Classification rationale
1
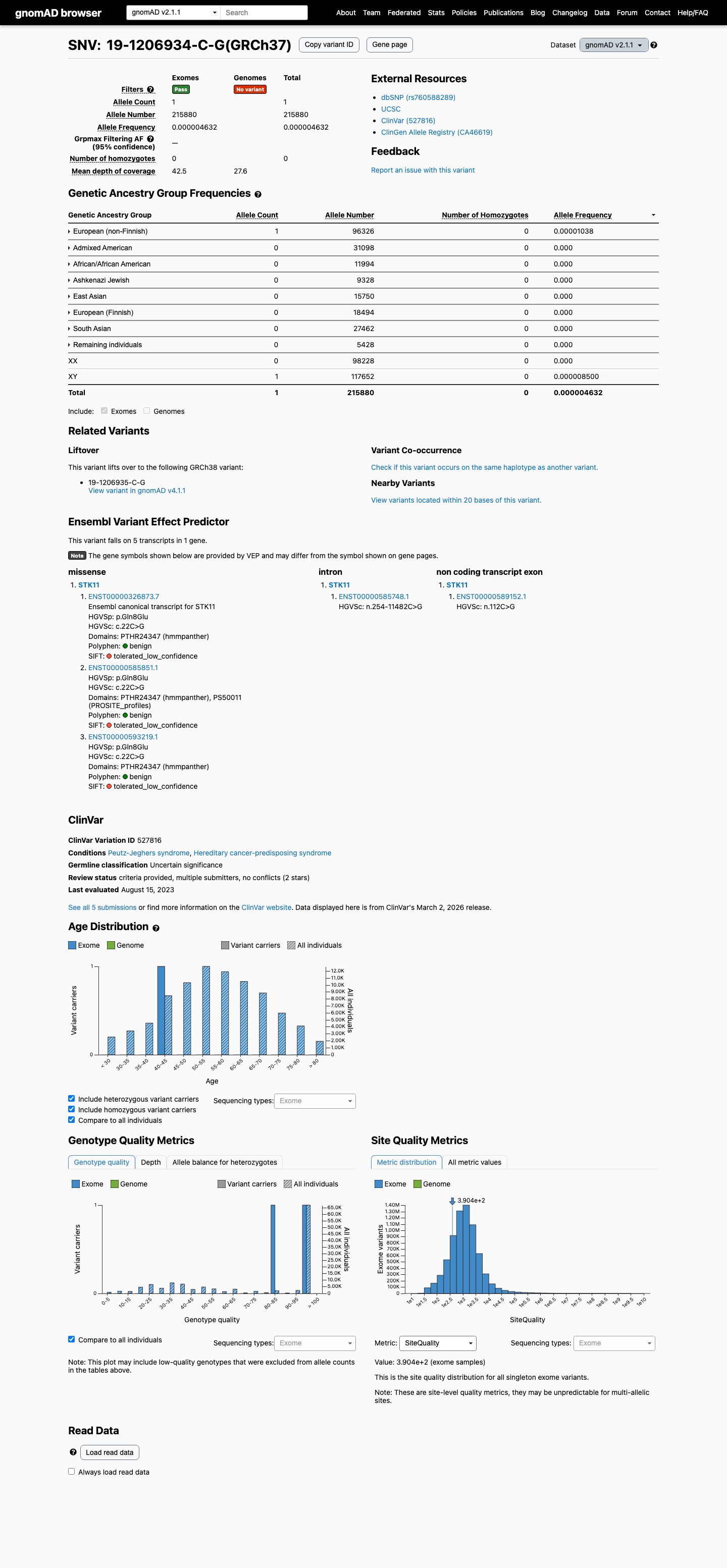
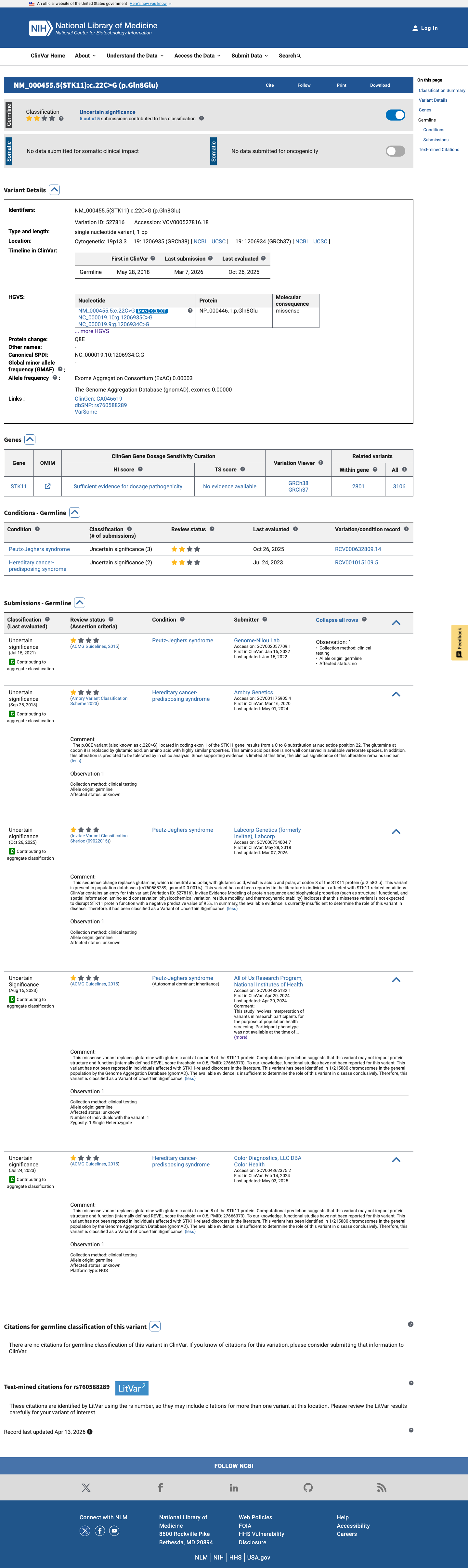
The STK11 c.22C>G (p.Gln8Glu) variant has been reported in ClinVar as uncertain significance by five clinical laboratory submissions.
