Classification rationale
1
2
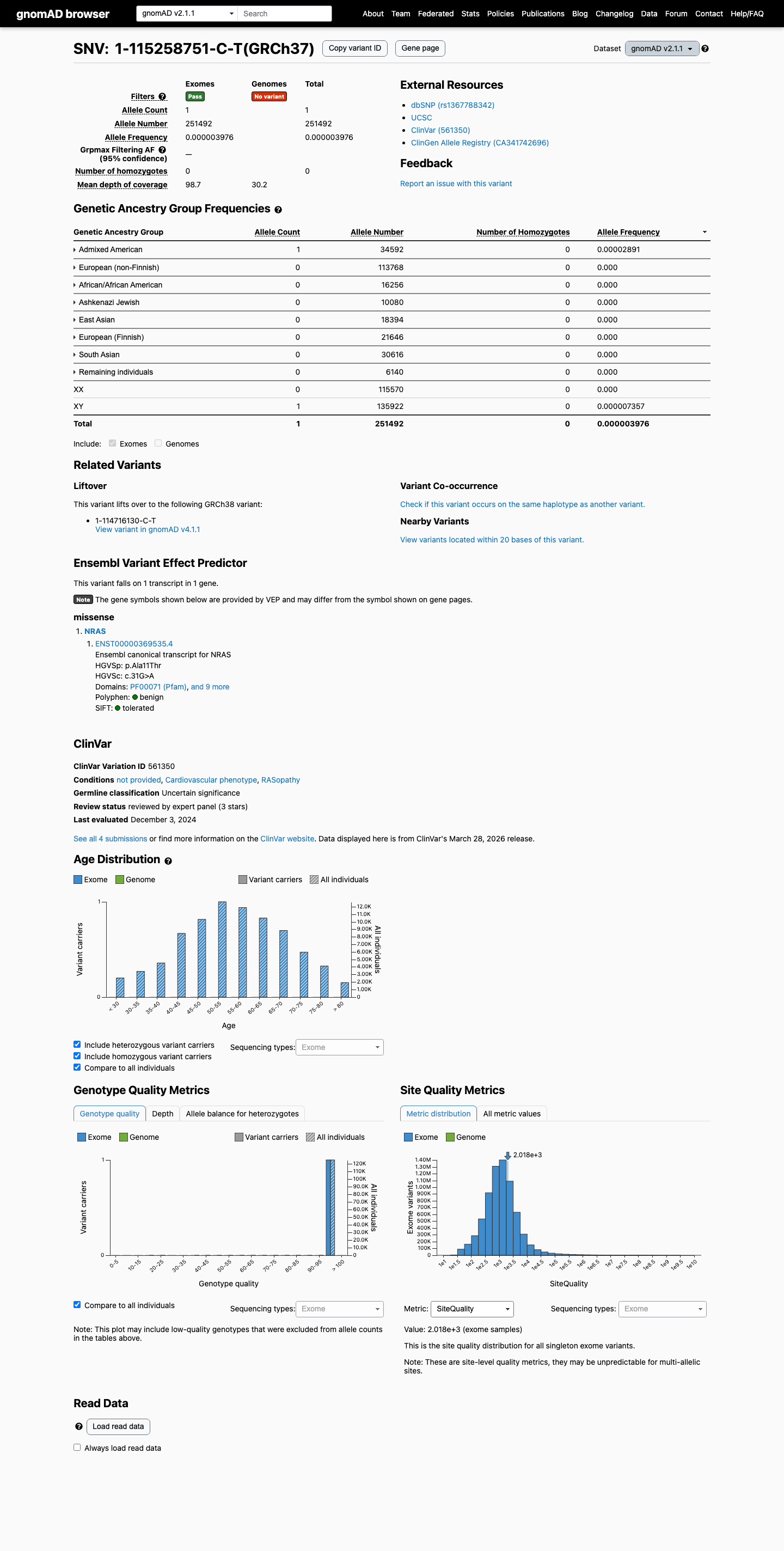
This variant is present at very low frequency in population databases, with 1/251492 alleles in gnomAD v2.1 (AF 0.00040%) and 1/1613858 alleles in gnomAD v4.1 (AF 0.000062%).
