The TP53 c.1136G>T (p.Arg379Leu) variant has been reported in ClinVar, where the ClinGen TP53 Variant Curation Expert Panel classifies it as Likely Benign, although additional clinical laboratory submissions are uncertain.
clinvar ↗This variant is absent from gnomAD v2.1 and is present at very low frequency in gnomAD v4.1 (3/1614000 alleles; AF 1.85874e-06), supporting PM2_Supporting and arguing against BS1 or BA1.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In the TP53 VCEP functional worksheet, p.Arg379Leu is recorded as partially functional in Kato-class data and noLOF in Giacomelli-class data, with a preassigned interpretation of no functional evidence, so PS3 and BS3 are not supported.
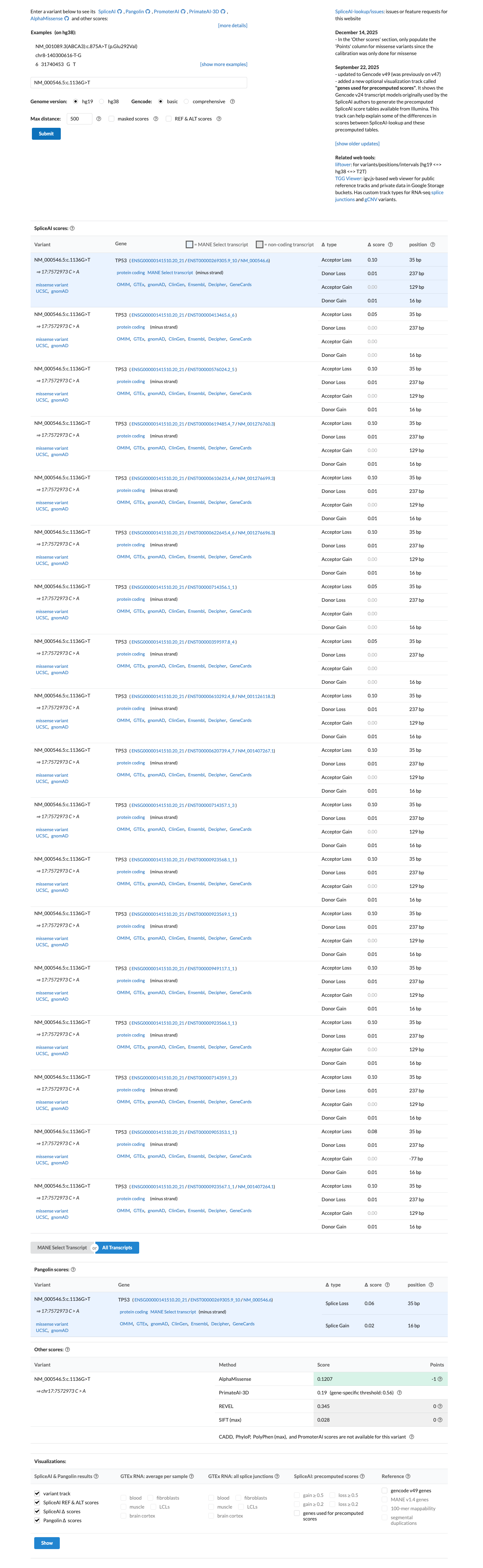
PMID:12826609 ↗ PMID:30224644 ↗TP53-specific in silico assessment lists c.1136G>T as Class C0 with BayesDel -0.0781755 and BP4_moderate, SpliceAI predicts no significant splice impact with max delta score 0.10, and the available REVEL score is 0.345; together these data support BP4_Moderate rather than PP3.
spliceai ↗