Classification rationale
1
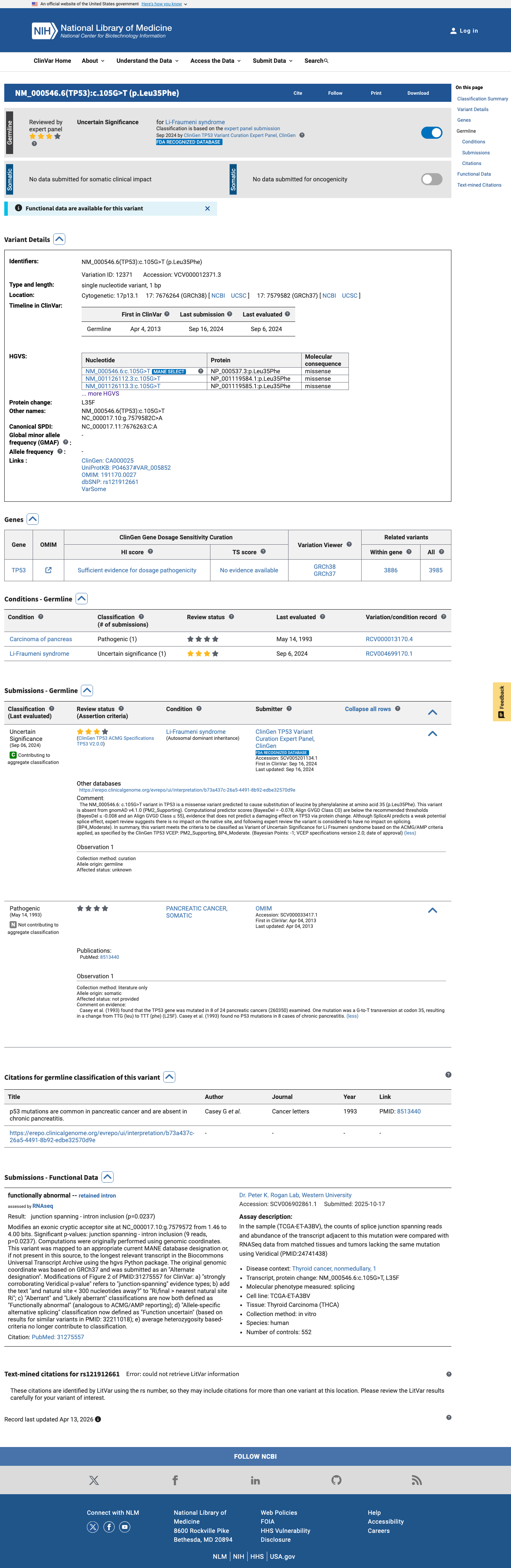
The TP53 c.105G>T (p.Leu35Phe, p.L35F) variant has been reported in ClinVar as a variant of uncertain significance by the ClinGen TP53 Variant Curation Expert Panel.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting marked rarity in population databases.
gnomad_v2 ↗ gnomad_v4 ↗3
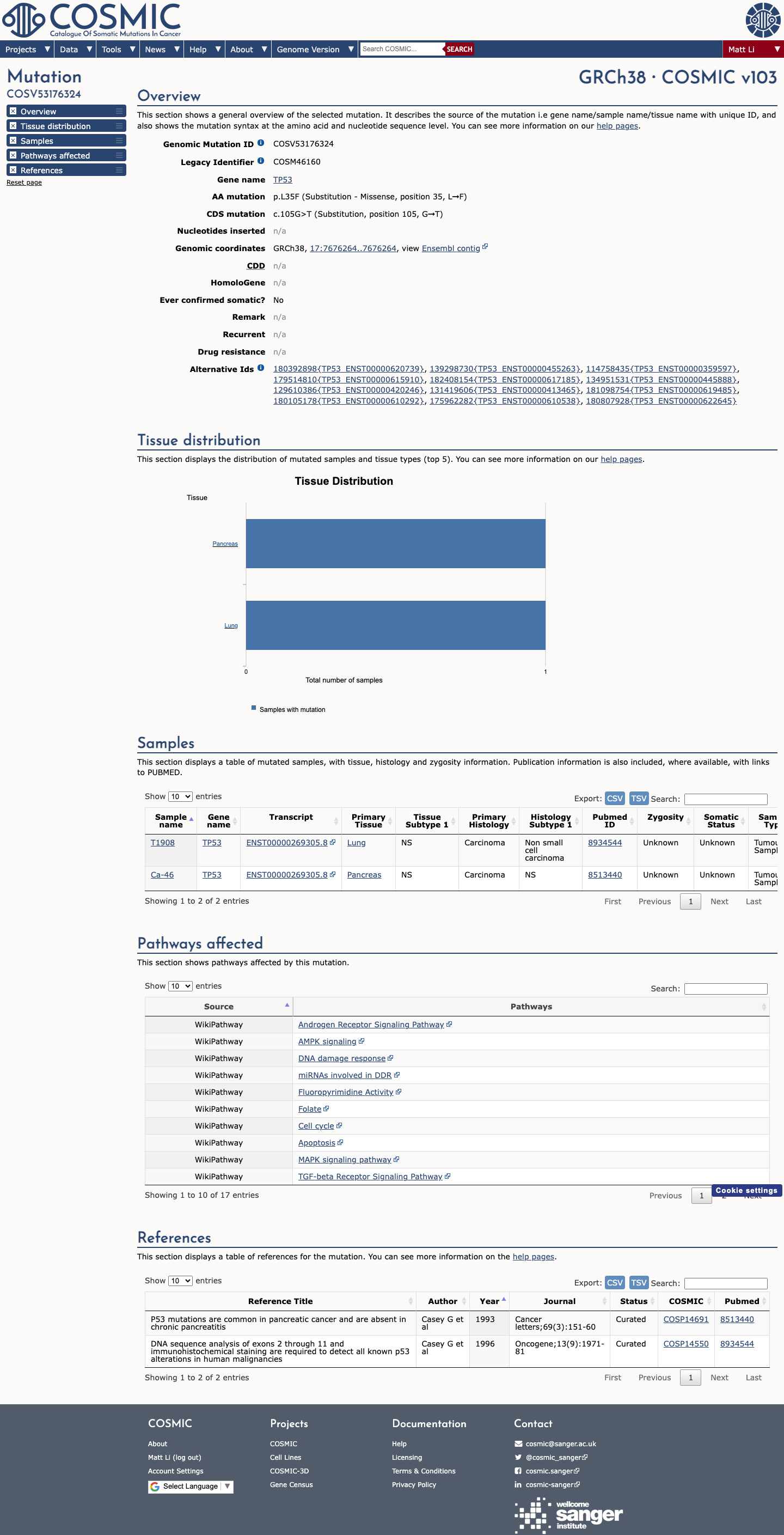
In the TP53 functional evidence framework, p.L35F is listed as functional with no loss of function, supporting BS3 and arguing against PS3.
