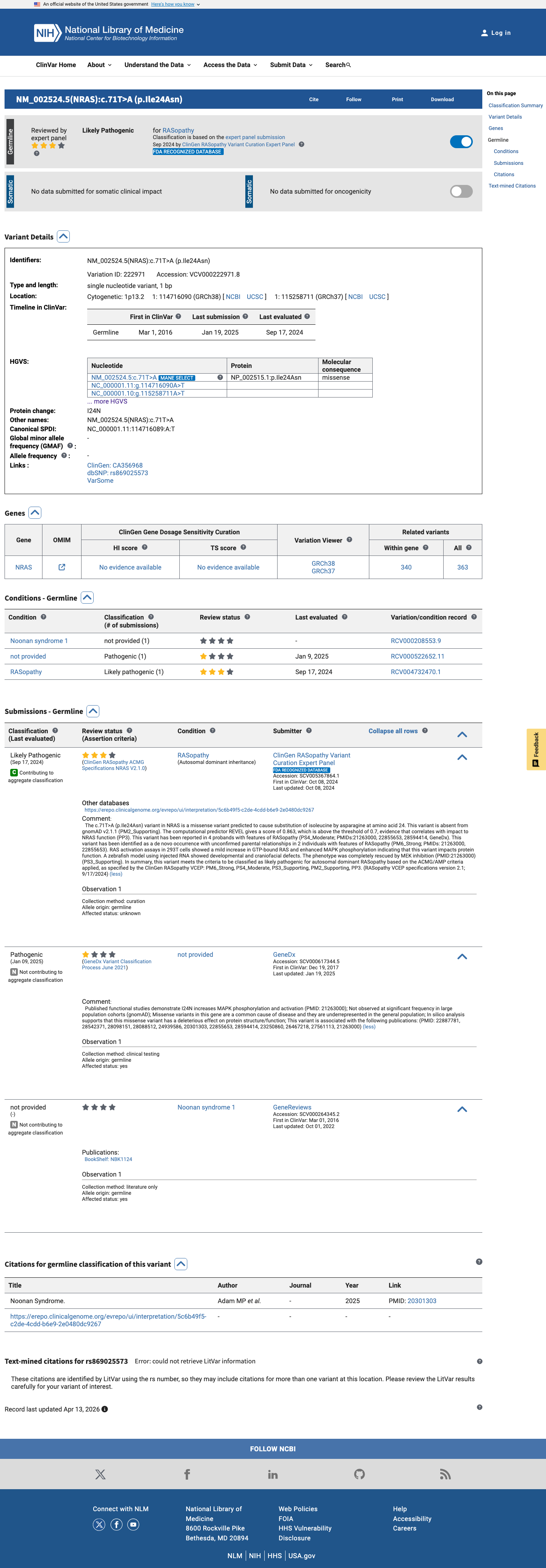
The NRAS c.71T>A (p.Ile24Asn) variant has been reported in ClinVar and is classified as Likely Pathogenic by the ClinGen RASopathy expert panel.
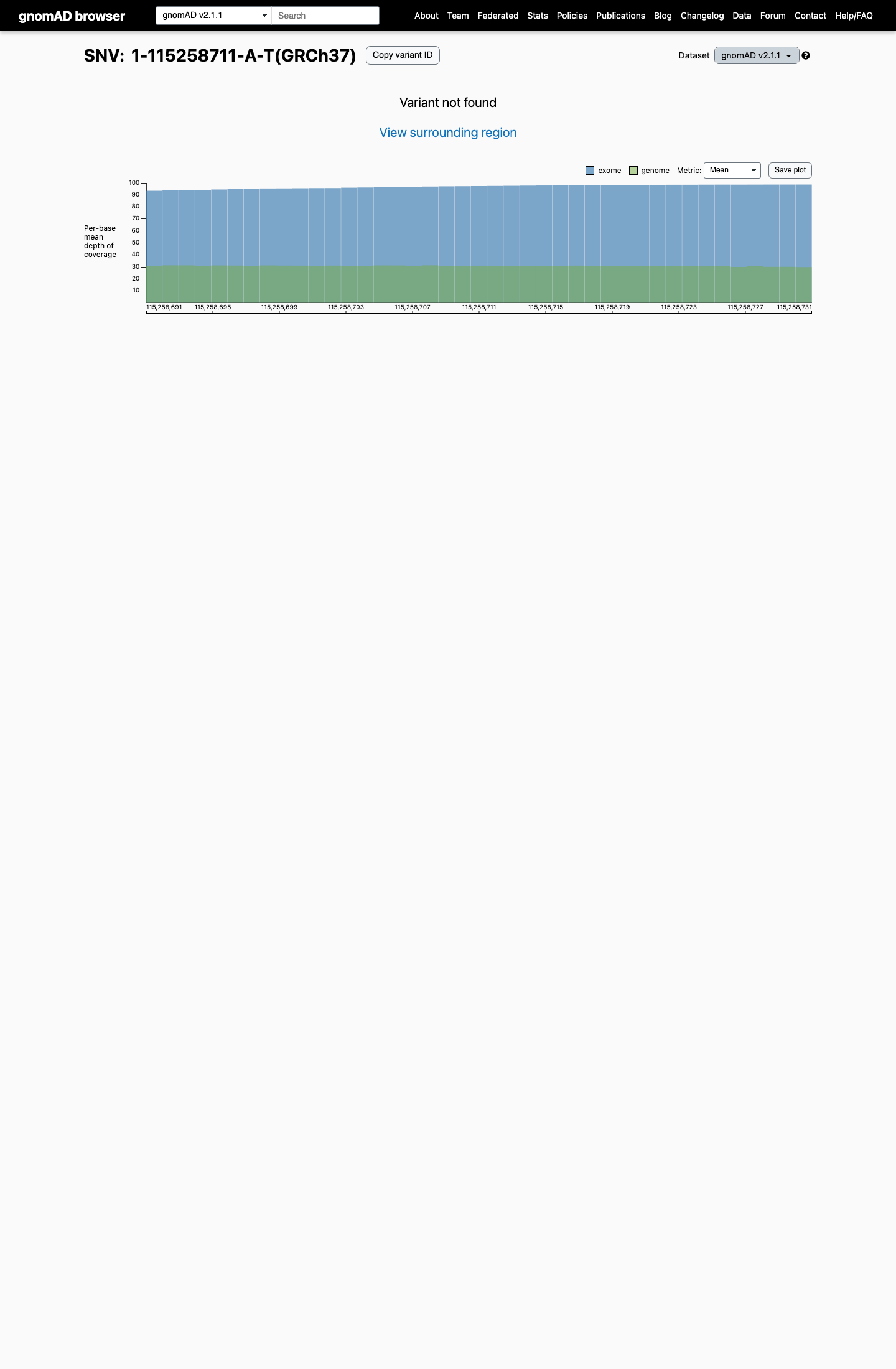
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity under the NRAS RASopathy framework.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In expert-panel-reviewed functional evidence, RAS activation assays showed mildly increased GTP-bound RAS with enhanced MAPK phosphorylation, and a zebrafish model showed developmental and craniofacial defects that were rescued by MEK inhibition, supporting an abnormal gain-of-function effect.
