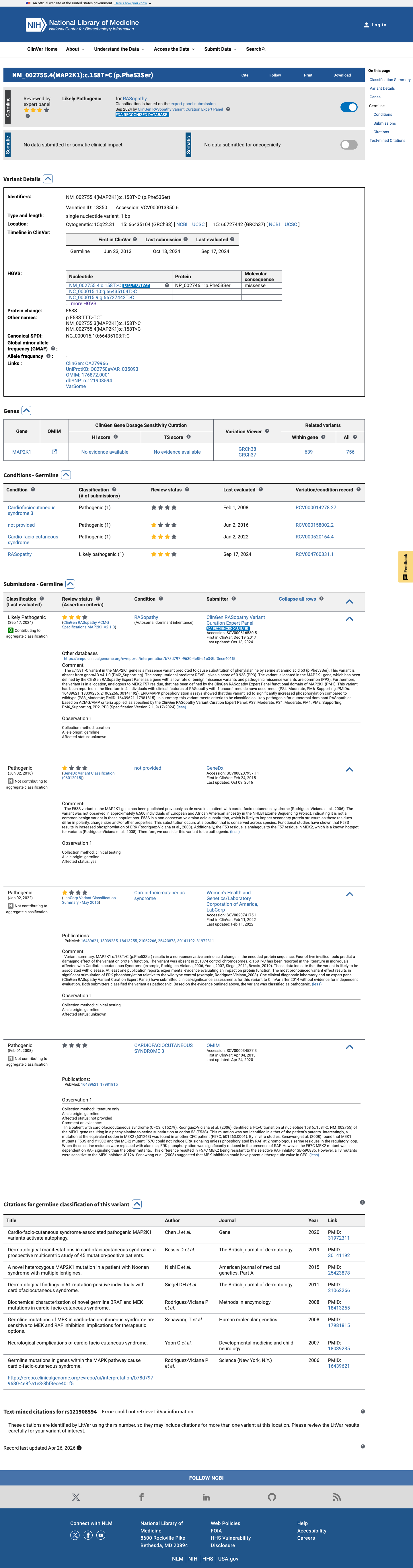
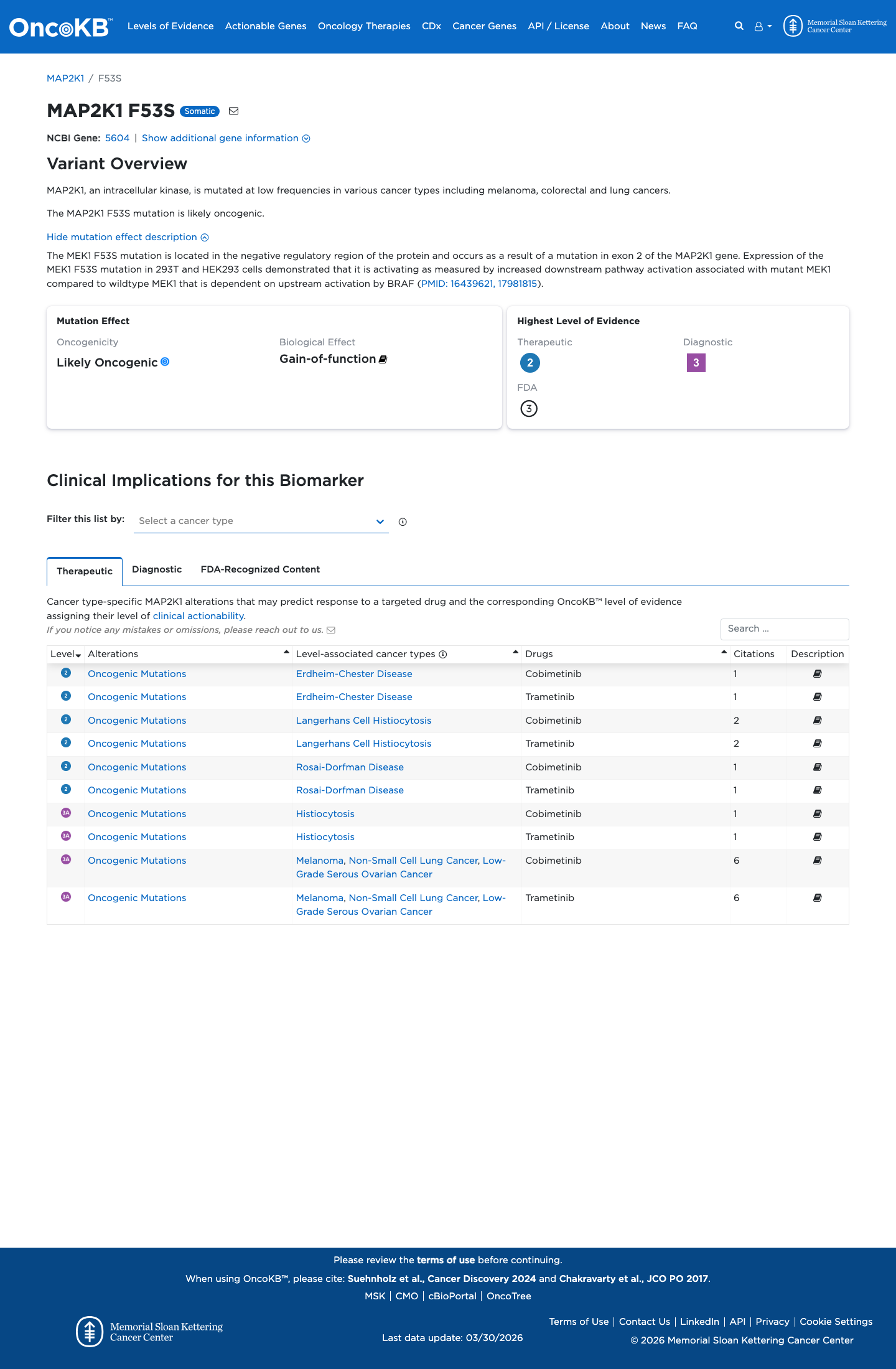
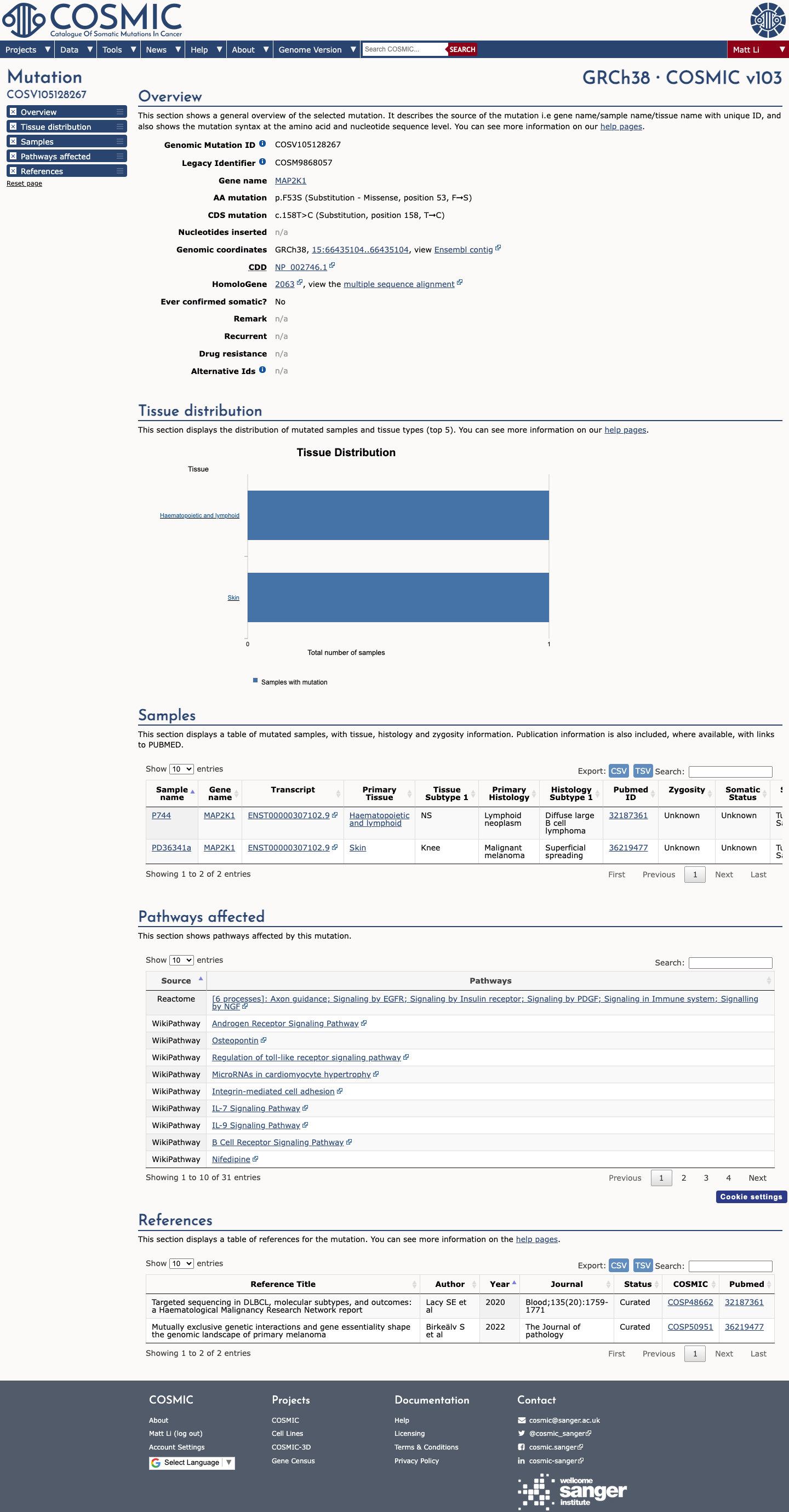
The MAP2K1 c.158T>C (p.Phe53Ser) variant has been observed in somatic cancers in COSMIC and has been reported in ClinVar, including an expert panel submission classifying it as likely pathogenic.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population reference datasets.
gnomad_v2 ↗ gnomad_v4 ↗Published functional studies of cardio-facio-cutaneous syndrome-associated MAP2K1 variants including F53S showed abnormal MAPK-pathway signaling, and the RASopathy VCEP recognizes MEK and ERK activation assays as approved functional assay types, supporting a damaging gain-of-function effect.