gnomad_v2 ↗ gnomad_v4 ↗3
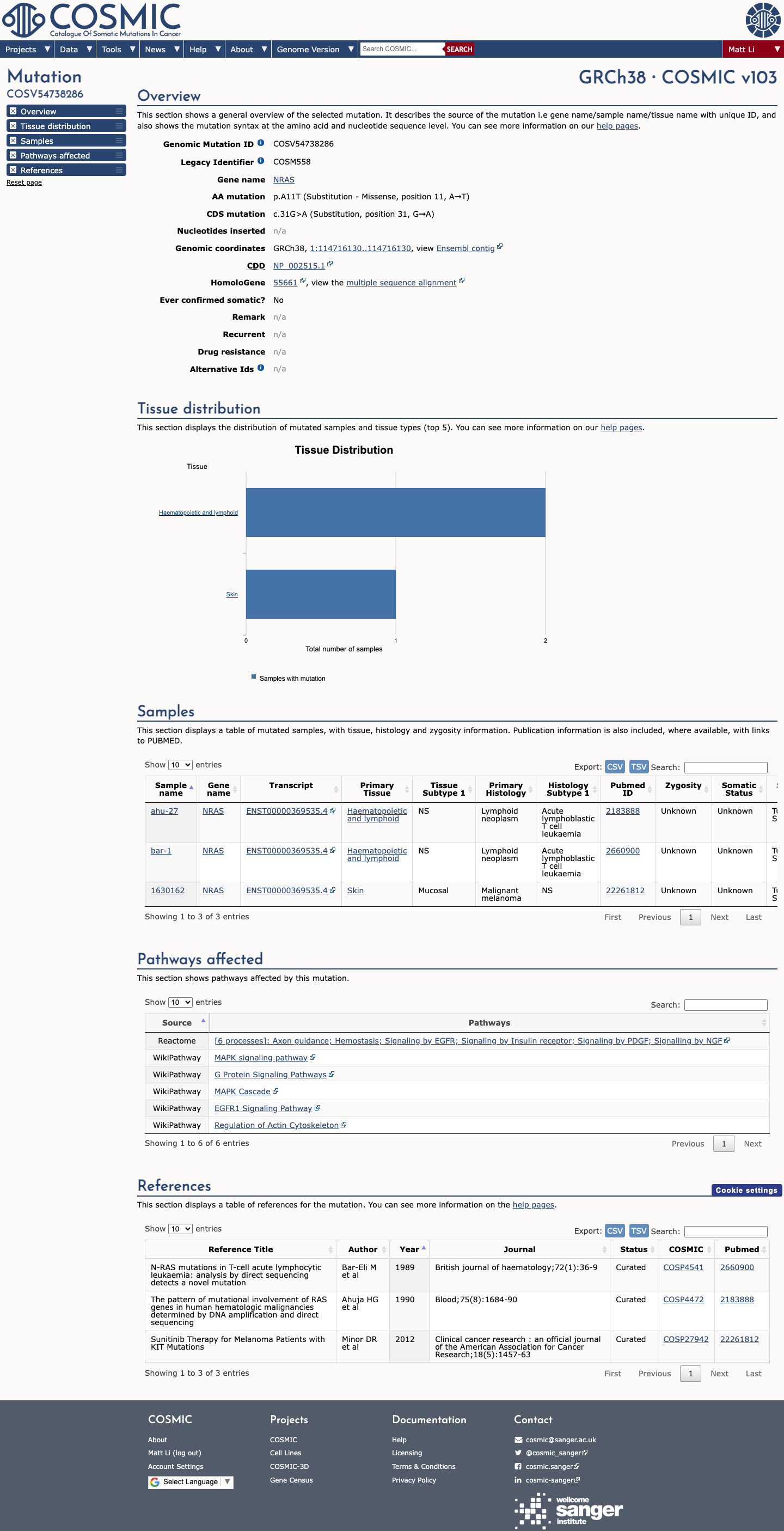
The p.Ala11Thr change lies within the NRAS P-loop (amino acids 10-17), a RASopathy VCEP-specified critical functional domain, which supports PM1.
cspec ↗4
SpliceAI predicts no significant splice impact for this variant (maximum delta score 0.00), the REVEL score is 0.412, and the BayesDel score is 0.00411725; these values do not meet the RASopathy VCEP thresholds for either PP3 or BP4.
cspec ↗ spliceai ↗