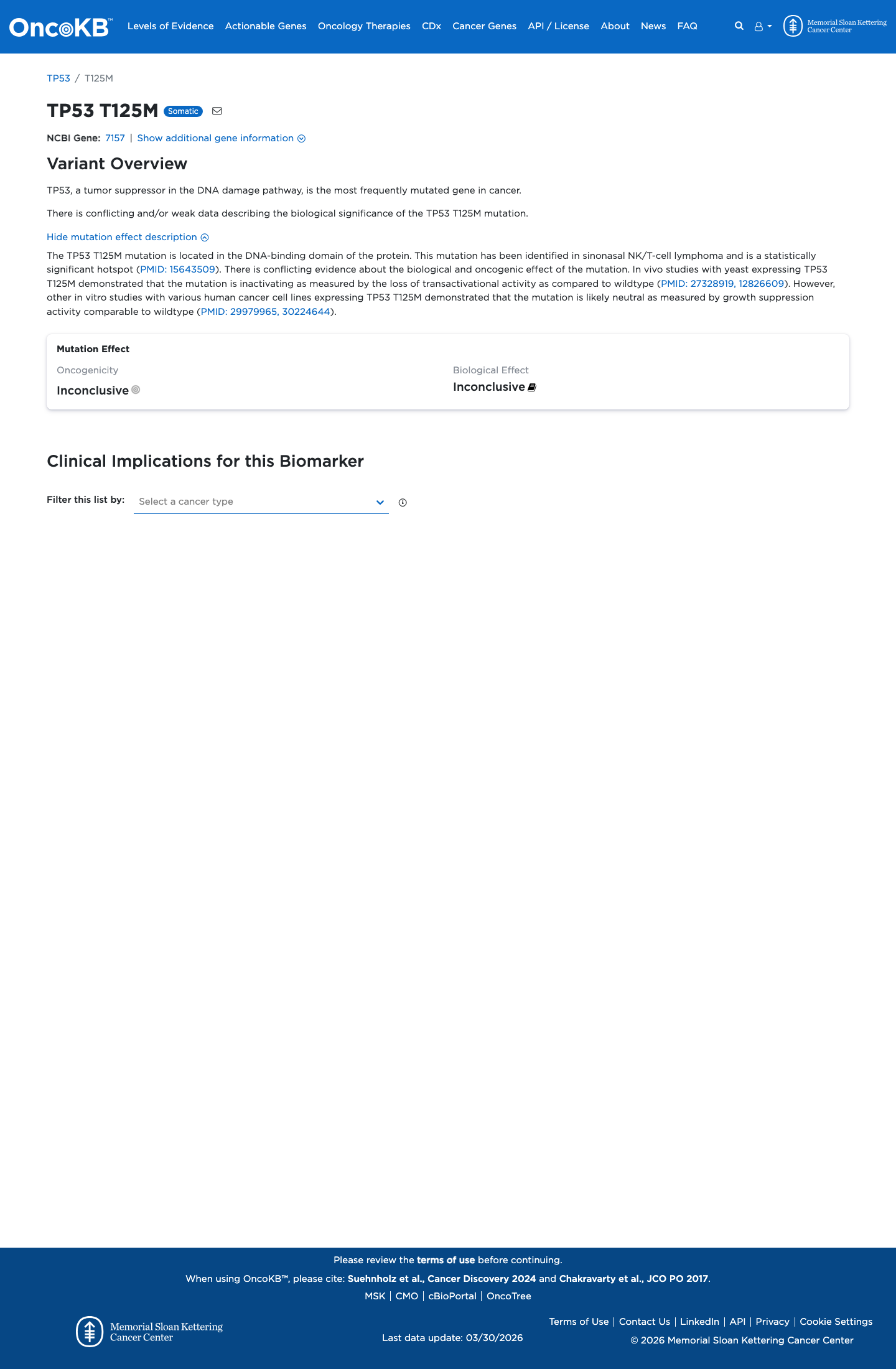
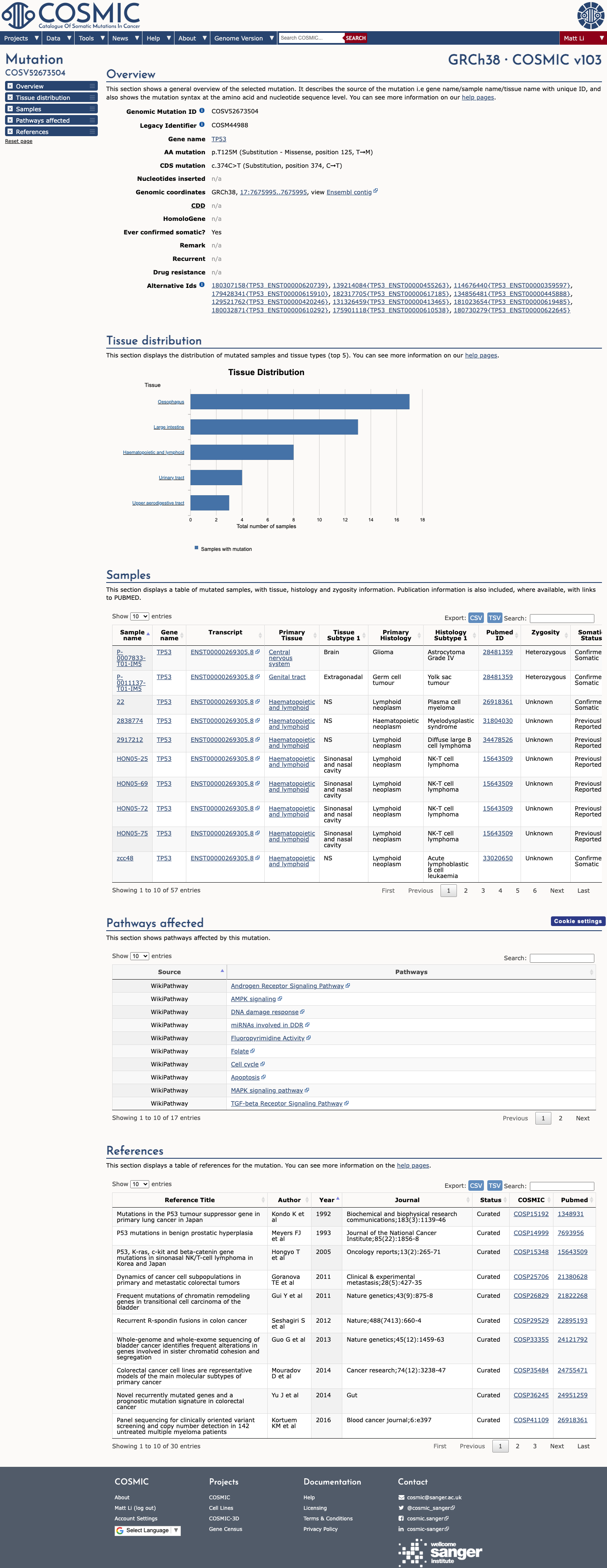
The TP53 c.374C>T (p.Thr125Met) variant has been reported in ClinVar, where most submissions classify it as likely pathogenic or pathogenic.
clinvar ↗This variant is rare in population databases, with gnomAD v4.1 showing an allele frequency of 1.86088e-06 (3/1612144) and a highest observed African/African American frequency of 1.33568e-05, which is below the TP53 VCEP PM2_Supporting threshold of 0.00003.
gnomad_v4 ↗ cspec ↗In the TP53 VCEP functional worksheet, p.Thr125Met is listed as non-functional in Kato et al. but as no loss of function in other eligible assays, leading to a preassigned interpretation of "No evidence" rather than PS3 or BS3.
