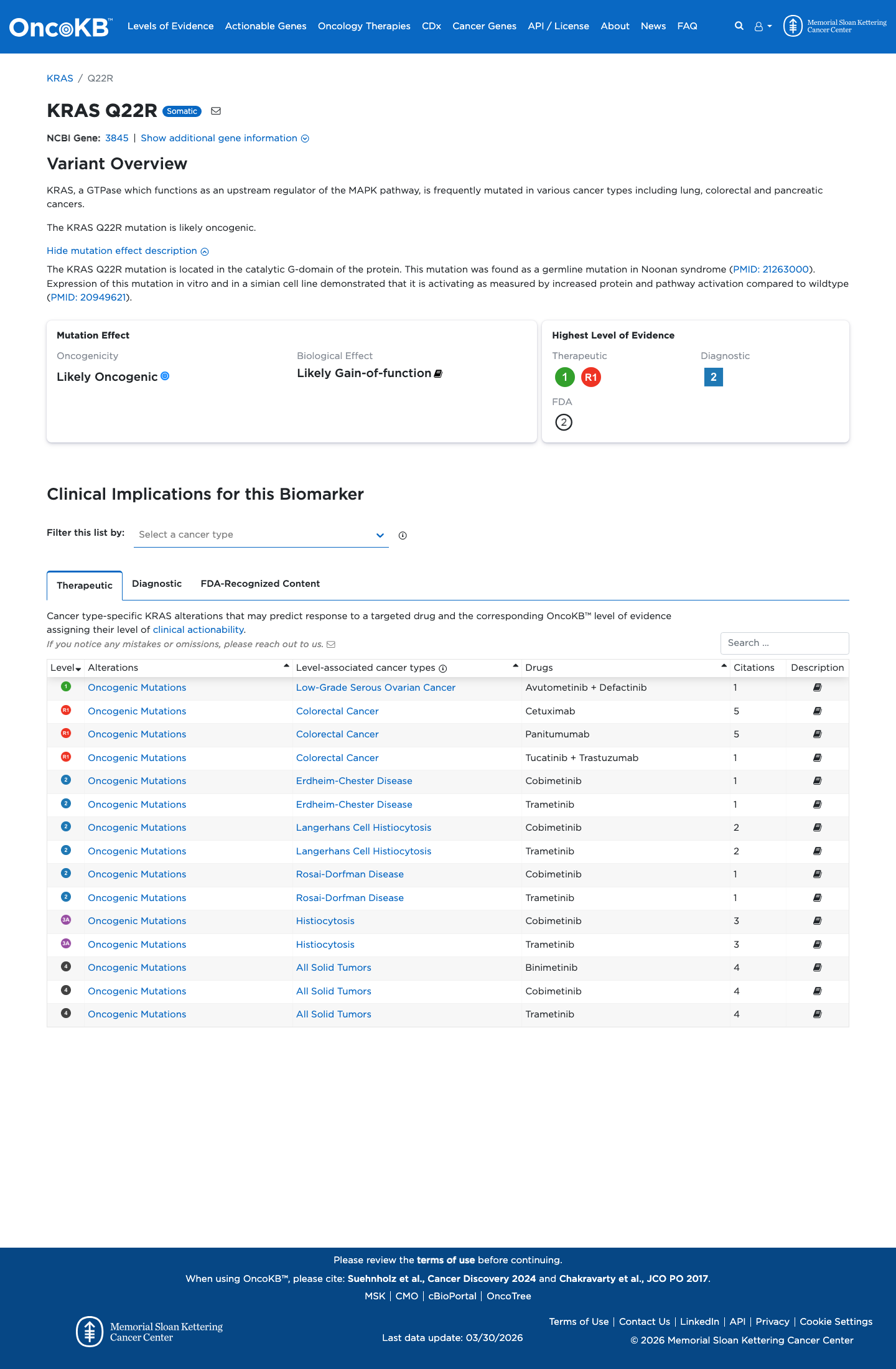
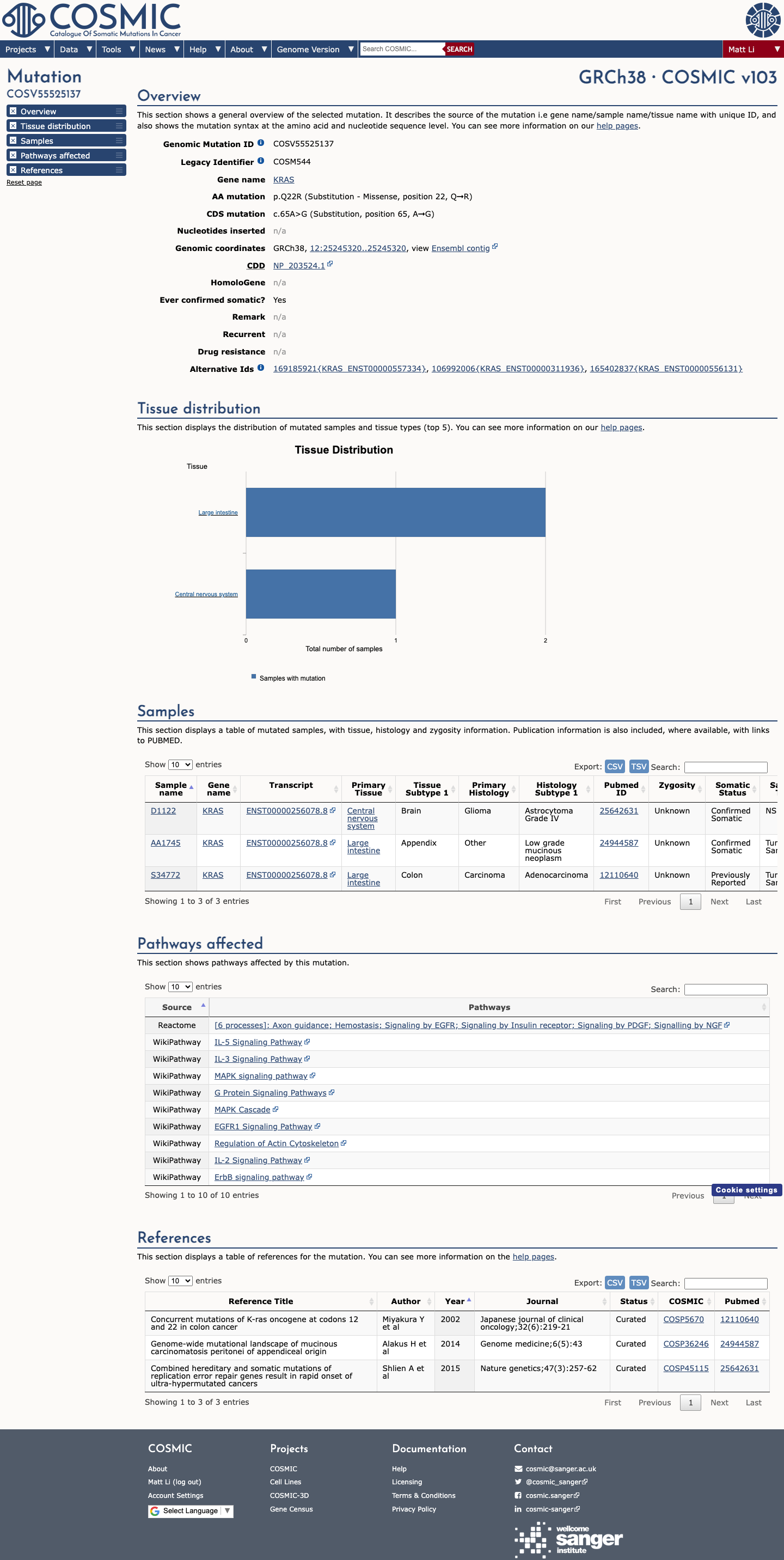
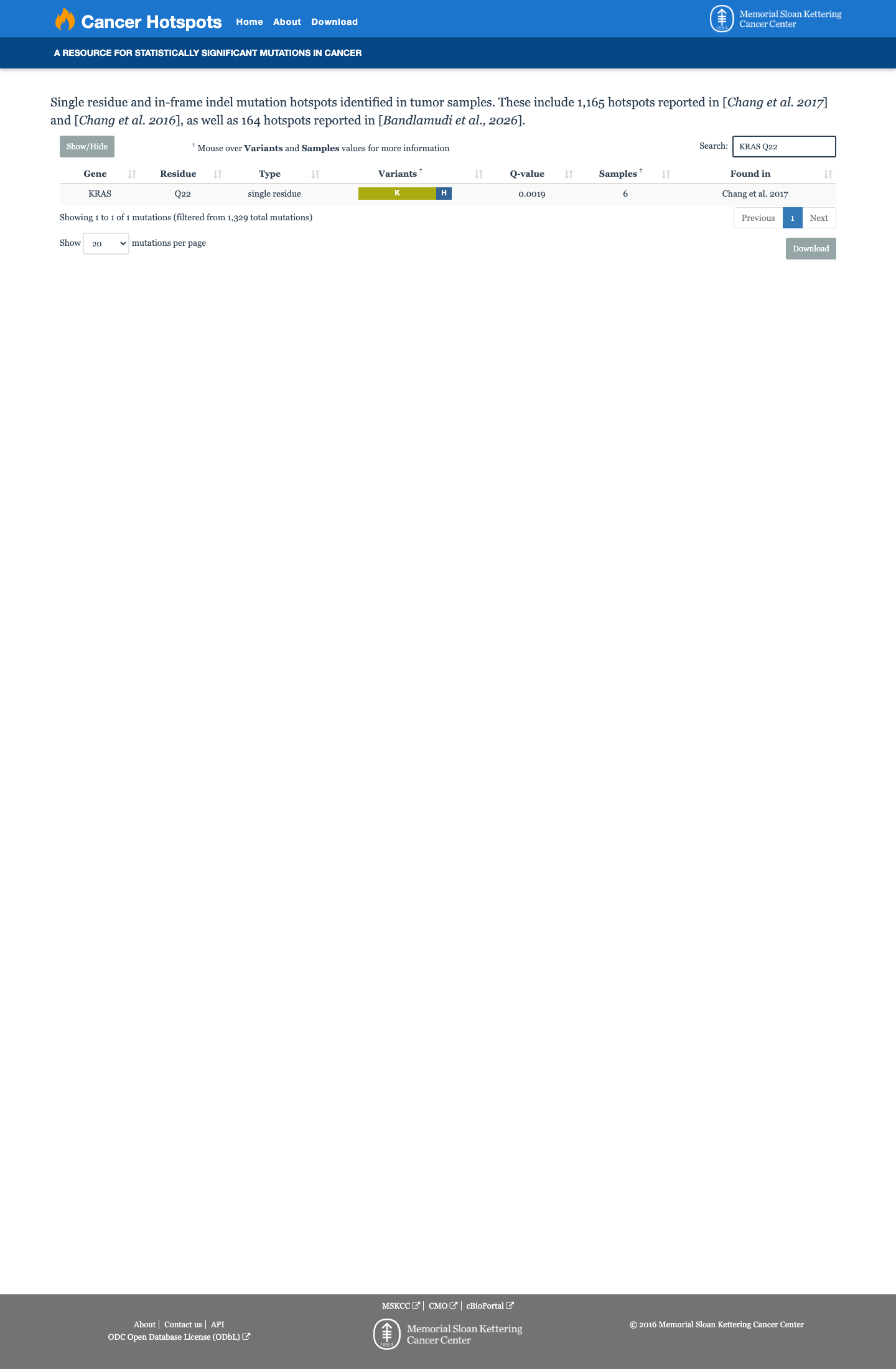
The KRAS NM_001369786.1:c.65A>G (p.Gln22Arg, p.Q22R) variant has been observed in somatic cancers in COSMIC (3 occurrences) and has been reported in ClinVar as pathogenic, including expert-panel review by the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which supports PM2 at supporting strength and is below the KRAS benign frequency thresholds for BS1 and BA1.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Published KRAS functional studies included p.Gln22Arg among germline KRAS variants with abnormal biochemical or signaling properties, but the currently reviewed RASopathy VCEP functional-study matrix did not clearly establish a qualifying approved-assay count for PS3 assignment.
PMID:20949621 ↗ cspec ↗Computational evidence supports a damaging missense effect, with REVEL 0.749 meeting the KRAS PP3 threshold of at least 0.7, while SpliceAI predicts no meaningful splice effect with a maximum delta score of 0.01; the BayesDel score is 0.273188.
spliceai ↗ cspec ↗