The BRCA1 c.4327C>G (p.Arg1443Gly; p.R1443G) variant has been reported in ClinVar, where the aggregate classification is Benign with expert panel review.
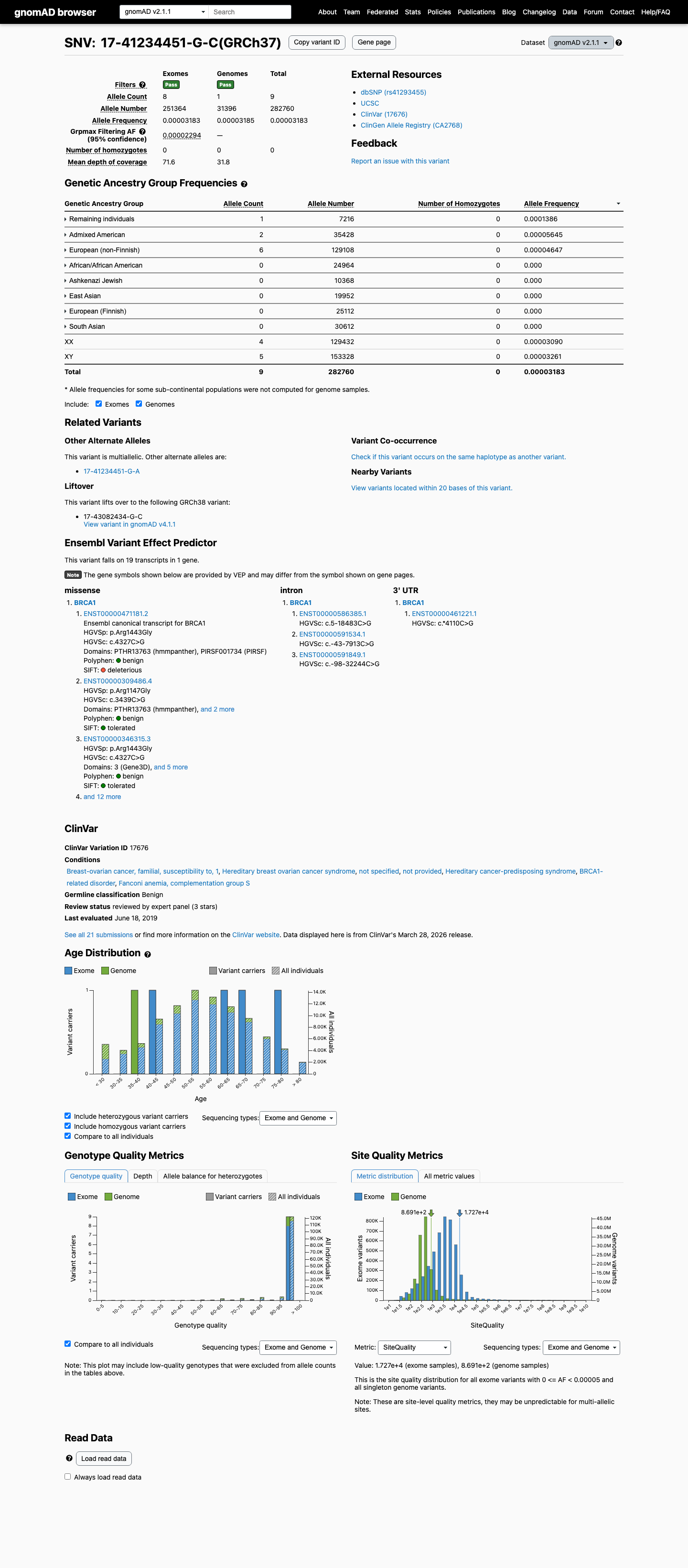
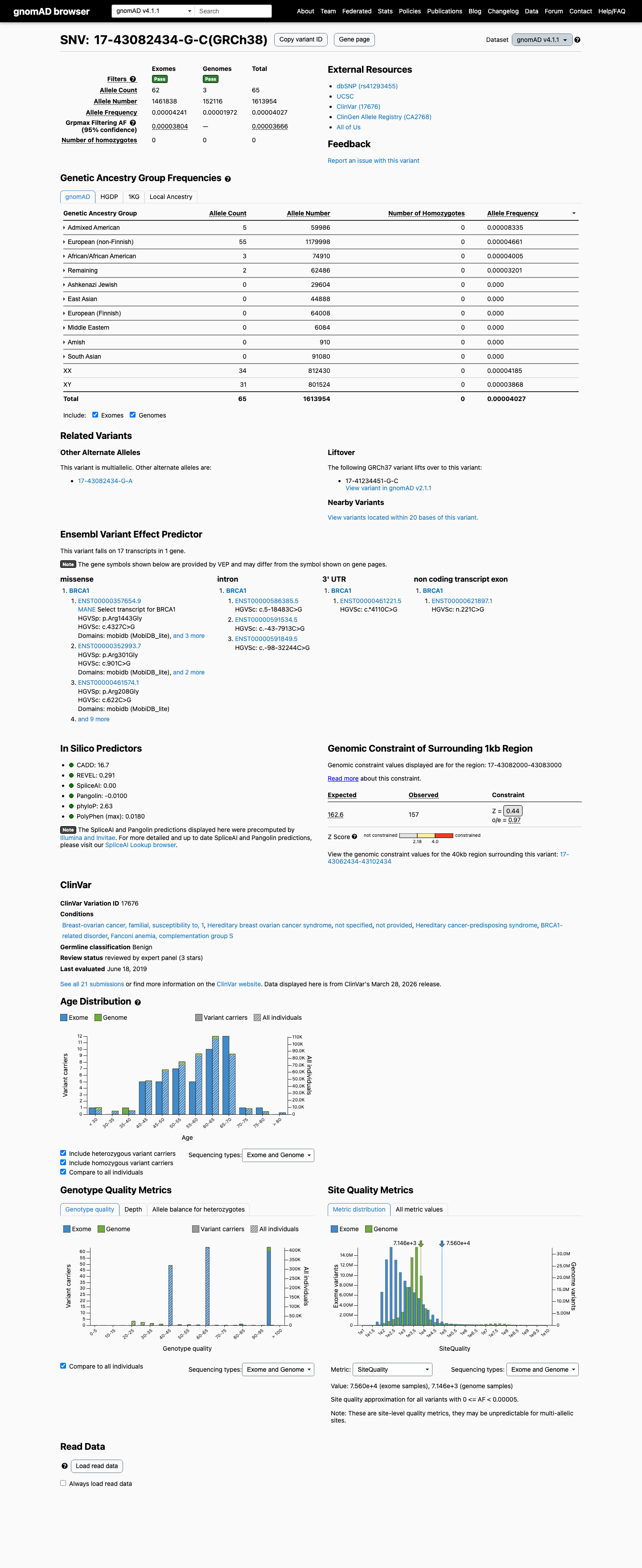
clinvar ↗This variant is present in population databases, including gnomAD v2.1 at 9/282760 alleles (AF 3.18291e-05; grpmax FAF 2.294e-05) and gnomAD v4.1 at 65/1613954 alleles (AF 4.02738e-05; grpmax FAF 3.666e-05), which exceeds the BRCA1 BS1 supporting threshold of 0.00002 but remains below the BA1 threshold of 0.001.