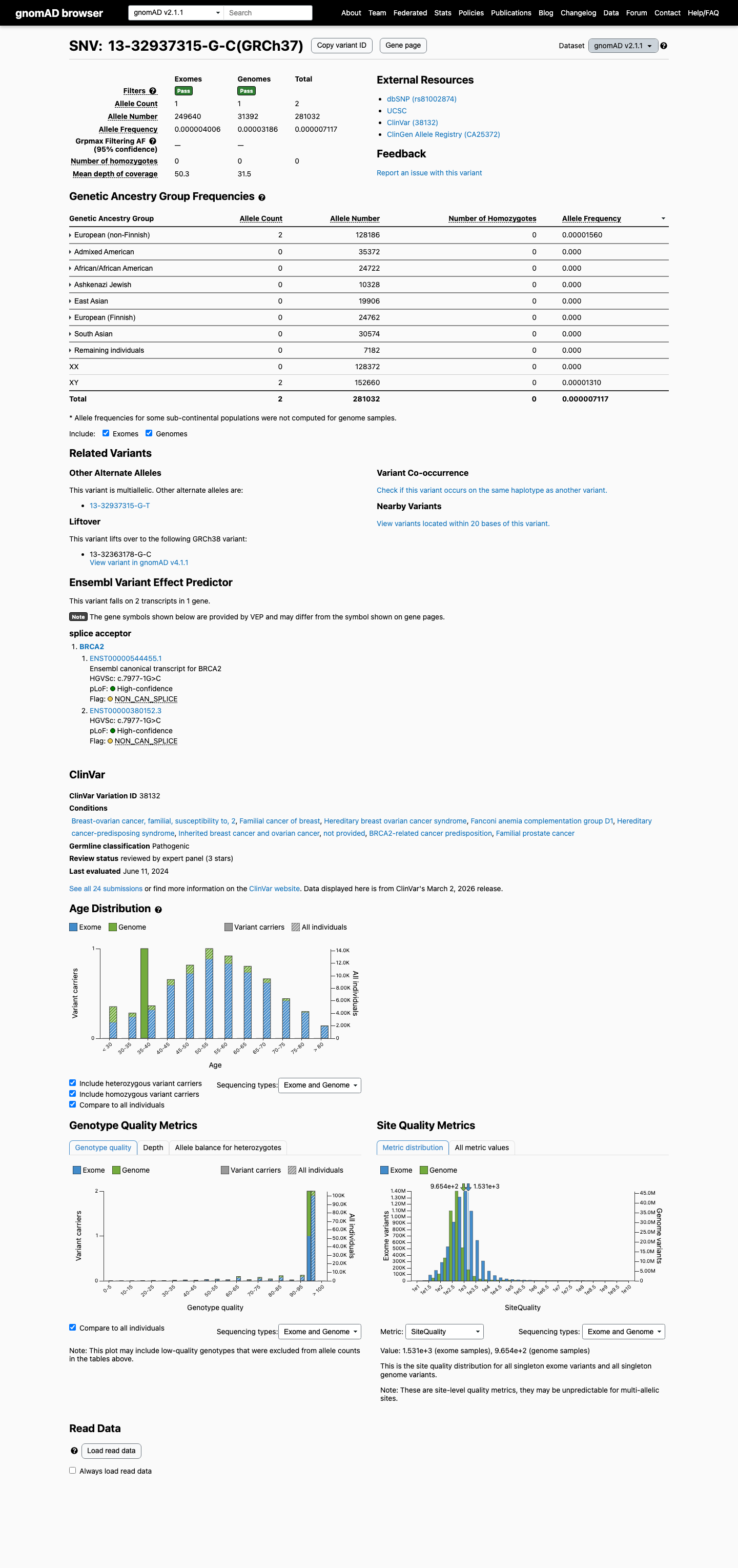
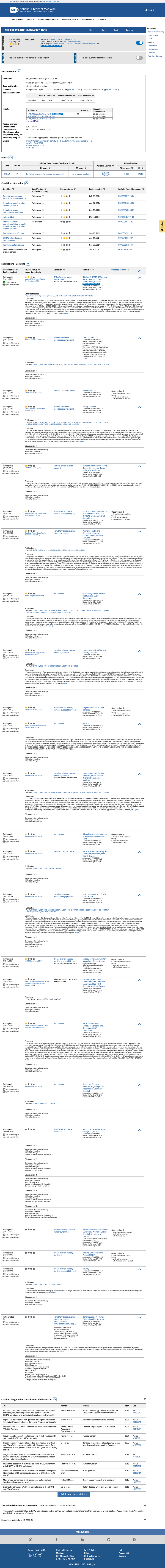
The BRCA2 c.7977-1G>C (p.?) variant has been reported in ClinVar as Pathogenic, including a Pathogenic expert-panel classification from ENIGMA.
clinvar ↗This variant is present at a very low frequency in population databases, with gnomAD v2.1 AF 7.11663e-06 (2/281032 alleles) and gnomAD v4.1 AF 3.10106e-06 (5/1612350 alleles), which is below benign frequency thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗RNA studies summarized in the ENIGMA BRCA2 materials reported abnormal splicing for this variant, including exon 18 deletion and exon 17/18 deletion, and ENIGMA Table 4 assigns PVS1_Strong (RNA) at this splice acceptor because variants at this site show leaky splicing.
