The NRAS c.368G>A (p.Arg123Lys) variant has not been observed in COSMIC and is reported in ClinVar as uncertain significance, including an expert panel submission from the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in the general population and meeting the NRAS RASopathy VCEP PM2_Supporting rule.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗The reviewed RASopathy VCEP functional study materials identify approved assay types for NRAS, but no variant-specific approved assay result for p.Arg123Lys was identified, so PS3 is not currently supported.
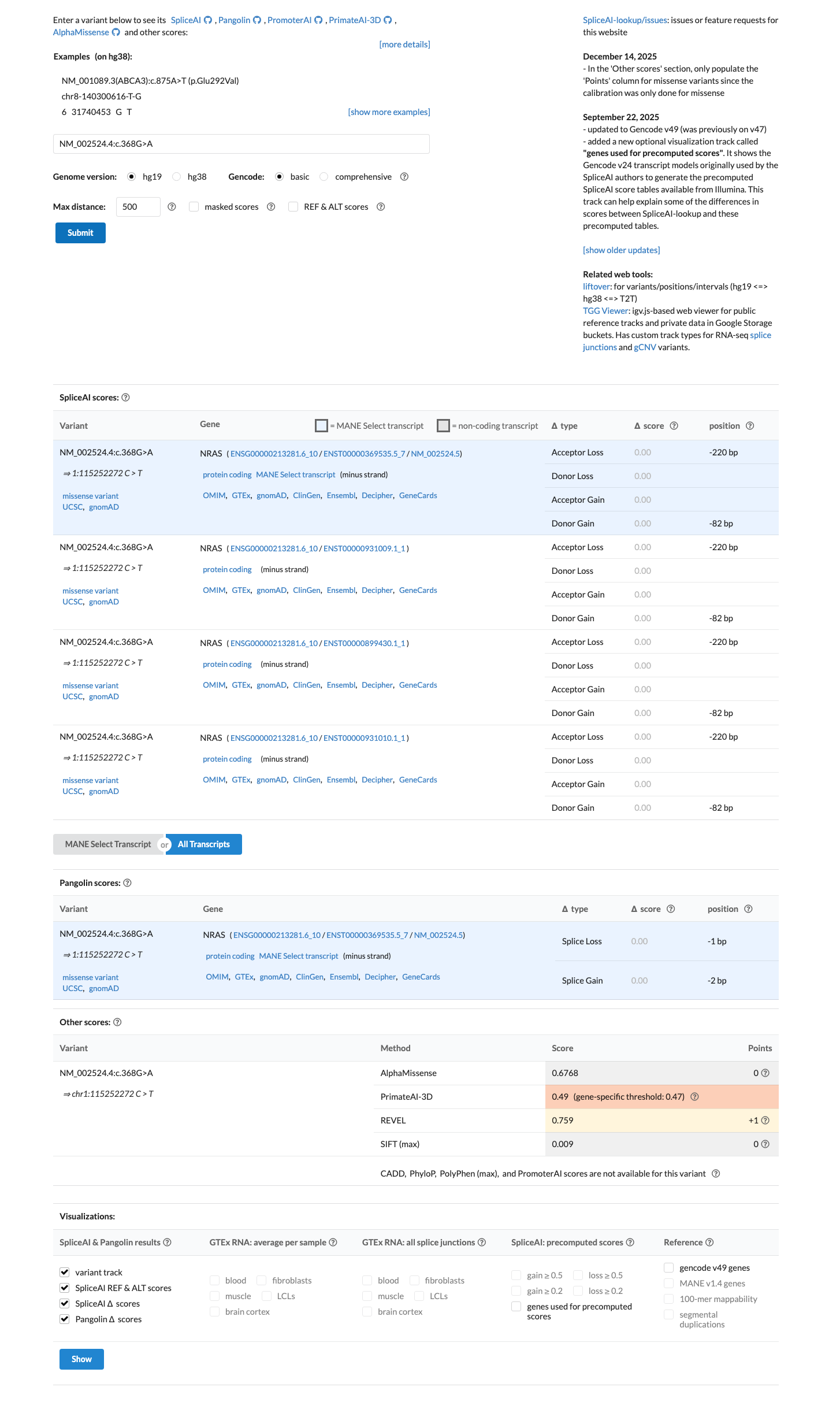
cspec ↗In silico evidence supports a deleterious missense effect, with REVEL 0.759 above the NRAS RASopathy VCEP PP3 threshold of 0.7, BayesDel 0.166482 directionally consistent with a damaging prediction, and SpliceAI predicting no significant splice impact with a maximum delta score of 0.00.
spliceai ↗ cspec ↗