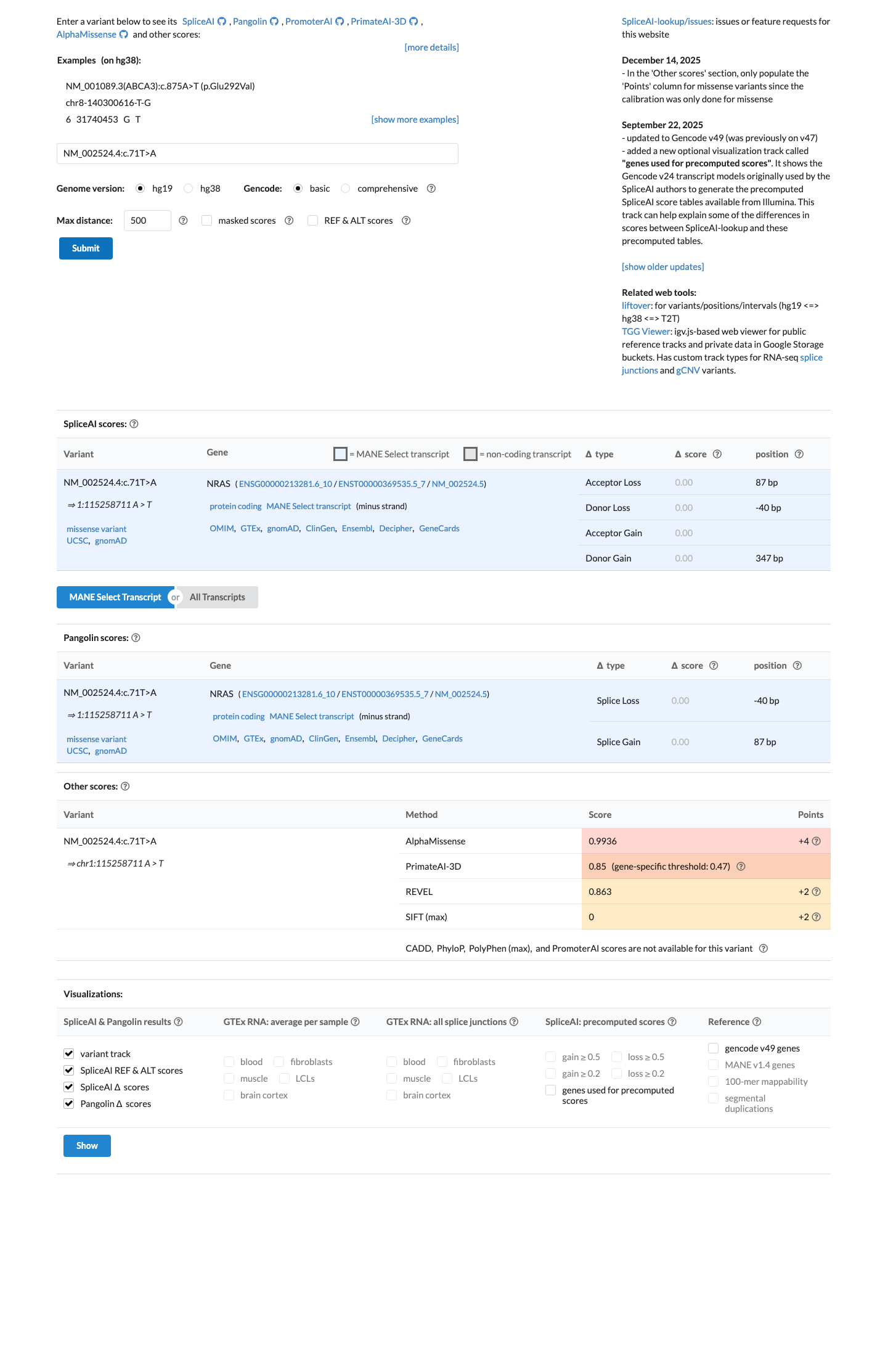
clinvar ↗Computational evidence supports a damaging missense effect, with REVEL 0.863 above the PP3 threshold of 0.7, SpliceAI showing no significant splice impact with a maximum delta score of 0.00, and BayesDel 0.159947.
spliceai ↗ cspec ↗