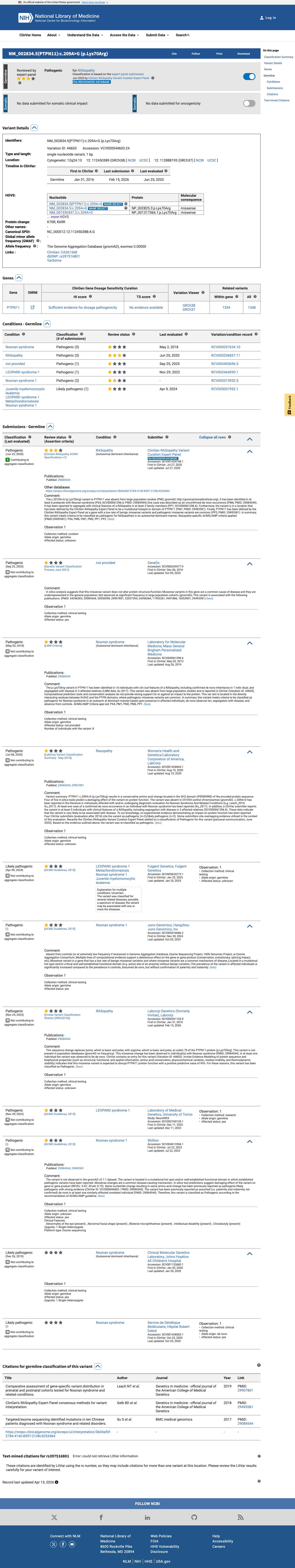
The PTPN11 c.209A>G (p.Lys70Arg) variant has been reported in ClinVar as Pathogenic by the ClinGen RASopathy Variant Curation Expert Panel and was also reported in a patient with Noonan syndrome in a published cohort.
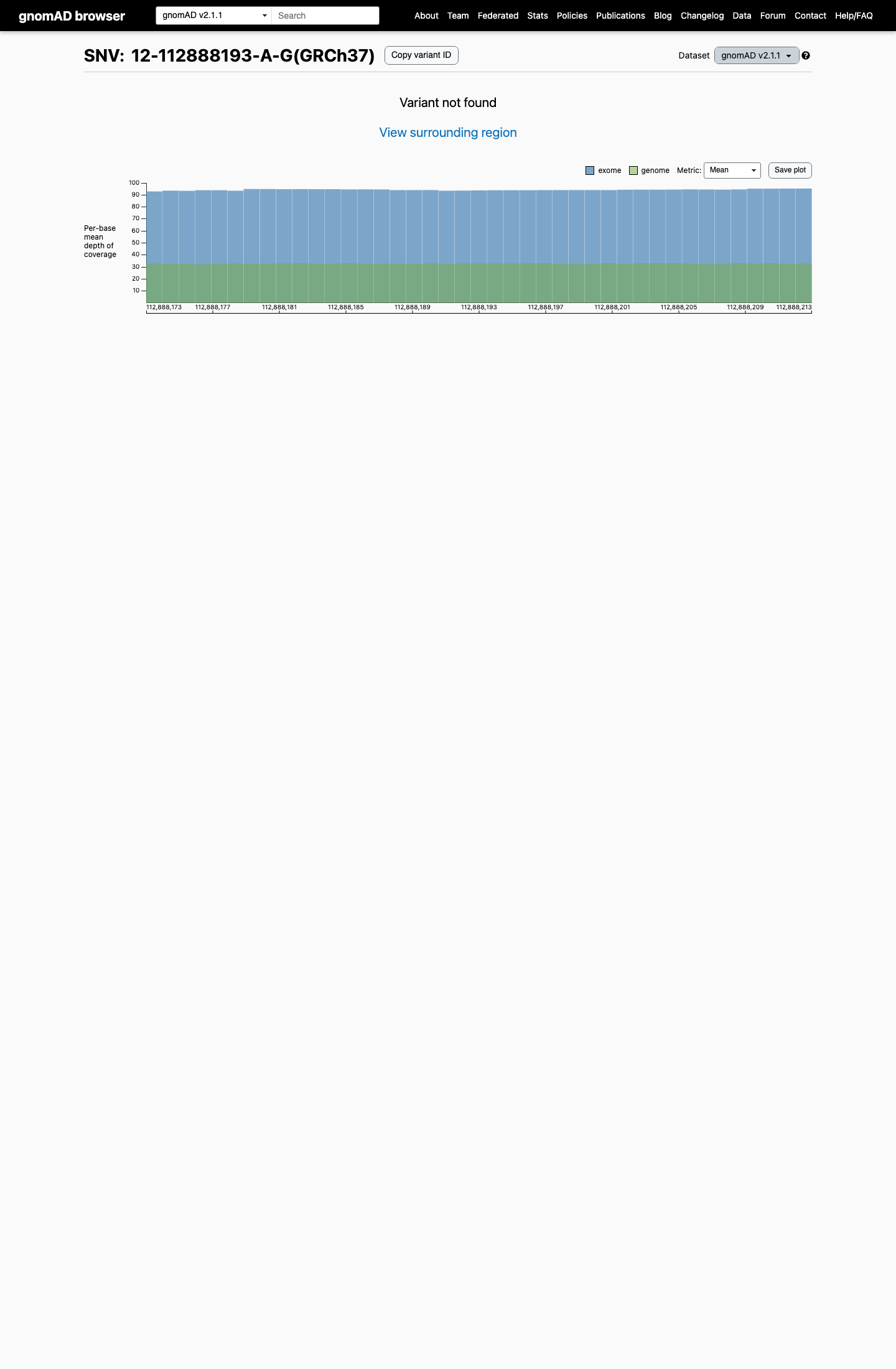
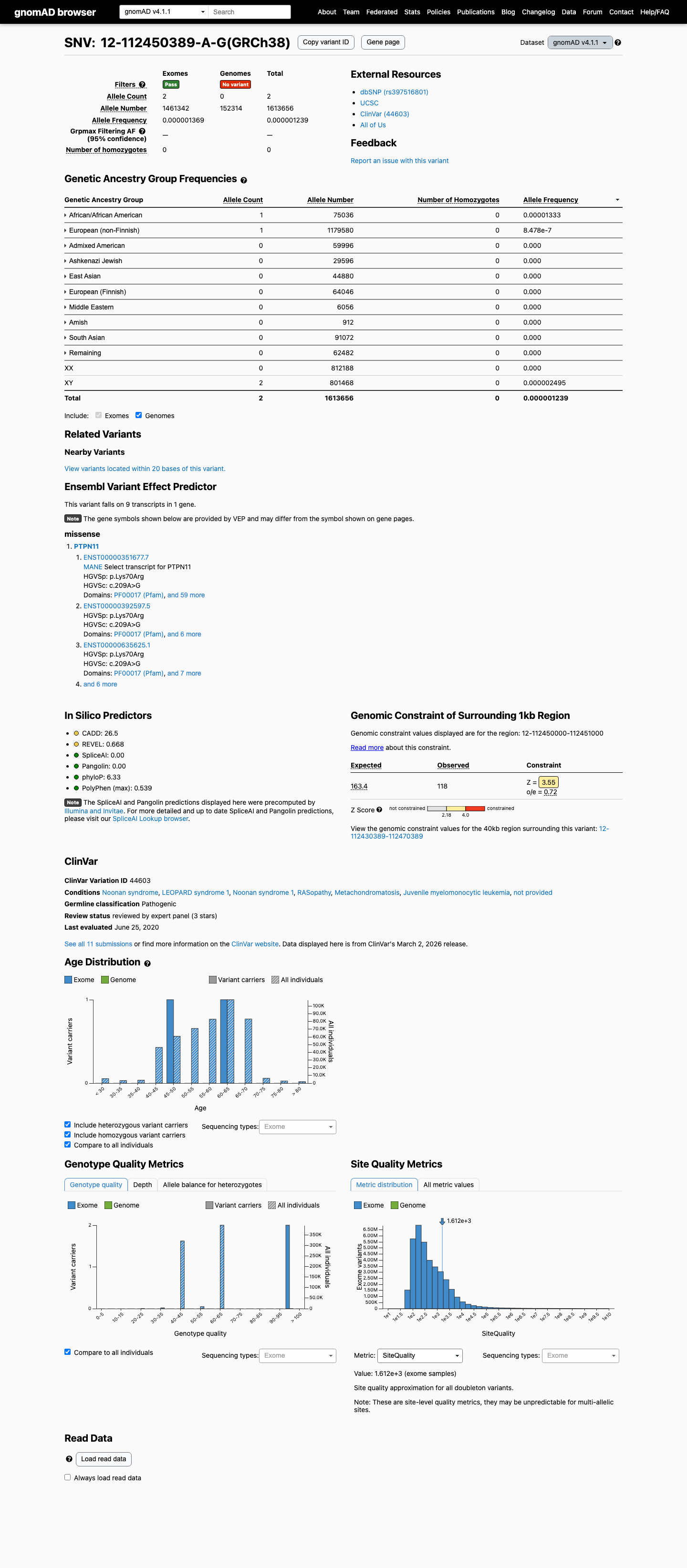
clinvar ↗ PMID:29084544 ↗This variant was absent from gnomAD v2.1 and is present only 2 times among 1613656 alleles in gnomAD v4.1 (AF 1.23942e-06; 0.00012%), which is far below the BS1 threshold of 0.025% and the BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗The affected Lys70 residue falls within the PTPN11 critical N-SH2/PTP interaction region explicitly designated by the RASopathy specification for PM1.
cspec ↗Available computational evidence does not meet PP3 or BP4 thresholds because REVEL is 0.668, BayesDel is 0.00798395, and SpliceAI predicts no significant splice impact with a maximum delta score of 0.00.
spliceai ↗ cspec ↗