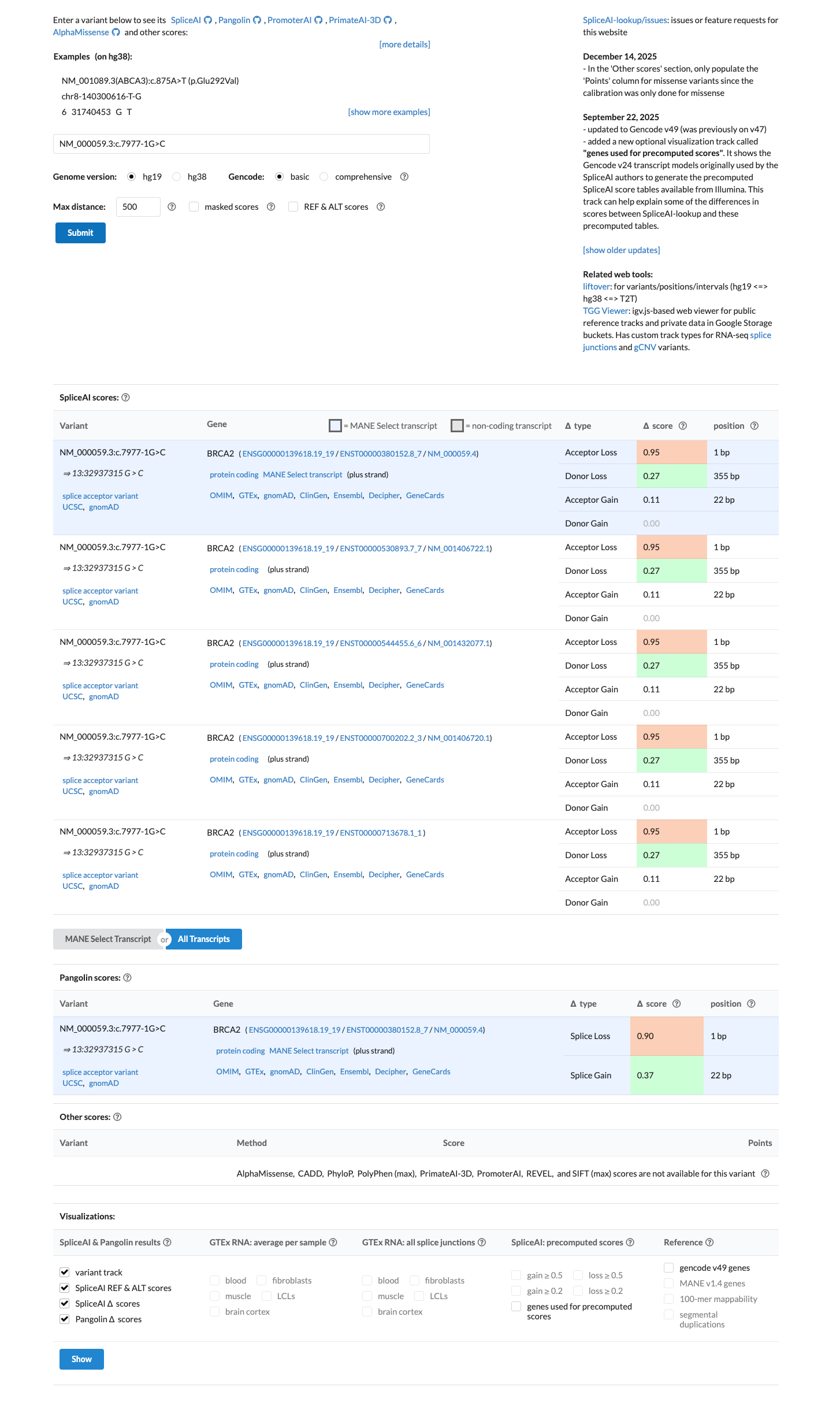
PMID:16211554 ↗Computational splicing analysis is also strongly abnormal, with a SpliceAI maximum delta score of 0.95, consistent with splice disruption, although this prediction is not counted separately because splice-impact evidence is already captured under PVS1 in the ENIGMA framework.
spliceai ↗ cspec ↗In the BRCA2 clinical-history likelihood-ratio dataset, this variant had an LR of 7.223 across 7 probands, which meets the ENIGMA threshold for PP4 at moderate strength.
PMID:31853058 ↗