4
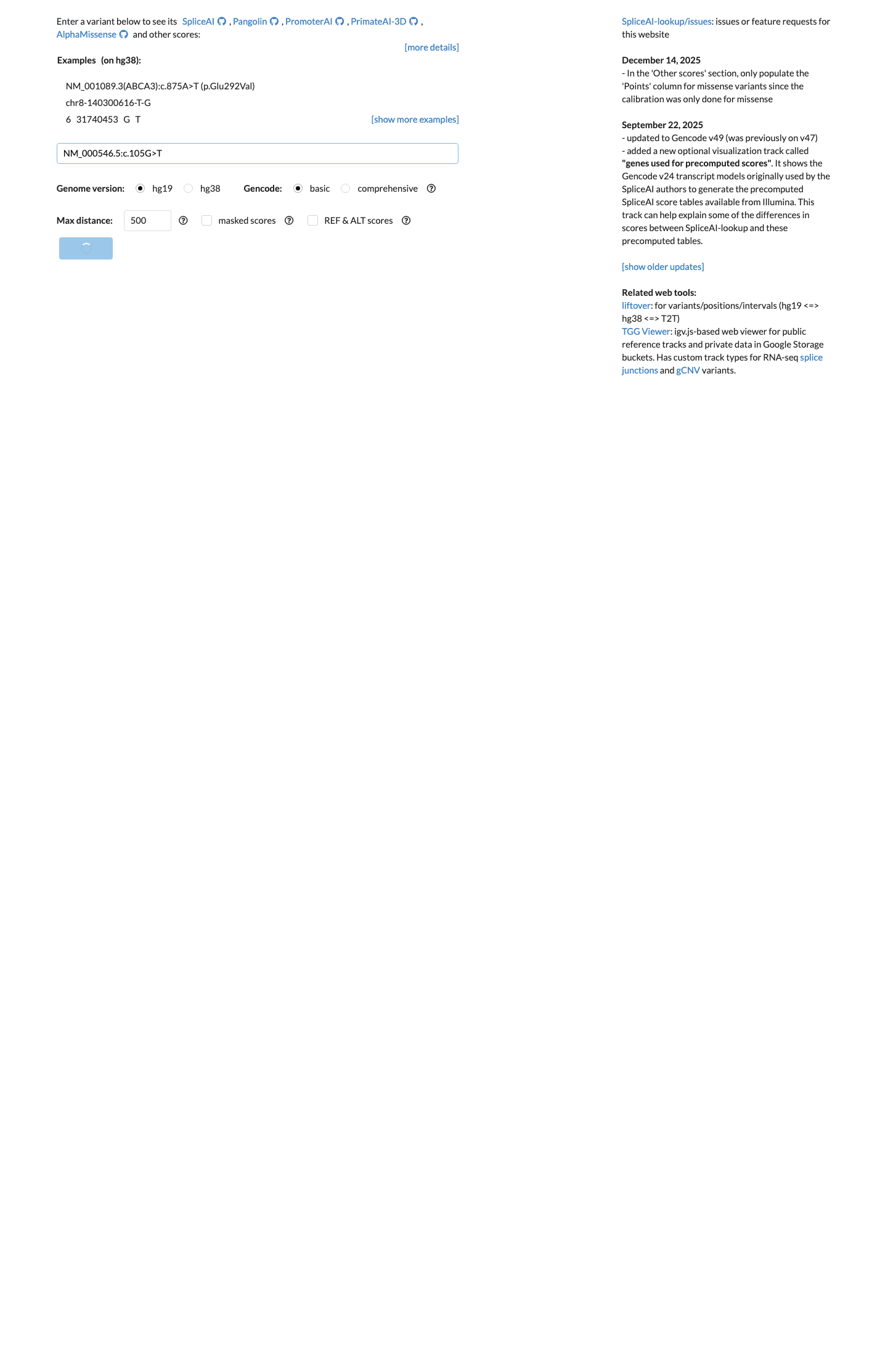
Computational evidence does not support a damaging effect: SpliceAI predicts no splice impact with a max delta score of 0.00, BayesDel is -0.0784713, REVEL is 0.486, and the TP53 bioinformatic worksheet assigns BP4_moderate.
spliceai ↗