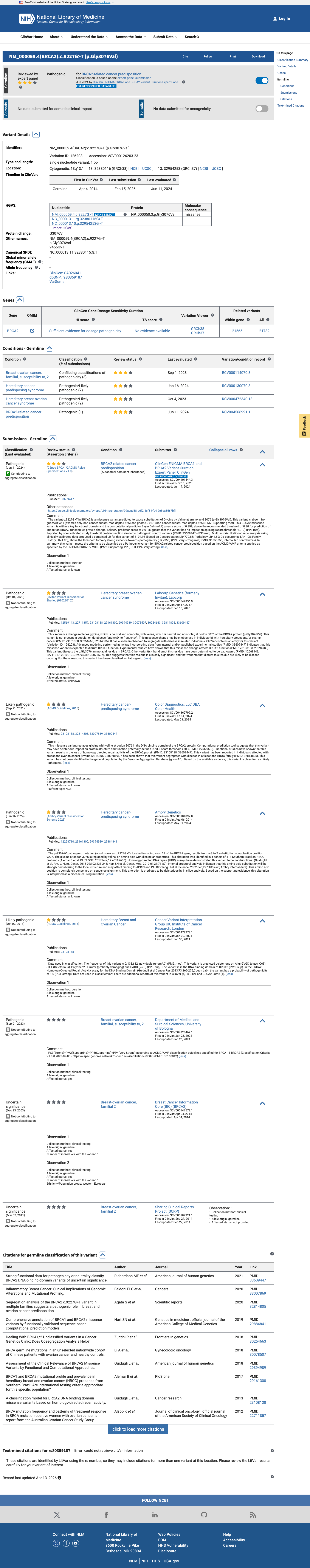
The BRCA2 c.9227G>T (p.Gly3076Val) variant has not been observed in COSMIC and has been reported in ClinVar, including a Pathogenic expert-panel classification from ClinGen ENIGMA.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which supports rarity, although the site-level depth information required for formal ENIGMA PM2_Supporting application was not identified here.
gnomad_v2 ↗ gnomad_v4 ↗In the ENIGMA BRCA2 calibrated functional dataset, p.(Gly3076Val) showed abnormal function similar to pathogenic control variants, supporting PS3 at strong strength.
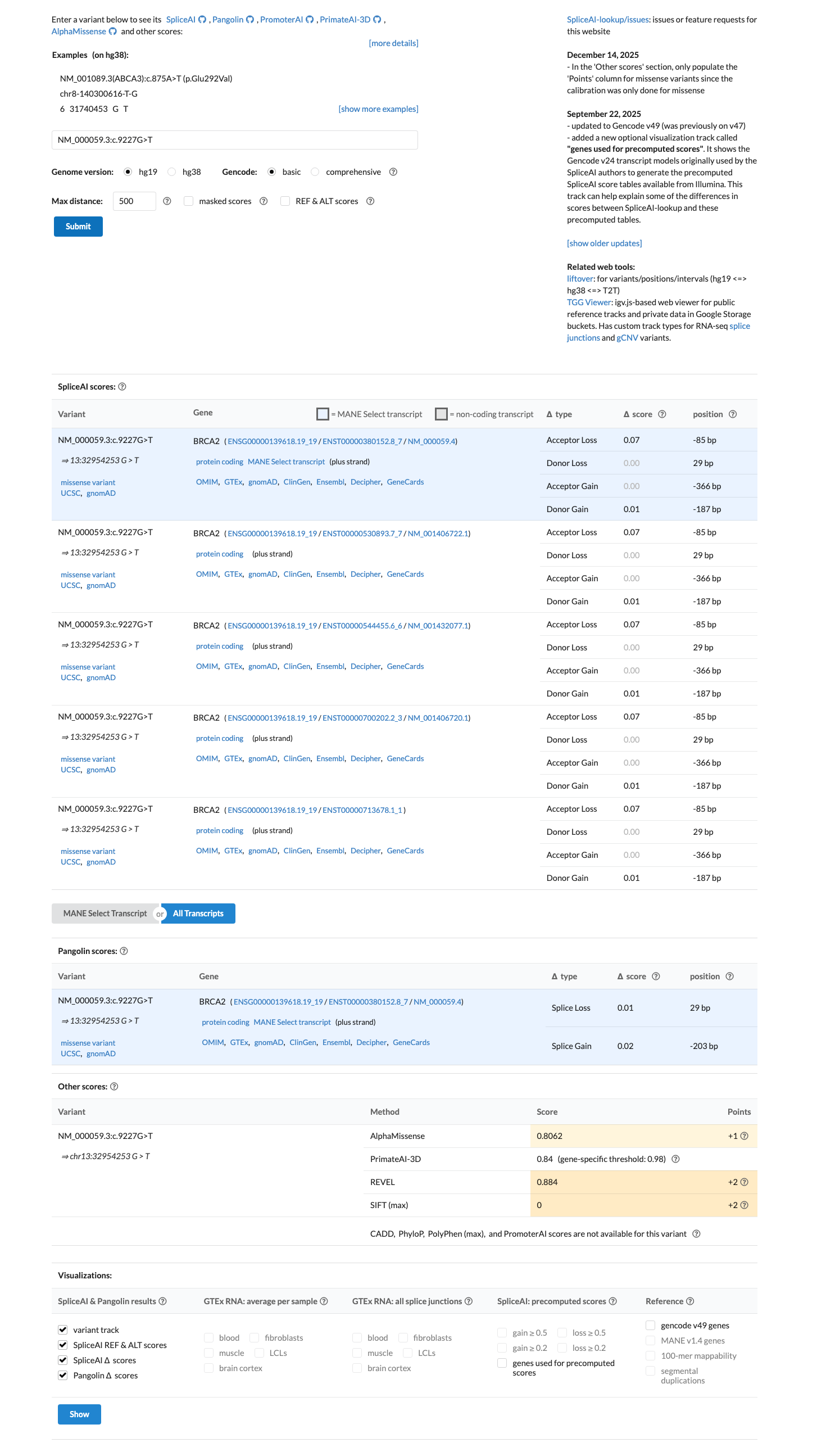
This missense change lies within the BRCA2 DNA-binding domain; BayesDel no-AF is 0.377938, REVEL is 0.884, and SpliceAI max delta is 0.07, supporting a damaging protein effect without a predicted splice effect and therefore supporting PP3 but not BP4.
cspec ↗ spliceai ↗