clinvar ↗2
This variant is very rare in population databases, with AF 4.6322e-06 in gnomAD v2.1 and AF 2.50785e-06 in gnomAD v4.1, supporting rarity but not a benign frequency threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
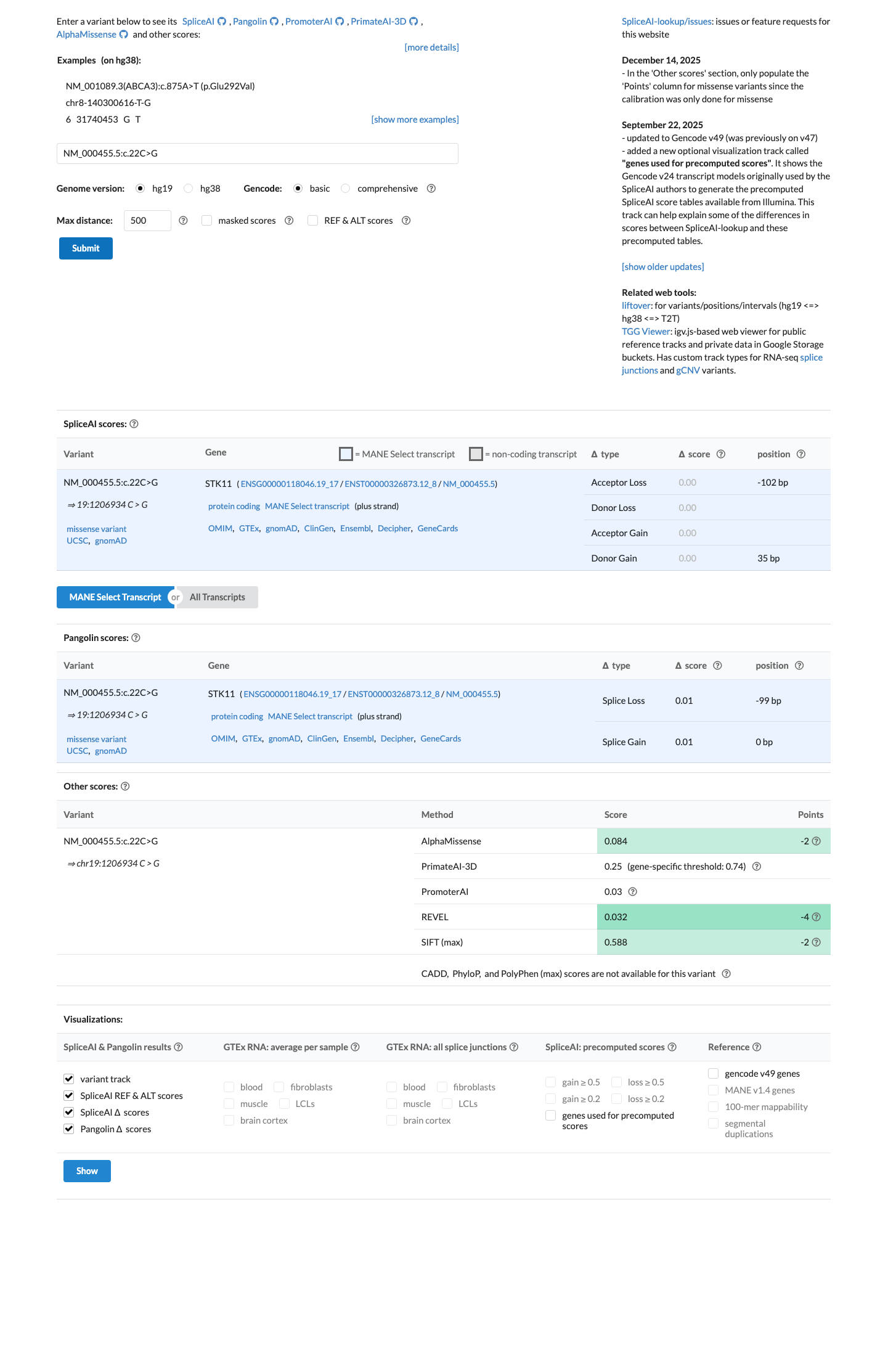
Computational evidence supports no significant impact on splicing or protein function, with SpliceAI max delta score 0.00, REVEL 0.032, and BayesDel -0.435498.
spliceai ↗