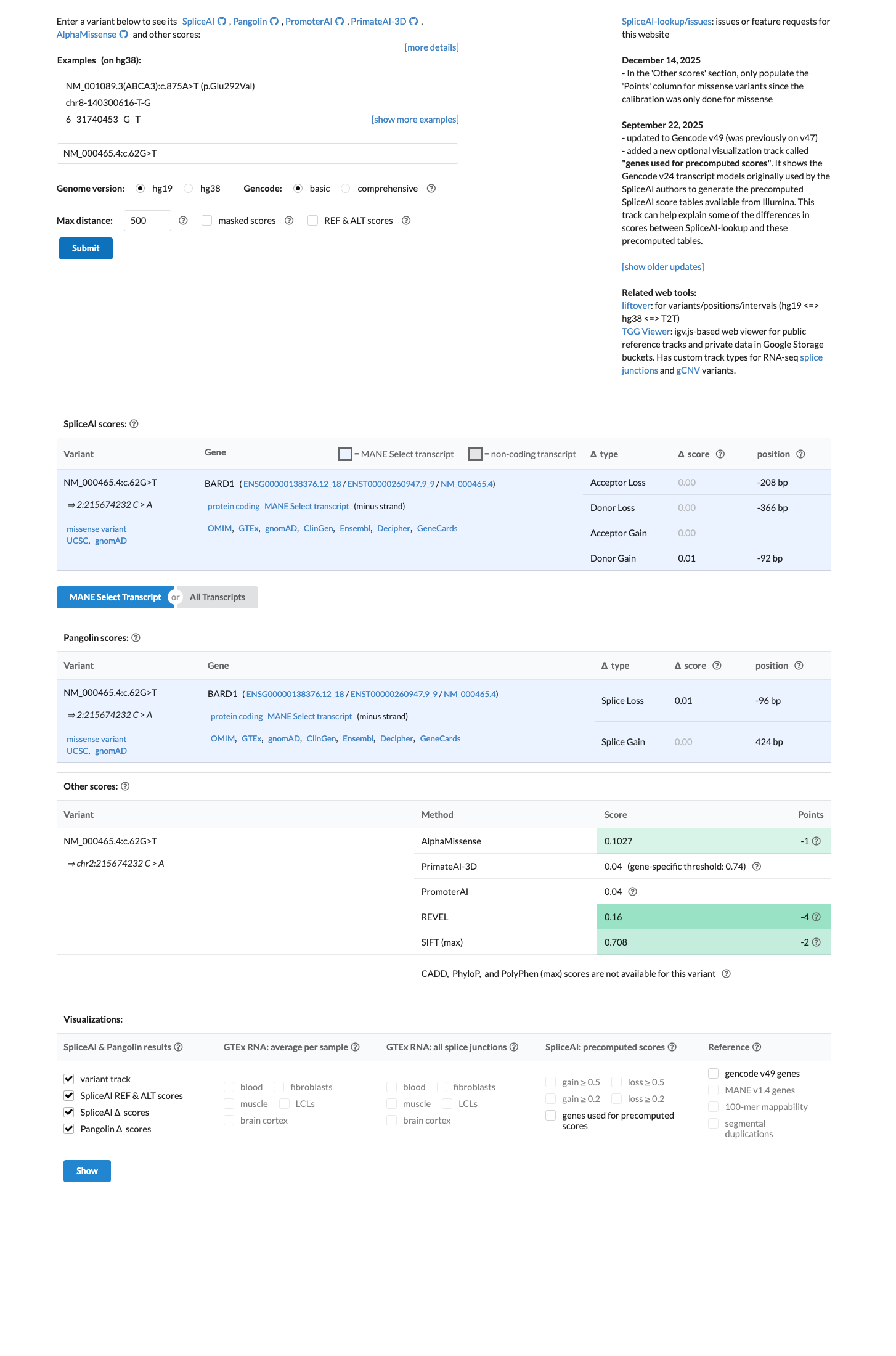
Available computational evidence does not support a damaging effect, with a REVEL score of 0.16 and a BayesDel score of -0.392235, supporting benign computational evidence rather than pathogenic computational evidence.