Classification rationale
1
The BRCA1 c.5089T>C (p.Cys1697Arg) variant has not been observed in COSMIC and has been reported in ClinVar, including a pathogenic expert-panel classification from ClinGen ENIGMA.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population controls.
gnomad_v2 ↗ gnomad_v4 ↗3
In calibrated functional studies summarized by the BRCA1 expert-panel resources, this variant showed loss of BRCA1 function, and the expert-panel table assigns PS3_Strong.
4
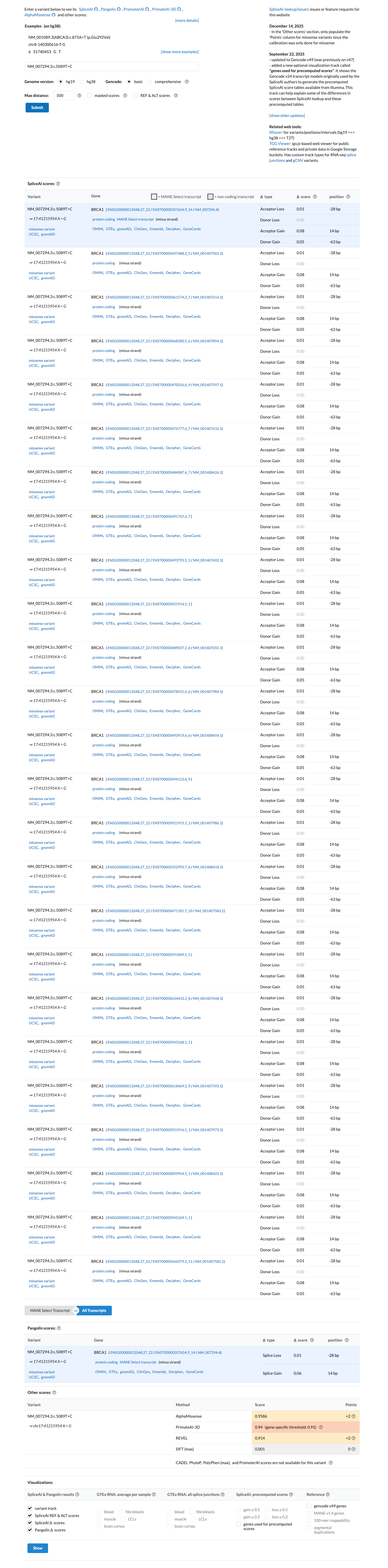
Computational evidence supports a damaging protein effect because the variant is in the BRCA1 BRCT repeats, BayesDel is 0.398132 (above the PP3 threshold of 0.28), and REVEL is 0.914, while SpliceAI predicts no significant splice effect with a max delta score of 0.08.
cspec ↗ spliceai ↗