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In two calibrated functional studies curated by ENIGMA, this variant showed no functional impact and protein behavior similar to benign control variants, supporting BS3 at strong strength.
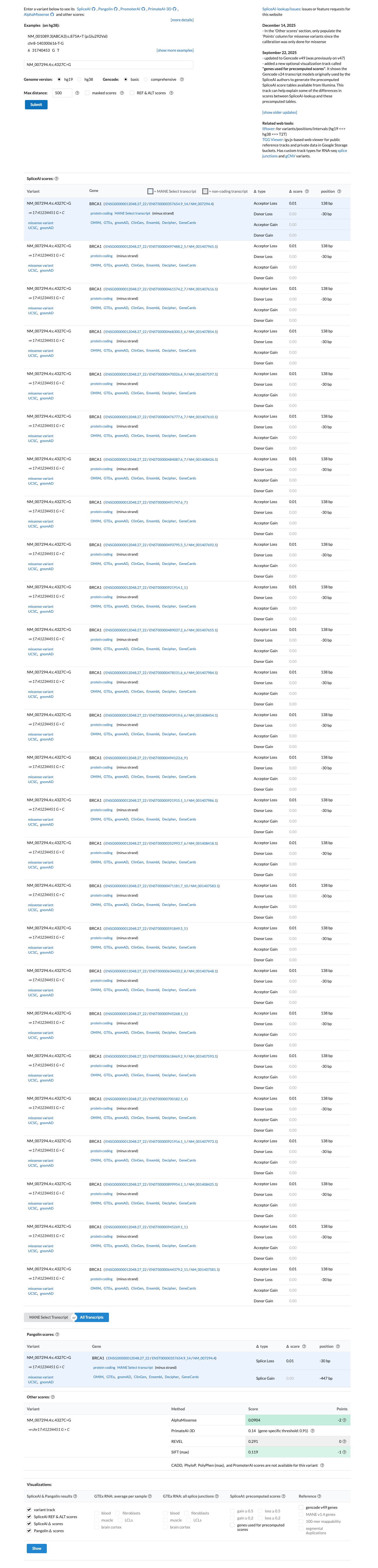
Computational evidence does not support a damaging effect: SpliceAI predicts no significant splice impact with a maximum delta score of 0.01, BayesDel no-AF is -0.285466, and REVEL is 0.007; because Arg1443 lies outside the BRCA1 ENIGMA clinically important functional domains, this profile supports BP1_Strong rather than PP3.
spliceai ↗ cspec ↗