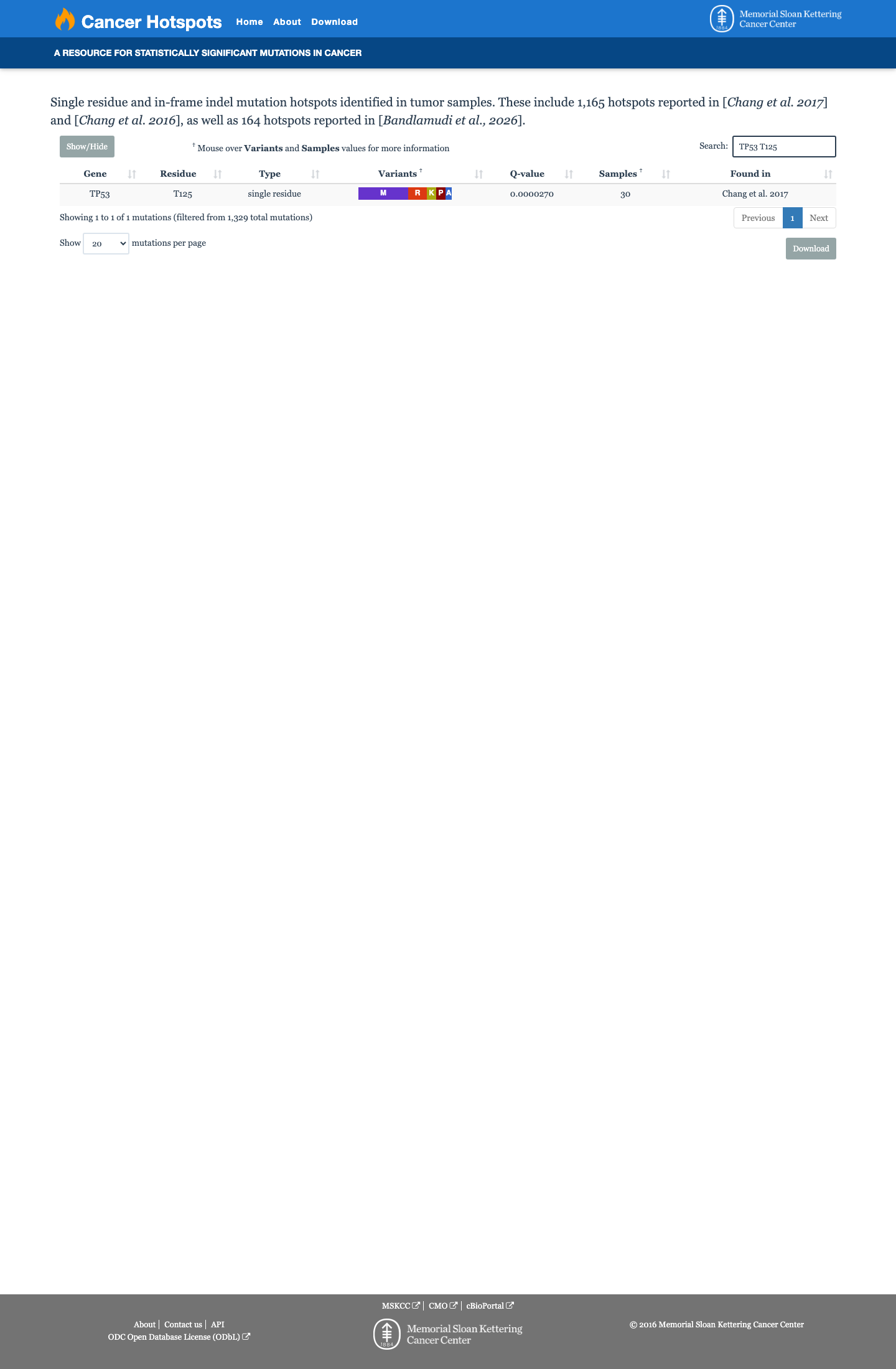
PMID:12826609 ↗ PMID:29979965 ↗ PMID:30224644 ↗TP53 VCEP computational evidence supports pathogenicity at the PP3_moderate level, with aGVGD Class C65 and BayesDel 0.500232 in the TP53 worksheet; REVEL is 0.925, while SpliceAI predicts no significant splice effect with a maximum delta score of 0.09.
spliceai ↗