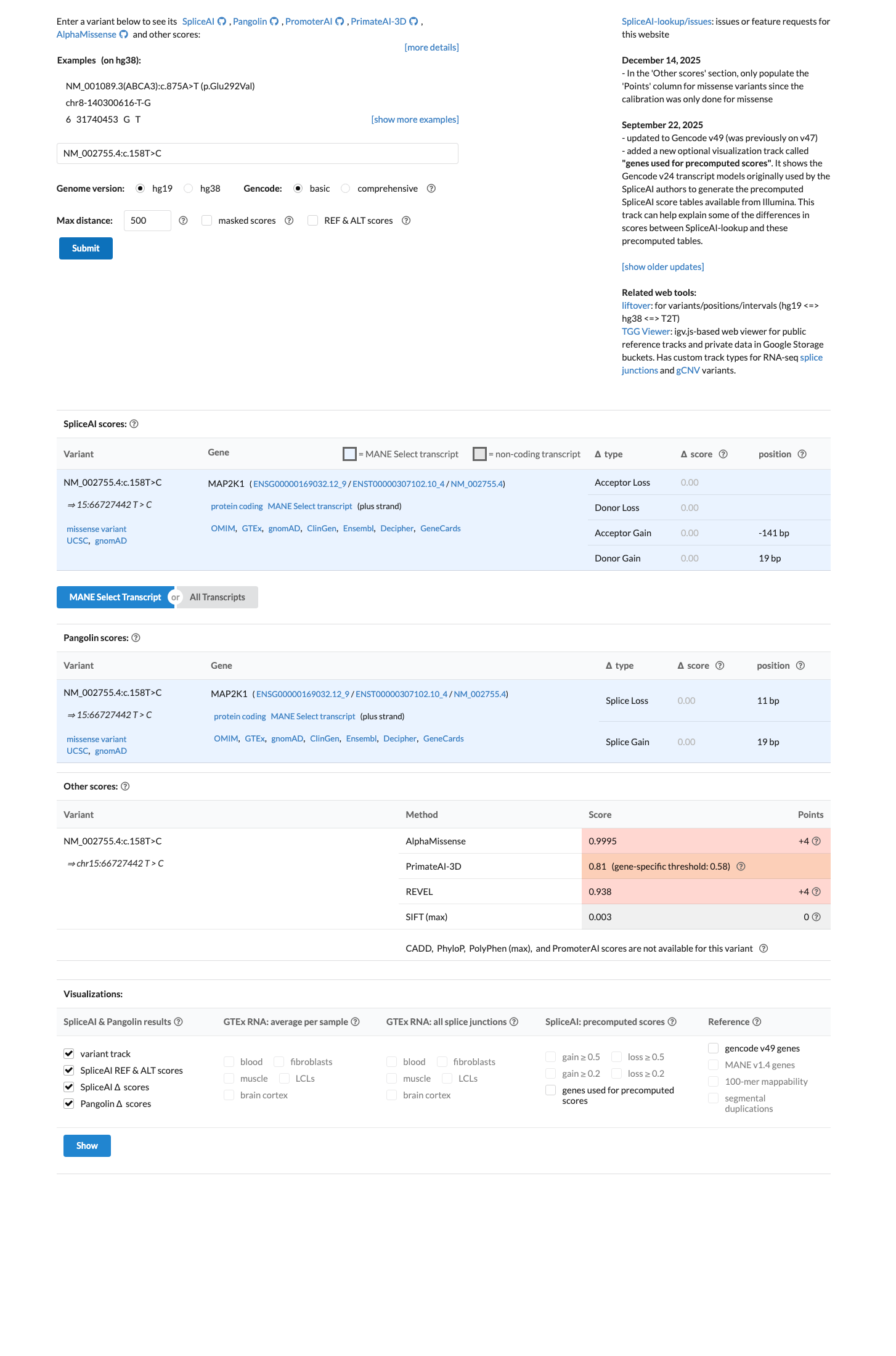
PMID:17981815 ↗ PMID:18413255 ↗Computational findings support a deleterious missense effect, with REVEL 0.938 above the VCEP PP3 threshold of 0.7, BayesDel 0.464442 in a damaging direction, and no predicted splice alteration by SpliceAI with a maximum delta score of 0.00.
spliceai ↗ cspec ↗