Classification rationale
1
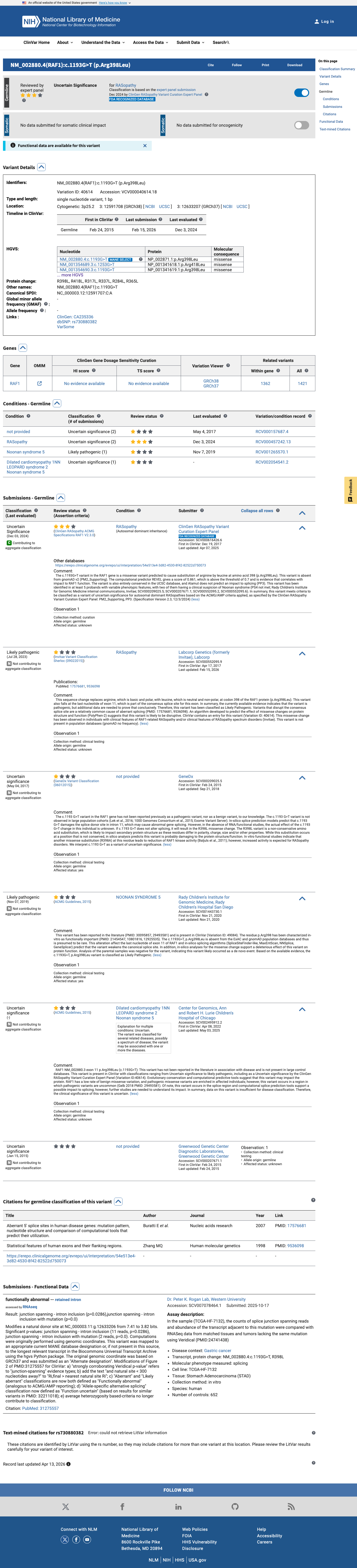
The RAF1 c.1193G>T (p.Arg398Leu) variant has been reported in ClinVar, where the ClinGen RASopathy Variant Curation Expert Panel classified it as uncertain significance and additional clinical laboratories have submitted both likely pathogenic and uncertain significance assertions.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, consistent with PM2_Supporting under the RAF1 RASopathy specification.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
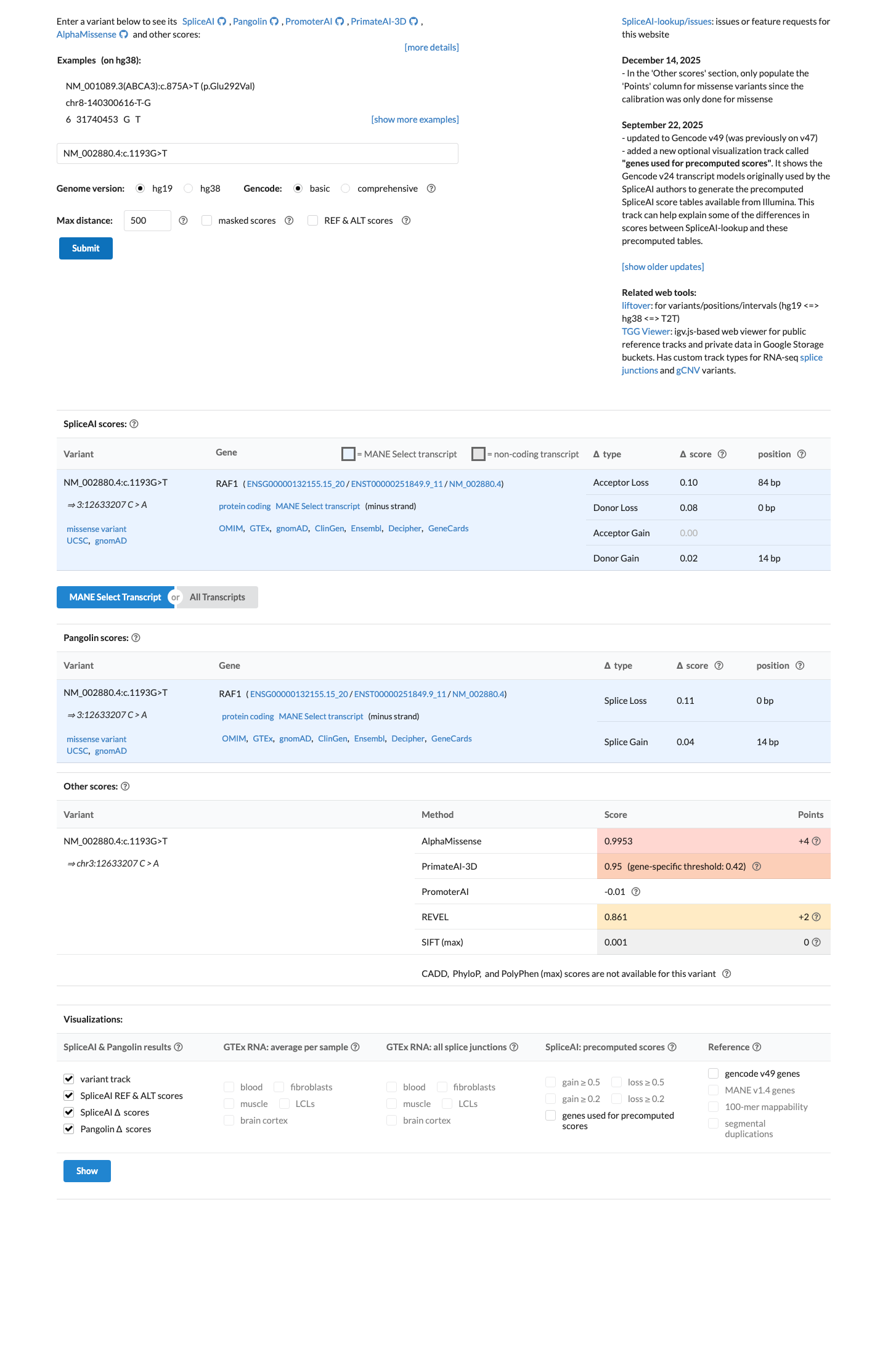
For this missense change, REVEL is 0.861, which exceeds the RAF1 PP3 threshold of 0.7; BayesDel is 0.342905; and SpliceAI predicts no significant splice effect with a maximum delta score of 0.10.
spliceai ↗ cspec ↗