Classification rationale
1
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, which supports rarity in population reference datasets.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
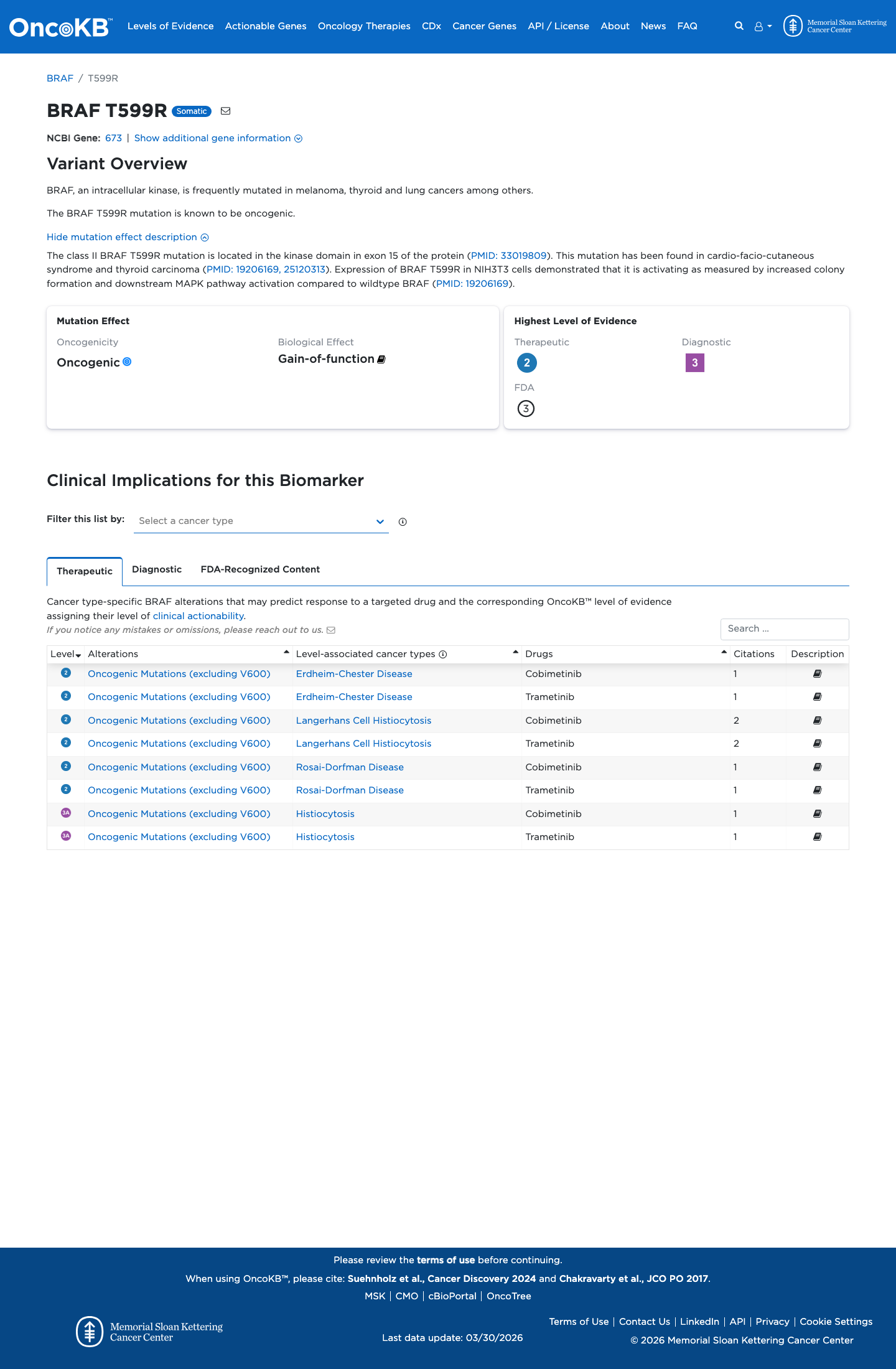
In approved functional studies, this variant was classified as pathogenic in both MEK activation and ERK activation assays, supporting abnormal MAPK pathway signaling consistent with the disease mechanism.
cspec ↗ PMID:19206169 ↗4
In silico evidence does not meet the BRAF RASopathy thresholds for PP3 or BP4 because REVEL is 0.359, SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, and BayesDel is 0.308415.
spliceai ↗ cspec ↗