Classification rationale
1
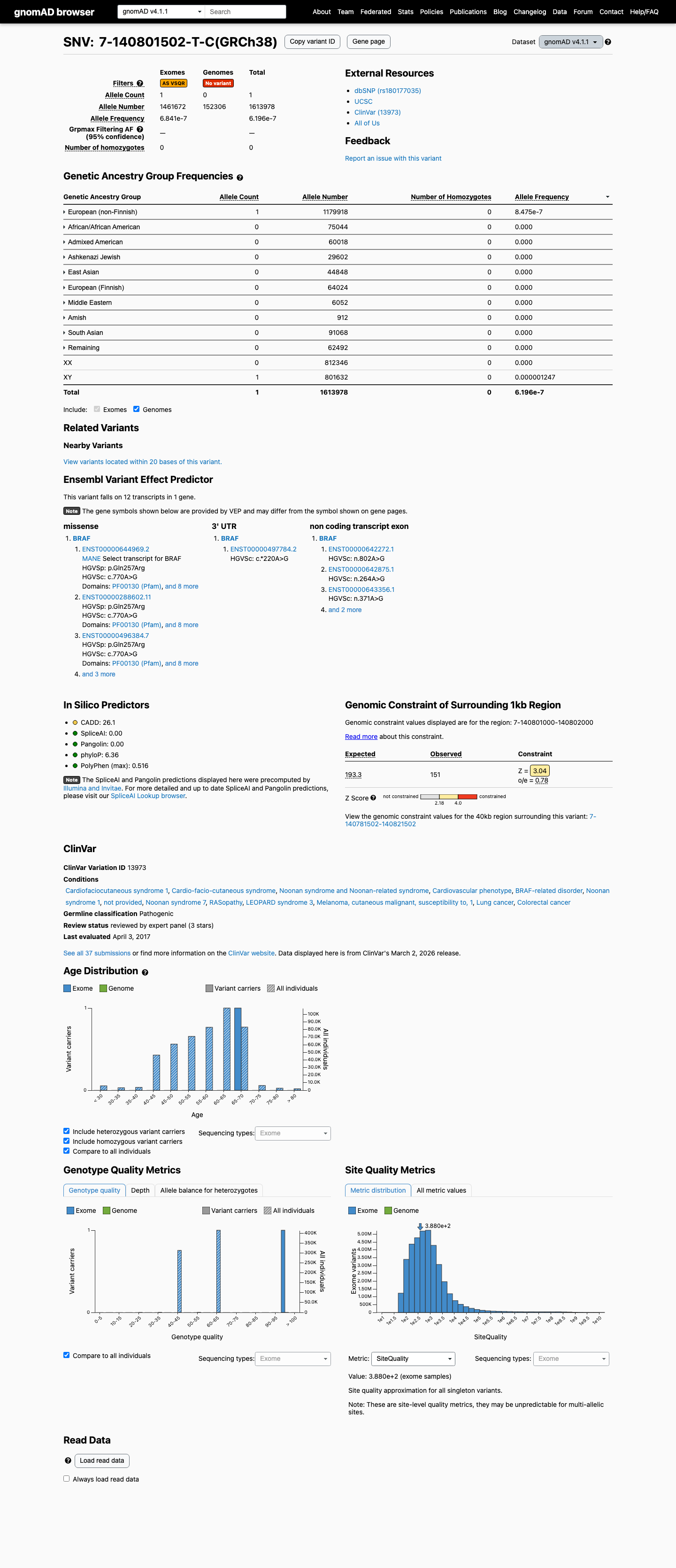
The BRAF c.770A>G (p.Gln257Arg) variant has been reported in ClinVar as pathogenic, including expert panel review.
clinvar ↗2
This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 (1/1,613,978 alleles; AF 0.00006%), which is below the BS1 threshold of 0.025% and the BA1 threshold of 0.05%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
In VCEP-approved functional studies, Q257R showed abnormal results in MEK activation, ERK activation, and BRAF kinase activity assays, consistent with a damaging gain-of-function effect.
cspec ↗4
Computational evidence supports a deleterious missense effect, with REVEL 0.838 above the PP3 threshold of 0.7, a positive BayesDel score of 0.476355, and SpliceAI showing only a low predicted splice effect with a maximum delta score of 0.21.
spliceai ↗ cspec ↗